[English] 日本語
 Yorodumi
Yorodumi- EMDB-3517: Structural reorganization of the chromatin remodeling enzyme Chd1... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3517 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structural reorganization of the chromatin remodeling enzyme Chd1 upon engagement with nucleosomes. | |||||||||
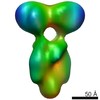
 Map data Map data | S.Cerevisiae Chd1 bound to 601-Nucleosome | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 20.0 Å | |||||||||
 Authors Authors | Sundaramoorthy R / Owen-Hughes T | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: Structural reorganization of the chromatin remodeling enzyme Chd1 upon engagement with nucleosomes. Authors: Ramasubramanian Sundaramoorthy / Amanda L Hughes / Vijender Singh / Nicola Wiechens / Daniel P Ryan / Hassane El-Mkami / Maxim Petoukhov / Dmitri I Svergun / Barbara Treutlein / Salina Quack ...Authors: Ramasubramanian Sundaramoorthy / Amanda L Hughes / Vijender Singh / Nicola Wiechens / Daniel P Ryan / Hassane El-Mkami / Maxim Petoukhov / Dmitri I Svergun / Barbara Treutlein / Salina Quack / Monika Fischer / Jens Michaelis / Bettina Böttcher / David G Norman / Tom Owen-Hughes /   Abstract: The yeast Chd1 protein acts to position nucleosomes across genomes. Here, we model the structure of the Chd1 protein in solution and when bound to nucleosomes. In the apo state, the DNA-binding ...The yeast Chd1 protein acts to position nucleosomes across genomes. Here, we model the structure of the Chd1 protein in solution and when bound to nucleosomes. In the apo state, the DNA-binding domain contacts the edge of the nucleosome while in the presence of the non-hydrolyzable ATP analog, ADP-beryllium fluoride, we observe additional interactions between the ATPase domain and the adjacent DNA gyre 1.5 helical turns from the dyad axis of symmetry. Binding in this conformation involves unravelling the outer turn of nucleosomal DNA and requires substantial reorientation of the DNA-binding domain with respect to the ATPase domains. The orientation of the DNA-binding domain is mediated by sequences in the N-terminus and mutations to this part of the protein have positive and negative effects on Chd1 activity. These observations indicate that the unfavorable alignment of C-terminal DNA-binding region in solution contributes to an auto-inhibited state. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3517.map.gz emd_3517.map.gz | 28.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3517-v30.xml emd-3517-v30.xml emd-3517.xml emd-3517.xml | 18.7 KB 18.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3517_fsc.xml emd_3517_fsc.xml | 7 KB | Display |  FSC data file FSC data file |
| Images |  emd_3517.png emd_3517.png | 40.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3517 http://ftp.pdbj.org/pub/emdb/structures/EMD-3517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3517 | HTTPS FTP |
-Validation report
| Summary document |  emd_3517_validation.pdf.gz emd_3517_validation.pdf.gz | 219.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3517_full_validation.pdf.gz emd_3517_full_validation.pdf.gz | 218.4 KB | Display | |
| Data in XML |  emd_3517_validation.xml.gz emd_3517_validation.xml.gz | 9.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3517 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3517 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3517 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3517 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3517.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3517.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | S.Cerevisiae Chd1 bound to 601-Nucleosome | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.58 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Chd1-Nucleosome complex
