8TU8
 | |
8PFQ
 
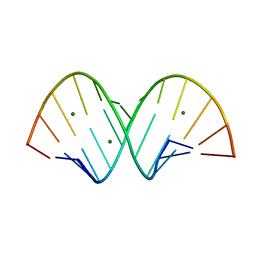 | | RNA structure with 1-methylpseudouridine, P1 space group | | Descriptor: | Chains: A,B, MAGNESIUM ION | | Authors: | Spingler, B, McAuley, K, Nievergelt, P, Thorn, A. | | Deposit date: | 2023-06-16 | | Release date: | 2024-01-31 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.01 Å) | | Cite: | RNA oligomers at atomic resolution containing 1-methylpseudouridine, an essential building block of mRNA vaccines.
Chemmedchem, 19, 2024
|
|
9N25
 
 | |
8TM8
 
 | |
9NEN
 
 | |
8PD0
 
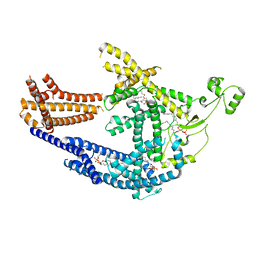 | | cryo-EM structure of Doa10 in MSP1E3D1 | | Descriptor: | 1,2-DIOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1,2-DIPALMITOYL-SN-GLYCERO-3-PHOSPHATE, ERAD-associated E3 ubiquitin-protein ligase DOA10 | | Authors: | Botsch, J.J, Braeuning, B, Schulman, B.A. | | Deposit date: | 2023-06-11 | | Release date: | 2024-01-17 | | Method: | ELECTRON MICROSCOPY (3.58 Å) | | Cite: | Doa10/MARCH6 architecture interconnects E3 ligase activity with lipid-binding transmembrane channel to regulate SQLE.
Nat Commun, 15, 2024
|
|
8TIL
 
 | |
9NFA
 
 | | Cryo-EM structure of Ro60/La/minimal misfolded pre-5S rRNA complex with Fab, composite map | | Descriptor: | Lupus La protein, MAGNESIUM ION, Minimal misfolded pre-5S rRNA, ... | | Authors: | Nam, H, Deme, J.C, Lea, S.M, Wolin, S.L. | | Deposit date: | 2025-02-21 | | Release date: | 2026-02-04 | | Last modified: | 2026-03-04 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Mechanistic insights into RNA chaperoning by Ro60 and La autoantigens.
Cell, 189, 2026
|
|
8PQN
 
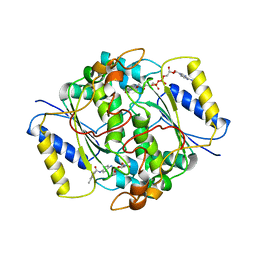 | | NQO1 bound to RBS-10 | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, N-[4-(3-methylbenzamido)phenyl]-5-nitrofuran-2-carboxamide, NAD(P)H dehydrogenase [quinone] 1 | | Authors: | Pous, J, Jose-Duran, F, Mayor-Ruiz, C, Riera, A. | | Deposit date: | 2023-07-11 | | Release date: | 2024-01-10 | | Last modified: | 2026-04-15 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Discovery and Mechanistic Elucidation of NQO1-Bioactivatable Small Molecules That Overcome Resistance to Degraders.
Angew.Chem.Int.Ed.Engl., 63, 2024
|
|
9NBC
 
 | |
8TQ4
 
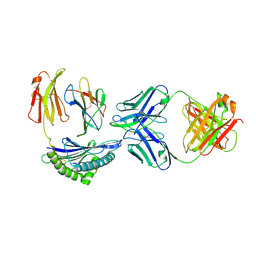 | | Crystal structure of Fab M142 in complex with MHC-I (H2-Dd) | | Descriptor: | Beta-2-microglobulin, Fab M142 Heavy Chain, Fab M142 Light Chain, ... | | Authors: | Jiang, J, Natarajan, K, Boyd, L.F, Margulies, D.H. | | Deposit date: | 2023-08-06 | | Release date: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (3.59 Å) | | Cite: | Crystal structure of Fab M142 in complex with MHC-I (H2-Dd)
To Be Published
|
|
8PTE
 
