-Search query
-Search result
Showing 1 - 50 of 78 items for (author: hart & fu)

EMDB-18110: 
The fibrillar and amorphous states of polyQ Q97
Method: electron tomography / : Zhao DY

EMDB-18114: 
phagophore in fibrillar polyQ
Method: electron tomography / : Zhao DY
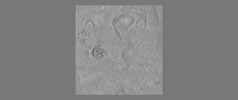
EMDB-18115: 
phagophore and lysosomes with amorphous polyQ
Method: electron tomography / : Zhao DY

EMDB-18116: 
autolysosome, lysosome next to polyQ fibrils are empty
Method: electron tomography / : Zhao DY

EMDB-18117: 
autophagosome and autolysosomes are empty next to fibrillar polyQ
Method: electron tomography / : Zhao DY
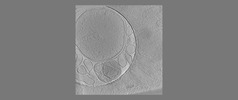
EMDB-18118: 
Isolated autophagosome
Method: electron tomography / : Zhao DY
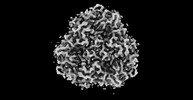
EMDB-32588: 
The Cryo-EM structure of siphonaxanthin chlorophyll a/b type light-harvesting complex II
Method: single particle / : Seki S, Nakaniwa T, Castro-Hartmann P, Sader K, Kawamoto A, Tanaka H, Qian P, Kurisu G, Fujii R

PDB-7wlm: 
The Cryo-EM structure of siphonaxanthin chlorophyll a/b type light-harvesting complex II
Method: single particle / : Seki S, Nakaniwa T, Castro-Hartmann P, Sader K, Kawamoto A, Tanaka H, Qian P, Kurisu G, Fujii R

EMDB-13739: 
Tomogram of a TauRD-YFP aggregate in a TauRD-YFP expressing Hek293T cell
Method: electron tomography / : Guo Q, Saha I, Fernandez-Busnadiego R, Hartl FU

EMDB-13740: 
Tomogram of a TauRD-YFP aggregate in a TauRD-YFP expressing primary mouse neuron
Method: electron tomography / : Trinkaus VA, Saha I, Fernandez-Busnadiego R, Hartl FU

EMDB-14532: 
Structure of P. luminescens TccC3-F-actin complex
Method: single particle / : Belyy A, Raunser S

EMDB-14533: 
Structure of ADP-ribosylated F-actin
Method: single particle / : Belyy A, Raunser S

PDB-7z7h: 
Structure of P. luminescens TccC3-F-actin complex
Method: single particle / : Belyy A, Raunser S

EMDB-32216: 
26S proteasome from the cell with TDP-25 inclusion
Method: subtomogram averaging / : Guo Q

EMDB-32217: 
tomogram of a rat primary neuron harboring TDP-25 inclusion
Method: electron tomography / : Guo Q

EMDB-12730: 
SSUL-gCAL mediating Synechococcus elongatus PCC 7942 M58 homo-demixing
Method: single particle / : Kun Z, Huping W, Ulrich FH, Manajit HH

EMDB-12731: 
Structure of the repeat unit in the network formed by CcmM full length isoform and Rubisco from Synechococcus elongatus
Method: single particle / : Zang K, Huping W, Ulrich FH, Manajit HH

EMDB-12732: 
CryoEM structure of the interaction between CcmM full length isoform (SSUL) domain with RbcL8 core from Synechococcus elongatus PCC 7942
Method: single particle / : Zang K, Huping W, Ulrich FH, Manajit HH

EMDB-11401: 
Tomogram of GFP-a-synuclein inclusion in primary mouse neuron expressing GFP-a-synuclein, seeded with PFFs
Method: electron tomography / : Trinkaus VA, Hartl FU, Fernandez-Busnadiego R

EMDB-11416: 
Tomogram of GFP-a-synuclein inclusion in primary mouse neuron expressing GFP-a-synuclein, seeded with MSA aggregates
Method: electron tomography / : Trinkaus VA, Hartl FU, Fernandez-Busnadiego R

EMDB-11417: 
Tomogram of endogenous a-synuclein inclusion in primary mouse neurons seeded with PFFs
Method: electron tomography / : Trinkaus VA, Hartl FU, Fernandez-Busnadiego R

EMDB-11028: 
CryoEM structure of Rubisco Activase with its substrate Rubisco from Nostoc sp. (strain PCC7120)
Method: single particle / : Wang H, Bracher A, Flecken M, Popilka L, Hartl FU, Hayer-Hartl M

EMDB-11029: 
CryoEM structure of the interaction between Rubisco Activase small-subunit-like (SSUL) domain with Rubisco from Nostoc sp. (strain PCC7120)
Method: single particle / : Wang H, Bracher A

