3FMF
 
 | |
3F4F
 
 | | Crystal structure of dUT1p, a dUTPase from Saccharomyces cerevisiae | | Descriptor: | 1,2-ETHANEDIOL, 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Singer, A.U, Evdokimova, E, Kudritska, M, Edwards, A.M, Yakunin, A.F, Savchenko, A. | | Deposit date: | 2008-10-31 | | Release date: | 2008-11-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and activity of the Saccharomyces cerevisiae dUTP pyrophosphatase DUT1, an essential housekeeping enzyme.
Biochem.J., 437, 2011
|
|
3F8M
 
 | | Crystal Structure of PhnF from Mycobacterium smegmatis | | Descriptor: | GLYCEROL, GntR-family protein transcriptional regulator | | Authors: | Busby, J.N, Gebhard, S, Cook, G.M, Lott, S.J, Baker, E.N, Money, V.A. | | Deposit date: | 2008-11-12 | | Release date: | 2009-11-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of PhnF, a GntR-family transcription regulator in Mycobacterium smegmatis
To be Published
|
|
3FZ1
 
 | | Crystal structure of a benzthiophene inhibitor bound to human Cyclin-dependent Kinase-2 (CDK-2) | | Descriptor: | (3R)-3-(aminomethyl)-9-methoxy-1,2,3,4-tetrahydro-5H-[1]benzothieno[3,2-e][1,4]diazepin-5-one, Cell division protein kinase 2 | | Authors: | Kurumbail, R.G, Caspers, N. | | Deposit date: | 2009-01-23 | | Release date: | 2009-04-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Benzothiophene inhibitors of MK2. Part 2: improvements in kinase selectivity and cell potency.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3GBL
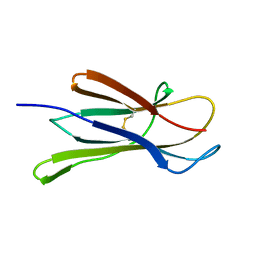 
 | |
3G5Y
 
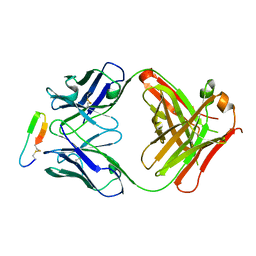 | |
3GDB
 
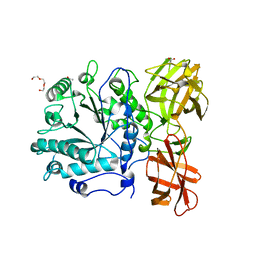 | | Crystal structure of Spr0440 glycoside hydrolase domain, Endo-D from Streptococcus pneumoniae R6 | | Descriptor: | ACETIC ACID, Putative uncharacterized protein spr0440, TRIETHYLENE GLYCOL | | Authors: | Jiang, Y.-L, Frolet, C, Di-guilmi, A.-M, Zhou, C.-Z, Vernet, T, Chen, Y.-X. | | Deposit date: | 2009-02-24 | | Release date: | 2009-03-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Crystal structure of Spr0440 glycoside hydrolase domain, Endo-D from Streptococcus pneumoniae R6
To be published
|
|
7XOL
 
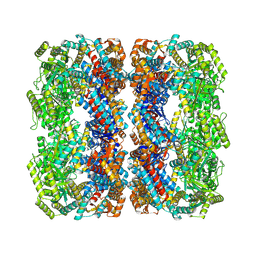 | |
7XOP
 
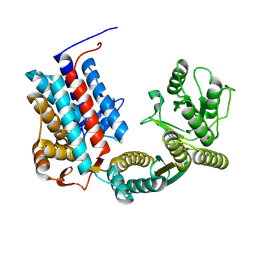 | |
7XOQ
 
 | |
7XOO
 
 | |
7XOS
 
 | |
7XOR
 
 | |
3GIZ
 
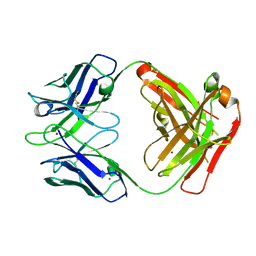 | | Crystal structure of the Fab fragment of anti-CD20 antibody Ofatumumab | | Descriptor: | Fab fragment of anti-CD20 antibody Ofatumumab, heavy chain, light chain, ... | | Authors: | Du, J, Yang, H, Ding, J. | | Deposit date: | 2009-03-07 | | Release date: | 2009-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of the Fab fragment of therapeutic antibody Ofatumumab provides insights into the recognition mechanism with CD20
Mol.Immunol., 46, 2009
|
|
7XOJ
 
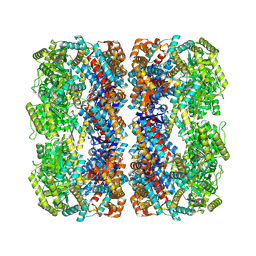 | |
7XON
 
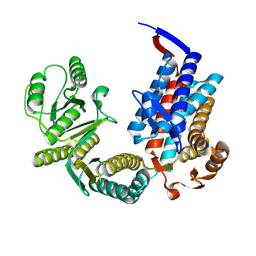 | |
7XOM
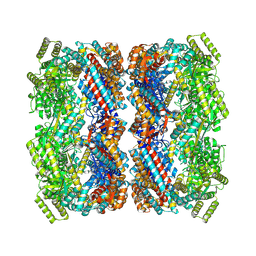 
 | |
7XOK
 