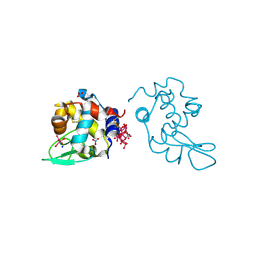 | | Polyoxidovanadate interaction with proteins: crystal structure of lysozyme bound to octadecavanadate ion (structure B) | | Descriptor: | Lysozyme C, NITRATE ION, Polyoxidovanadate complex, ... | | Authors: | Tito, G, Ferraro, G, Merlino, A. | | Deposit date: | 2023-07-14 | | Release date: | 2024-01-10 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.797 Å) | | Cite: | Stabilization and Binding of [V 4 O 12 ] 4- and Unprecedented [V 20 O 54 (NO 3 )] n- to Lysozyme upon Loss of Ligands and Oxidation of the Potential Drug V IV O(acetylacetonato) 2.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8TIO
 
 | |
8PMP
 
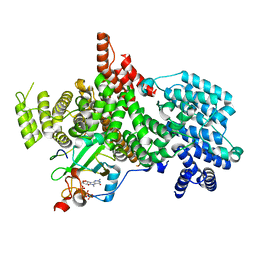 | | Structure of the human nuclear cap-binding complex bound to ARS2[147-871] and m7GTP | | Descriptor: | 7N-METHYL-8-HYDROGUANOSINE-5'-TRIPHOSPHATE, Nuclear cap-binding protein subunit 1, Nuclear cap-binding protein subunit 2, ... | | Authors: | Dubiez, E, Pellegrini, E, Foucher, A.E, Cusack, S, Kadlec, J. | | Deposit date: | 2023-06-29 | | Release date: | 2024-01-17 | | Method: | ELECTRON MICROSCOPY (3.43 Å) | | Cite: | Structural basis for competitive binding of productive and degradative co-transcriptional effectors to the nuclear cap-binding complex.
Cell Rep, 43, 2024
|
|
9NDD
 
 | |
8TII
 
 | |
8PO5
 | | Lactobacillus plantarum LpdD | | Descriptor: | MANGANESE (II) ION, Protein LpdD | | Authors: | Gahloth, D, Leys, D. | | Deposit date: | 2023-07-03 | | Release date: | 2024-01-17 | | Last modified: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The prFMNH 2 -binding chaperone LpdD assists UbiD decarboxylase activation.
J.Biol.Chem., 300, 2024
|
|
9NLM
 
 | |
8U8D
 
 | | Cryo-EM structure of the TREX-2 complex in complex with the N-terminal motif of Sub2 | | Descriptor: | 26S proteasome complex subunit SEM1, Nuclear mRNA export factor, RNA helicase, ... | | Authors: | Clarke, B.P, Xie, Y, Ren, Y. | | Deposit date: | 2023-09-17 | | Release date: | 2025-02-05 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (3.04 Å) | | Cite: | Structures and mRNP remodeling mechanism of the TREX-2 complex.
Structure, 33, 2025
|
|
9NEJ
 
 | | AZD-3 bound EFPA transporter of Mycobacterium tuberculosis | | Descriptor: | (2P)-2-(2-methoxyphenyl)-N-(3-methoxyphenyl)-1-methyl-1H-pyrrolo[3,2-c]pyridin-4-amine, (2R)-3-{[(S)-[(2R)-2,3-dihydroxypropoxy](hydroxy)phosphoryl]oxy}-2-{[(10E)-hexadec-10-enoyl]oxy}propyl (9E,14E)-octadeca-9,14-dienoate, (2S)-3-{[(R)-[(2R)-2,3-dihydroxypropoxy](hydroxy)phosphoryl]oxy}-2-{[(2E,9E,13E)-hexadeca-2,9,13-trienoyl]oxy}propyl (4E,9E,11Z,15E)-octadeca-4,9,11,15-tetraenoate, ... | | Authors: | Khandelwal, N.K, Gupta, M, Stroud, R.M. | | Deposit date: | 2025-02-19 | | Release date: | 2026-03-04 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | AZD-3 bound EFPA transporter of Mycobacterium tuberculosis
To Be Published
|
|
8PZH
 
 | | LpdD (H61A) mutant | | Descriptor: | MANGANESE (II) ION, PHOSPHATE ION, Protein LpdD | | Authors: | Gahloth, D, Leys, D. | | Deposit date: | 2023-07-27 | | Release date: | 2024-01-17 | | Last modified: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | The prFMNH 2 -binding chaperone LpdD assists UbiD decarboxylase activation.
J.Biol.Chem., 300, 2024
|
|
8TIN
 
 | |
9MZM
 
 | |
8Q5L
 
 | |
9NJ0
 
 | |