| Entire | Name: Chd1-Nucleosome complex |
|---|---|
| Components |
|
-Supramolecule #1: Chd1-Nucleosome complex
| Supramolecule | Name: Chd1-Nucleosome complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: S.Cerevisiae chromatin remodelling enzyme in complex with 601-nucleosome |
|---|---|
| Molecular weight | Theoretical: 400 KDa |
-Supramolecule #2: Chd1
| Supramolecule | Name: Chd1 / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Supramolecule #3: Nucleosome
| Supramolecule | Name: Nucleosome / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism: |
| Recombinant expression | Organism:  |
-Macromolecule #1: Chromodomain Helicase DNA binding protein
| Macromolecule | Name: Chromodomain Helicase DNA binding protein / type: other / ID: 1 / Details: Protein / Classification: other |
|---|---|
| Sequence | String: MAAKDISTEV LQNPELYGLR RSHRAAAHQQ NYFNDSDDED DEDNIKQSRR KRMTTIEDDE DEFEDEEGE EDSGEDEDEE DFEEDDDYYG SPIKQNRSKP KSRTKSKSKS KPKSQSEKQS T VKIPTRFS NRQNKTVNYN IDYSDDDLLE SEDDYGSEEA LSEENVHEAS ...String: MAAKDISTEV LQNPELYGLR RSHRAAAHQQ NYFNDSDDED DEDNIKQSRR KRMTTIEDDE DEFEDEEGE EDSGEDEDEE DFEEDDDYYG SPIKQNRSKP KSRTKSKSKS KPKSQSEKQS T VKIPTRFS NRQNKTVNYN IDYSDDDLLE SEDDYGSEEA LSEENVHEAS ANPQPEDFHG ID IVINHRL KTSLEEGKVL EKTVPDLNNC KENYEFLIKW TDESHLHNTW ETYESIGQVR GLK RLDNYC KQFIIEDQQV RLDPYVTAED IEIMDMERER RLDEFEEFHV PERIIDSQRA SLED GTSQL QYLVKWRRLN YDEATWENAT DIVKLAPEQV KHFQNRENSK ILPQYSSNYT SQRPR FEKL SVQPPFIKGG ELRDFQLTGI NWMAFLWSKG DNGILADEMG LGKTVQTVAF ISWLIF ARR QNGPHIIVVP LSTMPAWLDT FEKWAPDLNC ICYMGNQKSR DTIREYEFYT NPRAKGK KT MKFNVLLTTY EYILKDRAEL GSIKWQFMAV DEAHRLKNAE SSLYESLNSF KVANRMLI T GTPLQNNIKE LAALVNFLMP GRFTIDQEID FENQDEEQEE YIHDLHRRIQ PFILRRLKK DVEKSLPSKT ERILRVELSD VQTEYYKNIL TKNYSALTAG AKGGHFSLLN IMNELKKASN HPYLFDNAE ERVLQKFGDG KMTRENVLRG LIMSSGKMVL LDQLLTRLKK DGHRVLIFSQ M VRMLDILG DYLSIKGINF QRLDGTVPSA QRRISIDHFN SPDSNDFVFL LSTRAGGLGI NL MTADTVV IFDSDWNPQA DLQAMARAHR IGQKNHVMVY RLVSKDTVEE EVLERARKKM ILE YAIISL GVTDGNKYTK KNEPNAGELS AILKFGAGNM FTATDNQKKL EDLNLDDVLN HAED HVTTP DLGESHLGGE EFLKQFEVTD YKADIDWDDI IPEEELKKLQ DEEQKRKDEE YVKEQ LEMM NRRDNALKKI KNSVNGDGTA ANSDSDDDST SRSSRRRARA NDMDSIGESE VRALYK AIL KFGNLKEILD ELIADGTLPV KSFEKYGETY DEMMEAAKDC VHEEEKNRKE ILEKLEK HA TAYRAKLKSG EIKAENQPKD NPLTRLSLKK REKKAVLFNF KGVKSLNAES LLSRVEDL K YLKNLINSNY KDDPLKFSLG NNTPKPVQNW SSNWTKEEDE KLLIGVFKYG YGSWTQIRD DPFLGITDKI FLNEVHNPVA KKSASSSDTT PTPSKKGKGI TGSSKKVPGA IHLGRRVDYL LSFLRGGLN TKSPSADIGS KKLPTGPSKK RQRKPANHSK SMTPEI |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: Solutions were made fresh. | |||||||||
| Grid | Model: Quantifoil / Material: COPPER/RHODIUM / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 101.325 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: Grids are double side blotted with 2sec blotting time and blotting force 10. | |||||||||
| Details | This sample was monodisperse. One Chd1 molecule bound to one Nucleosome. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 20 |
|---|---|
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Details | Preliminary grid screen was done manually and then data collection was performed automatically using EMTools-EMMenu |
| Image recording | Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Sampling interval: 15.6 µm / Number grids imaged: 1 / Number real images: 140 / Average exposure time: 2.0 sec. / Average electron dose: 22.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 5.0 µm / Calibrated defocus min: 1.5 µm / Calibrated magnification: 68000 / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 68000 |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller