[English] 日本語
 Yorodumi
Yorodumi- EMDB-2463: The electron cryo-miscroscopy reconstruction of the connector of ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2463 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The electron cryo-miscroscopy reconstruction of the connector of lactococcal phage p2 | |||||||||
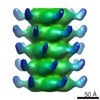
 Map data Map data | Reconstruction of the connector of lactococcal phage p2 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Lactococcus lactis / Siphoviridae / 936 phages / Bacteriophage / electron microscopy / single-particle / p2 | |||||||||
| Biological species |  Lactococcus lactis phage p2 (virus) Lactococcus lactis phage p2 (virus) | |||||||||
| Method | single particle reconstruction / Resolution: 21.0 Å | |||||||||
 Authors Authors | Bebeacua C / Tremblay D / Farenc C / Chapot MP / Sadovskaya I / van Heel M / Veesler D / Moineau S / Cambillau C | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2013 Journal: J Virol / Year: 2013Title: Structure, adsorption to host, and infection mechanism of virulent lactococcal phage p2. Authors: Cecilia Bebeacua / Denise Tremblay / Carine Farenc / Marie-Pierre Chapot-Chartier / Irina Sadovskaya / Marin van Heel / David Veesler / Sylvain Moineau / Christian Cambillau /  Abstract: Lactococcal siphophages from the 936 and P335 groups infect the Gram-positive bacterium Lactococcus lactis using receptor binding proteins (RBPs) attached to their baseplate, a large multiprotein ...Lactococcal siphophages from the 936 and P335 groups infect the Gram-positive bacterium Lactococcus lactis using receptor binding proteins (RBPs) attached to their baseplate, a large multiprotein complex at the distal part of the tail. We have previously reported the crystal and electron microscopy (EM) structures of the baseplates of phages p2 (936 group) and TP901-1 (P335 group) as well as the full EM structure of the TP901-1 virion. Here, we report the complete EM structure of siphophage p2, including its capsid, connector complex, tail, and baseplate. Furthermore, we show that the p2 tail is characterized by the presence of protruding decorations, which are related to adhesins and are likely contributed by the major tail protein C-terminal domains. This feature is reminiscent of the tail of Escherichia coli phage λ and Bacillus subtilis phage SPP1 and might point to a common mechanism for establishing initial interactions with their bacterial hosts. Comparative analyses showed that the architecture of the phage p2 baseplate differs largely from that of lactococcal phage TP901-1. We quantified the interaction of its RBP with the saccharidic receptor and determined that specificity is due to lower k(off) values of the RBP/saccharidic dissociation. Taken together, these results suggest that the infection of L. lactis strains by phage p2 is a multistep process that involves reversible attachment, followed by baseplate activation, specific attachment of the RBPs to the saccharidic receptor, and DNA ejection. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2463.map.gz emd_2463.map.gz | 4.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2463-v30.xml emd-2463-v30.xml emd-2463.xml emd-2463.xml | 9 KB 9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2463.png emd_2463.png | 113.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2463 http://ftp.pdbj.org/pub/emdb/structures/EMD-2463 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2463 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2463 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2463.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2463.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of the connector of lactococcal phage p2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.64 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Connector of Lactococcal phage p2
| Entire | Name: Connector of Lactococcal phage p2 |
|---|---|
| Components |
|
-Supramolecule #1000: Connector of Lactococcal phage p2
| Supramolecule | Name: Connector of Lactococcal phage p2 / type: sample / ID: 1000 / Oligomeric state: Dodecamer / Number unique components: 1 |
|---|
-Supramolecule #1: Lactococcus lactis phage p2
| Supramolecule | Name: Lactococcus lactis phage p2 / type: virus / ID: 1 / Name.synonym: Lactococcal phage p2 Details: The connector was selected from a sample containing entire Lactococcal phages p2. NCBI-ID: 100641 / Sci species name: Lactococcus lactis phage p2 / Virus type: OTHER / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No / Syn species name: Lactococcal phage p2 |
|---|---|
| Host (natural) | Organism: Lactococcal lactis / Strain: NZ9000 / synonym: BACTERIA(EUBACTERIA) |
| Virus shell | Shell ID: 1 / Name: T7 / Diameter: 660 Å / T number (triangulation number): 7 |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 50 mM Tris-HCl, 100 mM NaCl, 8 mM MgSO4 |
|---|---|
| Grid | Details: Quantifoil grids |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200T |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 100,000 times magnification. |
| Date | Jul 1, 2009 |
| Image recording | Category: CCD / Film or detector model: GENERIC TVIPS (4k x 4k) / Digitization - Sampling interval: 4.64 µm / Number real images: 1000 / Average electron dose: 10 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.2 mm / Nominal defocus max: 0.002 µm / Nominal defocus min: 0.001 µm / Nominal magnification: 38000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
- Image processing
Image processing
| CTF correction | Details: Images |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C12 (12 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 21.0 Å / Resolution method: OTHER / Software - Name: IMAGIC / Number images used: 1500 |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















