4OKA
 
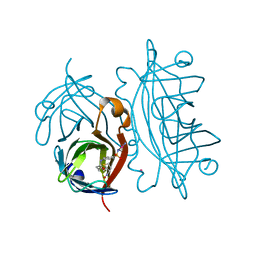 | | Structural-, Kinetic- and Docking Studies of Artificial Imine Reductases Based on the Biotin-Streptavidin Technology: An Induced Lock-and-Key Hypothesis | | Descriptor: | IRIDIUM ION, Streptavidin, [N-(4-{[2-(amino-kappaN)ethyl]sulfamoyl-kappaN}phenyl)-5-(2-oxohexahydro-1H-thieno[3,4-d]imidazol-4-yl)pentanamidato]iridium(III) | | Authors: | Schirmer, T, Heinisch, T. | | Deposit date: | 2014-01-22 | | Release date: | 2014-11-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.505 Å) | | Cite: | Structural, Kinetic, and Docking Studies of Artificial Imine Reductases Based on Biotin-Streptavidin Technology: An Induced Lock-and-Key Hypothesis
J.Am.Chem.Soc., 136, 2014
|
|
4K46
 
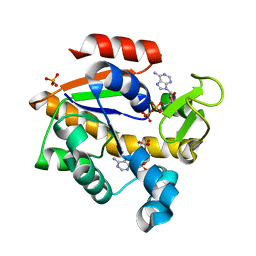 | | Crystal Structure of Adenylate Kinase from Photobacterium profundum | | Descriptor: | ADENOSINE MONOPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, Adenylate kinase, ... | | Authors: | Cho, Y.-J, Kerns, S.J, Kern, D. | | Deposit date: | 2013-04-12 | | Release date: | 2014-07-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Alike but Different: Adenylate kinases from E. Coli, Aquifex, and P. profundum
To be Published
|
|
4KE6
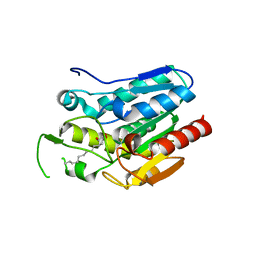 
 | | Crystal structure D196N mutant of Monoglyceride lipase from Bacillus sp. H257 in complex with 1-rac-lauroyl glycerol | | Descriptor: | (2R)-2,3-dihydroxypropyl dodecanoate, (4S)-2-METHYL-2,4-PENTANEDIOL, Thermostable monoacylglycerol lipase | | Authors: | Rengachari, S, Aschauer, P, Gruber, K, Dreveny, I, Oberer, M. | | Deposit date: | 2013-04-25 | | Release date: | 2013-09-18 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Conformational plasticity and ligand binding of bacterial monoacylglycerol lipase.
J.Biol.Chem., 288, 2013
|
|
2RKO
 
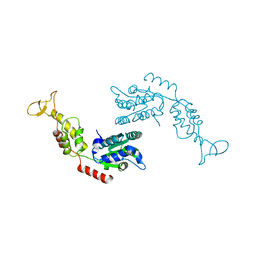 | | Crystal Structure of the Vps4p-dimer | | Descriptor: | Vacuolar protein sorting-associated protein 4 | | Authors: | Hartmann, C, Gruetter, M.G. | | Deposit date: | 2007-10-17 | | Release date: | 2008-02-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.35 Å) | | Cite: | Vacuolar protein sorting: two different functional states of the AAA-ATPase Vps4p
J.Mol.Biol., 377, 2008
|
|
4K7D
 
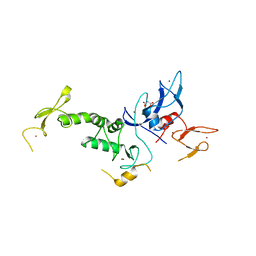 | | Crystal Structure of Parkin C-terminal RING domains | | Descriptor: | CHLORIDE ION, E3 ubiquitin-protein ligase parkin, MALONATE ION, ... | | Authors: | Sauve, V, Trempe, J.-F, Menade, M, Gehring, K. | | Deposit date: | 2013-04-17 | | Release date: | 2013-05-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of parkin reveals mechanisms for ubiquitin ligase activation.
Science, 340, 2013
|
|
4O7Y
 
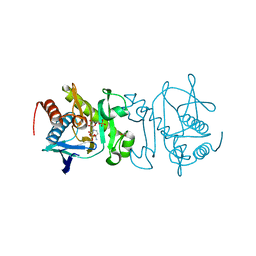 | |
4KGH
 
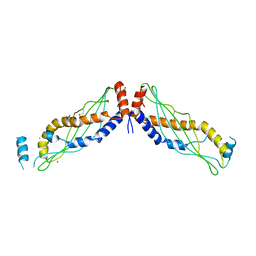 | |
4OJI
 
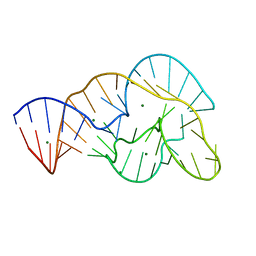 | | Crystal Structure of Twister Ribozyme | | Descriptor: | MAGNESIUM ION, RNA (52-MER) | | Authors: | Liu, Y, Wilson, T.J, McPhee, S.A, Lilley, D.M.J. | | Deposit date: | 2014-01-21 | | Release date: | 2014-07-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure and mechanistic investigation of the twister ribozyme.
Nat.Chem.Biol., 10, 2014
|
|
4MAZ
 
