-検索条件
-検索結果
検索 (著者・登録者: brown & l)の結果733件中、1から50件目までを表示しています

EMDB-17164: 
Structure of human terminal uridylyltransferase 4 (TUT4, ZCCHC11) in complex with pre-let7g miRNA and Lin28A

EMDB-17377: 
Structure of human SIT1 (focussed map / refinement)

EMDB-17378: 
Structure of human SIT1:ACE2 complex (open PD conformation)

EMDB-17379: 
Structure of human SIT1:ACE2 complex (closed PD conformation)

EMDB-17380: 
Structure of human SIT1 bound to L-pipecolate (focussed map / refinement)

EMDB-17381: 
Structure of human SIT1:ACE2 complex (open PD conformation) bound to L-pipecolate

EMDB-17382: 
Structure of human SIT1:ACE2 complex (closed PD conformation) bound to L-pipecolate

PDB-8p2w: 
Structure of human SIT1 (focussed map / refinement)

PDB-8p2x: 
Structure of human SIT1:ACE2 complex (open PD conformation)

PDB-8p2y: 
Structure of human SIT1:ACE2 complex (closed PD conformation)

PDB-8p2z: 
Structure of human SIT1 bound to L-pipecolate (focussed map / refinement)

PDB-8p30: 
Structure of human SIT1:ACE2 complex (open PD conformation) bound to L-pipecolate

PDB-8p31: 
Structure of human SIT1:ACE2 complex (closed PD conformation) bound to L-pipecolate

EMDB-17197: 
Human TPC2 in Complex with Antagonist (S)-SG-094

EMDB-19108: 
Human TPC2 in Complex withAntagonist (R)-SG-094

PDB-8ouo: 
Human TPC2 in Complex with Antagonist (S)-SG-094

EMDB-41271: 
Consensus map of 96nm repeat of human respiratory doublet microtubule, RS1-2 region

EMDB-41152: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus SZ3 in Closed Conformation

PDB-8tc5: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus SZ3 in Closed Conformation

EMDB-41149: 
Cryo-EM Structure of Spike Glycoprotein from Bat Coronavirus WIV1 in Closed Conformation

EMDB-41150: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus 007 in Closed Conformation

PDB-8tc0: 
Cryo-EM Structure of Spike Glycoprotein from Bat Coronavirus WIV1 in Closed Conformation

PDB-8tc1: 
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus 007 in Closed Conformation

EMDB-43889: 
Chlamydomonas reinhardtii mastigoneme (constituent map 1)

EMDB-43890: 
Chlamydomonas reinhardtii mastigoneme (constituent map 2)

EMDB-43891: 
Chlamydomonas reinhardtii mastigoneme (constituent map 3)

EMDB-43892: 
Composite cryo-EM map of the Chlamydomonas reinhardtii mastigoneme

PDB-9b4h: 
Chlamydomonas reinhardtii mastigoneme filament

EMDB-41363: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5

PDB-8tl6: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5

EMDB-43083: 
S. aureus TarL H300N in complex with CDP-ribitol (single tetramer)

EMDB-43084: 
S. aureus TarL H300N in complex with CDP-ribitol (two tetramers with CHAPS micelle)

PDB-8va1: 
S. aureus TarL H300N in complex with CDP-ribitol (single tetramer)
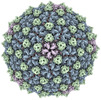
EMDB-40271: 
Mycobacterium phage Adjutor

EMDB-42135: 
CryoEM structure of Sec7 autoinhibited conformation

EMDB-42182: 
Focused map of Sec7 monomer autoinhibited conformation

EMDB-42183: 
Consensus map of Sec7 dimer autoinhibited conformation

PDB-8ucq: 
CryoEM structure of Sec7 autoinhibited conformation

EMDB-43224: 
Structure of the HKU1 RBD bound to the human TMPRSS2 receptor

PDB-8vgt: 
Structure of the HKU1 RBD bound to the human TMPRSS2 receptor

EMDB-41933: 
CryoEM map of horse spleen apoferritin determined as a reference for benchmarking square and rectangular apertures for cryo-EM

EMDB-41936: 
CryoEM map of horse spleen apoferritin determined as a reference for benchmarking square and rectangular apertures for cryo-EM (Falcon IV round beam)

EMDB-41937: 
CryoEM map of horse spleen apoferritin determined for benchmarking square and rectangular apertures for cryo-EM (Falcon IV square beam)

EMDB-41938: 
CryoEM map of horse spleen apoferritin determined for benchmarking square and rectangular apertures for cryo-EM (Gatan K3 rectangular beam)

EMDB-42882: 
Amylin 1 receptor bound to salmon calcitonin (Amy1R:sCT) reconstructed from cryo-EM datasets recorded using a rectangular aperture

EMDB-19067: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).

EMDB-19076: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).

EMDB-19077: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon and EF-Tu(GDP) (structure 3).

PDB-8rd8: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).

PDB-8rdv: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
