-Search query
-Search result
Showing 1 - 50 of 1,122 items for (author: shin & s)

EMDB-37112: 
Cryo-EM structure of human SIDT1 bound to cholesterol
Method: single particle / : Hirano Y, Ohto U, Shimizu T

EMDB-37113: 
Cryo-EM structure of human SIDT1
Method: single particle / : Hirano Y, Ohto U, Shimizu T

PDB-8kcw: 
Cryo-EM structure of human SIDT1 bound to cholesterol
Method: single particle / : Hirano Y, Ohto U, Shimizu T

EMDB-19638: 
YlmH bound to PtRNA-50S
Method: single particle / : Paternoga H, Dimitrova-Paternoga L, Wilson DN

EMDB-19641: 
YlmH bound to stalled 50S subunits with RqcH and PtRNA
Method: single particle / : Paternoga H, Wilson DN

PDB-8s1p: 
YlmH bound to PtRNA-50S
Method: single particle / : Paternoga H, Dimitrova-Paternoga L, Wilson DN

PDB-8s1u: 
YlmH bound to stalled 50S subunits with RqcH and PtRNA
Method: single particle / : Paternoga H, Wilson DN

EMDB-19926: 
CTE type I tau filament from vacuolar tauopathy
Method: helical / : Qi C, Scheres HWS, Goedert M

EMDB-19927: 
CTE type II tau filament from vacuolar tauopathy
Method: helical / : Qi C, Scheres HWS, Goedert M

EMDB-19928: 
CTE type II tau filament from vacuolar tauopathy
Method: helical / : Qi C, Scheres HWS, Goedert M

PDB-9erm: 
CTE type I tau filament from vacuolar tauopathy
Method: helical / : Qi C, Scheres HWS, Goedert M

PDB-9ern: 
CTE type II tau filament from vacuolar tauopathy
Method: helical / : Qi C, Scheres HWS, Goedert M

PDB-9ero: 
CTE type II tau filament from vacuolar tauopathy
Method: helical / : Qi C, Scheres HWS, Goedert M

EMDB-40436: 
48-nm doublet microtubule from Tetrahymena thermophila strain MEC17
Method: single particle / : Black CS, Kubo S, Yang SK, Bui KH

PDB-8sf7: 
48-nm doublet microtubule from Tetrahymena thermophila strain MEC17
Method: single particle / : Black CS, Kubo S, Yang SK, Bui KH
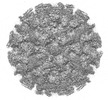
EMDB-36223: 
Cryo-EM structure of the GI.4 Chiba VLP complexed with the CV-1A1 Fv-clasp
Method: single particle / : Hosaka T, Katsura K, Kimura-Someya T, Someya Y, Shirouzu M

PDB-8jg5: 
Cryo-EM structure of the GI.4 Chiba VLP complexed with the CV-1A1 Fv-clasp
Method: single particle / : Hosaka T, Katsura K, Kimura-Someya T, Someya Y, Shirouzu M

EMDB-37850: 
Cryo-EM structure of native H. thermoluteolus TH-1 GroEL
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

EMDB-37853: 
Cryo-EM structure of H. thermoluteolus GroEL-GroES2 football complex
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

EMDB-37862: 
Cryo-EM structure of H. thermophilus GroEL-GroES2 asymmetric football complex
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

EMDB-37863: 
Cryo-EM structure of H. thermophilus GroEL-GroES bullet complex
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

PDB-8wu4: 
Cryo-EM structure of native H. thermoluteolus TH-1 GroEL
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

PDB-8wuc: 
Cryo-EM structure of H. thermoluteolus GroEL-GroES2 football complex
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

PDB-8wuw: 
Cryo-EM structure of H. thermophilus GroEL-GroES2 asymmetric football complex
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

PDB-8wux: 
Cryo-EM structure of H. thermophilus GroEL-GroES bullet complex
Method: single particle / : Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S

EMDB-18534: 
Structure of SecM-stalled Escherichia coli 70S ribosome
Method: single particle / : Gersteuer F, Morici M, Wilson DN

EMDB-18590: 
Cryo-EM map of rotated SecM-stalled Escherichia coli 70S ribosome
Method: single particle / : Gersteuer F, Morici M, Wilson DN

PDB-8qoa: 
Structure of SecM-stalled Escherichia coli 70S ribosome
Method: single particle / : Gersteuer F, Morici M, Wilson DN

EMDB-18320: 
E. coli ApdP-stalled ribosomal complex
Method: single particle / : Morici M, Wilson DN

EMDB-18332: 
B. subtilis ApdA-stalled ribosomal complex
Method: single particle / : Morici M, Wilson DN

EMDB-18340: 
ApdP-SRC with P-tRNA only
Method: single particle / : Morici M, Wilson DN

EMDB-18341: 
ApdA-SRC with P-tRNA only
Method: single particle / : Morici M, Wilson DN

EMDB-17509: 
Cryo-EM structure of CAK in complex with inhibitor BS-181
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17510: 
Cryo-EM structure of CAK in complex with inhibitor BS-194
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17512: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0510-R
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17513: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0510-S
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17514: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0574
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17515: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0768
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17516: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0829
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17517: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0880 (ring-up conformation)
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17518: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0880 (ring-down conformation)
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17519: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0914
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17520: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0943
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17521: 
Cryo-EM structure of CAK in complex with inhibitor dinaciclib
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17522: 
Cryo-EM structure of CAK with averaged inhibitor density
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17523: 
Cryo-EM structure of apo CAK
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17524: 
Cryo-EM map of inhibitor-bound CAK (Krios G4 performance comparison dataset)
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search






 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
