2EYC
 
 | |
2EYD
 
 | |
2EYB
 
 | |
2EYA
 
 | |
4WLU
 
 | |
4WLO
 
 | |
4WLF
 
 | | Crystal structure of L-malate bound MDH2 | | Descriptor: | (2S)-2-hydroxybutanedioic acid, Malate dehydrogenase, mitochondrial, ... | | Authors: | Eo, Y.M, Han, B.G, Ahn, H.C. | | Deposit date: | 2014-10-07 | | Release date: | 2015-11-25 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of L-malate bound MDH2
To Be Published
|
|
4WLN
 
 | | Crystal structure of apo MDH2 | | Descriptor: | Malate dehydrogenase, mitochondrial, PHOSPHATE ION | | Authors: | Eo, Y.M, Han, B.G, Ahn, H.C. | | Deposit date: | 2014-10-07 | | Release date: | 2015-11-25 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Crystal structure of apo MDH2
To Be Published
|
|
4WLE
 
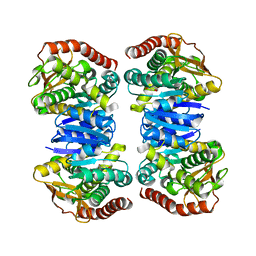 | |
4WLV
 
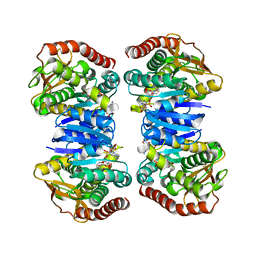 | | Crystal structure of NAD bound MDH2 | | Descriptor: | Malate dehydrogenase, mitochondrial, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Eo, Y.M, Han, B.G, Ahn, H.C. | | Deposit date: | 2014-10-08 | | Release date: | 2015-11-25 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of NAD bound MDH2
To Be Published
|
|
2Z84
 
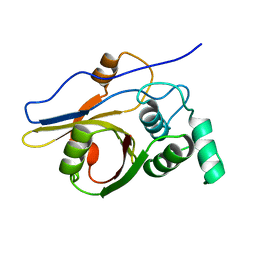 | | Insights from crystal and solution structures of mouse UfSP1 | | Descriptor: | Ufm1-specific protease 1 | | Authors: | Ha, B.H, Ahn, H.C, Kang, S.H, Tanaka, K, Chung, C.H, Kim, E.E. | | Deposit date: | 2007-08-30 | | Release date: | 2008-03-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for Ufm1 processing by UfSP1
J. Biol. Chem., 283, 2008
|
|
4KKH
 
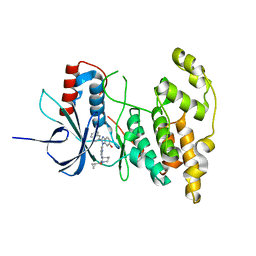 | | The crystal structure of inhibitor-bound JNK3 | | Descriptor: | Mitogen-activated protein kinase 10, cyclopropyl[(3R)-3-({4-[6-hydroxy-2-(naphthalen-2-yl)-1H-benzimidazol-1-yl]pyrimidin-2-yl}amino)piperidin-1-yl]methanone | | Authors: | Han, B.G, Shim, M.B, Ahn, H.C. | | Deposit date: | 2013-05-06 | | Release date: | 2014-05-21 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of AMP, apo, and inhibitor-bound JNK3
To be Published
|
|
4KKE
 
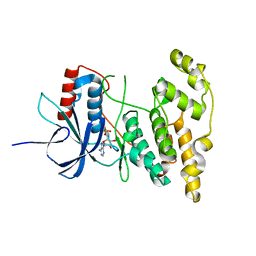 | |
4KKG
 
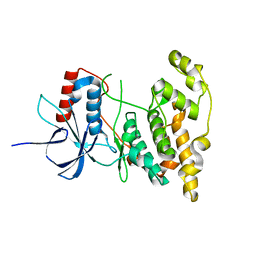 | |
7LUB
 
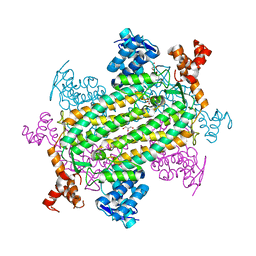 | |
7MBH
 
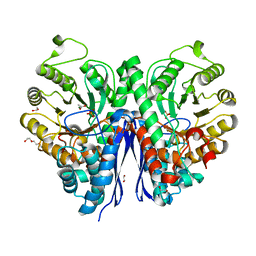 | | Structure of Human Enolase 2 in complex with phosphoserine | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Gamma-enolase, ... | | Authors: | Leonard, P.G, Hicks, K.G, Rutter, J. | | Deposit date: | 2021-03-31 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Protein-metabolite interactomics of carbohydrate metabolism reveal regulation of lactate dehydrogenase.
Science, 379, 2023
|
|
5XV9
 
 | |
2MNU
 
 | |
7E3W
 
 | | Metallo beta-lactamase fold protein (cAMP bound) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE, CHLORIDE ION, ... | | Authors: | Lee, K.-Y, Kim, D.-G, Lee, B.-J. | | Deposit date: | 2021-02-09 | | Release date: | 2022-02-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural and functional studies of SAV1707 from Staphylococcus aureus elucidate its distinct metal-dependent activity and a crucial residue for catalysis.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
7E3V
 
 | | Metallo beta-lactamase fold protein (cAMP free) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, NICKEL (II) ION, ... | | Authors: | Lee, K.-Y, Kim, D.-G, Lee, B.-J. | | Deposit date: | 2021-02-09 | | Release date: | 2022-02-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural and functional studies of SAV1707 from Staphylococcus aureus elucidate its distinct metal-dependent activity and a crucial residue for catalysis.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
6A4L
 
 | | AcrR from Mycobacterium tuberculosis | | Descriptor: | SODIUM ION, SULFATE ION, TetR family transcriptional regulator | | Authors: | Kang, S.M. | | Deposit date: | 2018-06-20 | | Release date: | 2019-06-26 | | Last modified: | 2020-04-29 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The crystal structure of AcrR from Mycobacterium tuberculosis reveals a one-component transcriptional regulation mechanism.
Febs Open Bio, 9, 2019
|
|
6A4W
 
 | | AcrR from Mycobacterium tuberculosis | | Descriptor: | GLYCEROL, SODIUM ION, SULFATE ION, ... | | Authors: | Kang, S.M, Kim, D.H. | | Deposit date: | 2018-06-21 | | Release date: | 2019-06-26 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.587 Å) | | Cite: | The crystal structure of AcrR from Mycobacterium tuberculosis reveals a one-component transcriptional regulation mechanism.
Febs Open Bio, 9, 2019
|
|