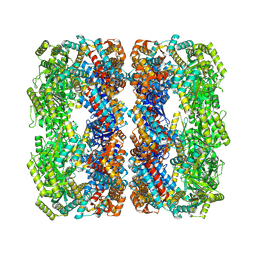 | |
1V57
 
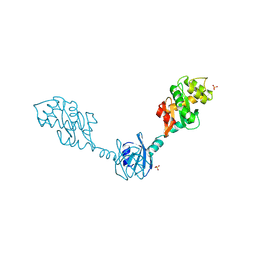 | | Crystal Structure of the Disulfide Bond Isomerase DsbG | | Descriptor: | SULFATE ION, Thiol:disulfide interchange protein dsbG | | Authors: | Heras, B, Edeling, M.A, Schirra, H.J, Raina, S, Martin, J.L. | | Deposit date: | 2003-11-21 | | Release date: | 2004-06-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of the DsbG disulfide isomerase reveal an unstable disulfide
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
3DGQ
 
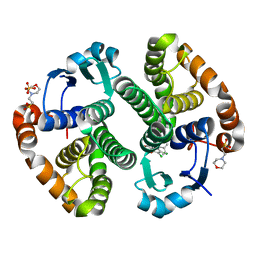 | |
3DU0
 
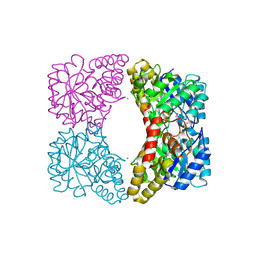 | | E. coli dihydrodipicolinate synthase with first substrate, pyruvate, bound in active site | | Descriptor: | CHLORIDE ION, Dihydrodipicolinate synthase, GLYCEROL, ... | | Authors: | Dobson, R.C.J, Devenish, S.R.A, Gerrard, J.A, Jameson, G.B. | | Deposit date: | 2008-07-16 | | Release date: | 2008-11-18 | | Last modified: | 2019-12-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The high-resolution structure of dihydrodipicolinate synthase from Escherichia coli bound to its first substrate, pyruvate.
Acta Crystallogr.,Sect.F, 64, 2008
|
|
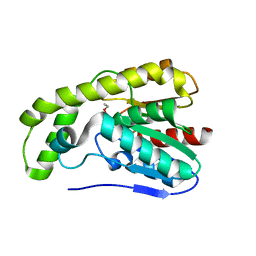
3E1G
 
 | |
3DJ4
 
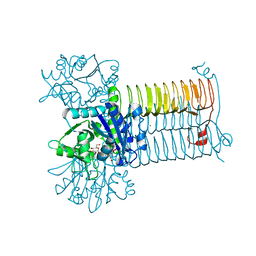 | | Crystal Structure of GlmU from Mycobacterium tuberculosis in complex with URIDINE-DIPHOSPHATE-N-ACETYLGLUCOSAMINE. | | Descriptor: | Bifunctional protein glmU, COBALT (II) ION, MAGNESIUM ION, ... | | Authors: | Verma, S.K, Prakash, B. | | Deposit date: | 2008-06-22 | | Release date: | 2009-05-19 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | PknB-mediated phosphorylation of a novel substrate, N-acetylglucosamine-1-phosphate uridyltransferase, modulates its acetyltransferase activity.
J.Mol.Biol., 386, 2009
|
|
3E17
 
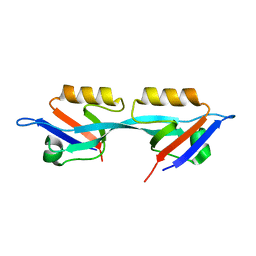 | | Crystal structure of the second PDZ domain from human Zona Occludens-2 | | Descriptor: | Tight junction protein ZO-2 | | Authors: | Chen, H, Tong, S.L, Teng, M.K, Niu, L.W. | | Deposit date: | 2008-08-01 | | Release date: | 2009-07-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure of the second PDZ domain from human zonula occludens 2
Acta Crystallogr.,Sect.F, 65, 2009
|
|
3GS0
 
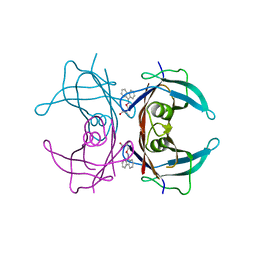 | | Human transthyretin (TTR) complexed with (S)-3-(9H-fluoren-9-ylideneaminooxy)-2-methylpropanoic acid (inhibitor 16) | | Descriptor: | (2S)-3-[(9H-fluoren-9-ylideneamino)oxy]-2-methylpropanoic acid, Transthyretin | | Authors: | Mohamedmohaideen, N.N, Palaninathan, S.K, Orlandini, E, Sacchettini, J.C. | | Deposit date: | 2009-03-26 | | Release date: | 2009-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Novel transthyretin amyloid fibril formation inhibitors: synthesis, biological evaluation, and X-ray structural analysis.
Plos One, 4, 2009
|
|