EMDB-11575: 
CryoEM Local map of Rubisco Activase from the complex with its substrate Rubisco from Nostoc sp. (strain PCC7120)
Method: single particle / : Wang H, Bracher A

EMDB-10223: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A

EMDB-10224: 
Cryo-EM Structure of T. kodakarensis 70S ribosome in TkNat10 deleted strain
Method: single particle / : Matzov D, Sas-Chen A

EMDB-10503: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A

PDB-6skf: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A, Thomas JM, Santangelo T, Meier JL, Schwartz S, Shalev-Benami M

PDB-6skg: 
Cryo-EM Structure of T. kodakarensis 70S ribosome in TkNat10 deleted strain
Method: single particle / : Matzov D, Sas-Chen A, Thomas JM, Santangelo T, Meier JL, Schwartz S, Shalev-Benami M

PDB-6th6: 
Cryo-EM Structure of T. kodakarensis 70S ribosome
Method: single particle / : Matzov D, Sas-Chen A, Thomas JM, Santangelo T, Meier JL, Schwartz S, Shalev-Benami M

EMDB-10528: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in the apo state
Method: single particle / : Bracher A, Wang H

EMDB-10529: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in complex with ADP
Method: single particle / : Bracher A, Wang H

EMDB-10530: 
Structure of the chaperonin gp146 from the bacteriophage EL (Pseudomonas aeruginosa) in complex with ATPgammaS
Method: single particle / : Bracher A, Wang H

EMDB-10495: 
Cryo-EM Structure of NADH reduced form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C

EMDB-10496: 
Cryo-EM Structure of as isolated form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C

PDB-6tg9: 
Cryo-EM Structure of NADH reduced form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C, Mittelstaedt G

PDB-6tga: 
Cryo-EM Structure of as isolated form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C, Mittelstaedt G

EMDB-4752: 
Yeast Vms1-60S ribosomal subunit complex (post-state)
Method: single particle / : Su T, Izawa T, Cheng J, Yamashita Y, Berninghausen O, Inada T, Neupert W, Beckmann R

PDB-6r86: 
Yeast Vms1-60S ribosomal subunit complex (post-state)
Method: single particle / : Su T, Izawa T, Cheng J, Yamashita Y, Berninghausen O, Inada T, Neupert W, Beckmann R

EMDB-4751: 
Yeast Vms1 (Q295L)-60S ribosomal subunit complex (pre-state with Arb1)
Method: single particle / : Su T, Izawa T, Cheng J, Yamashita Y, Berninghausen O, Inada T, Neupert W, Beckmann R

EMDB-4753: 
Yeast Vms1 (Q295L)-60S ribosomal subunit complex (pre-state without Arb1)
Method: single particle / : Su T, Izawa T, Cheng J, Yamashita Y, Berninghausen O, Inada T, Neupert W, Beckmann R

PDB-6r84: 
Yeast Vms1 (Q295L)-60S ribosomal subunit complex (pre-state with Arb1)
Method: single particle / : Su T, Izawa T, Cheng J, Yamashita Y, Berninghausen O, Inada T, Neupert W, Beckmann R

PDB-6r87: 
Yeast Vms1 (Q295L)-60S ribosomal subunit complex (pre-state without Arb1)
Method: single particle / : Su T, Izawa T, Cheng J, Yamashita Y, Berninghausen O, Inada T, Neupert W, Beckmann R

EMDB-0180: 
Structure of the repeat unit in the network formed by CcmM and Rubisco from Synechococcus elongatus
Method: single particle / : Wang H, Yan X

EMDB-0015: 
Symmetry-free cryo-EM map of GroEL-actin
Method: single particle / : Milicic G, Balchin D, Hayer-Hartl M, Hartl FU

EMDB-0016: 
Symmetry-free cryo-EM map of TRiC in apo state (nucleotide free)
Method: single particle / : Milicic G, Balchin D, Hayer-Hartl M, Hartl FU

EMDB-0017: 
Symmetry-free cryo-EM map of TRiC-actin
Method: single particle / : Milicic G, Balchin D, Hayer-Hartl M, Hartl FU

EMDB-0018: 
Symmetry-free cryo-EM map of TRiC-actin-alpha_CCT1
Method: single particle / : Milicic G, Balchin D, Hayer-Hartl M, Hartl FU

EMDB-0022: 
Symmetry-free cryo-EM map of TRiC-ADP-BeFx
Method: single particle / : Strauss M, Milicic G, Balchin D, Hayer-Hartl M, Hartl FU
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