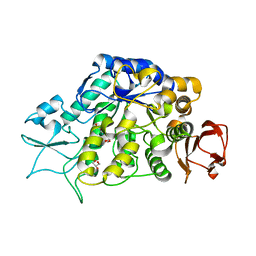 | | The Structure of MalL mutant enzyme V200S from Bacillus subtilus | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Hobbs, J.K, Jiao, W, Easter, A.D, Parker, E.J, Schipper, L.A, Arcus, V.L. | | Deposit date: | 2013-08-18 | | Release date: | 2013-09-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Change in heat capacity for enzyme catalysis determines temperature dependence of enzyme catalyzed rates.
Acs Chem.Biol., 8, 2013
|
|
4MB1
 
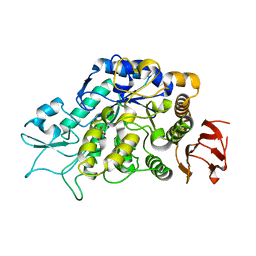 | | The Structure of MalL mutant enzyme G202P from Bacillus subtilus | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, Oligo-1,6-glucosidase 1 | | Authors: | Hobbs, J.K, Jiao, W, Easter, A.D, Parker, E.J, Schipper, L.A, Arcus, V.L. | | Deposit date: | 2013-08-19 | | Release date: | 2013-09-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Change in heat capacity for enzyme catalysis determines temperature dependence of enzyme catalyzed rates.
Acs Chem.Biol., 8, 2013
|
|
4QHM
 
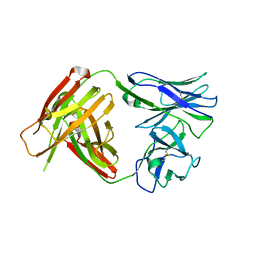 | | I3.1 (unbound) from CH103 Lineage | | Descriptor: | I2 light chain, I3 heavy chain | | Authors: | Fera, D, Harrison, S.C. | | Deposit date: | 2014-05-28 | | Release date: | 2014-06-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.23 Å) | | Cite: | Affinity maturation in an HIV broadly neutralizing B-cell lineage through reorientation of variable domains.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
4M0H
 
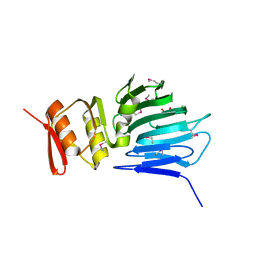 | |
4M0N
 
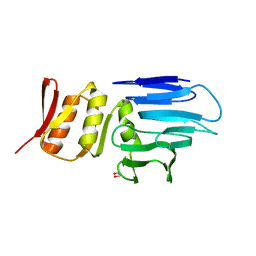 | |
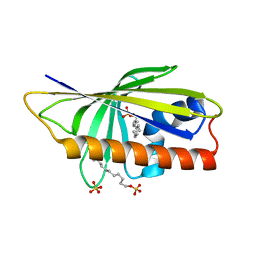
4QIP
 
 | |
4MFR
 
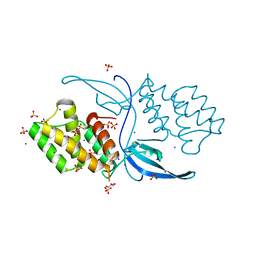 | | Crystal structure of Mycobacterium tuberculosis CarD | | Descriptor: | GLYCEROL, IODIDE ION, RNA polymerase-binding transcription factor CarD, ... | | Authors: | Kaur, G, Thakur, K.G. | | Deposit date: | 2013-08-28 | | Release date: | 2013-11-06 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of Mycobacterium tuberculosis CarD, an essential RNA polymerase binding protein, reveals a quasidomain-swapped dimeric structural architecture.
Proteins, 82, 2014
|
|
4M8U
 
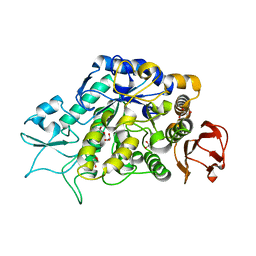 | | The Structure of MalL mutant enzyme V200A from Bacillus subtilus | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, GLYCEROL, ... | | Authors: | Hobbs, J.K, Jiao, W, Easter, A.D, Parker, E.J, Schipper, L.A, Arcus, V.L. | | Deposit date: | 2013-08-13 | | Release date: | 2013-09-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Change in heat capacity for enzyme catalysis determines temperature dependence of enzyme catalyzed rates.
Acs Chem.Biol., 8, 2013
|
|
4M99
 
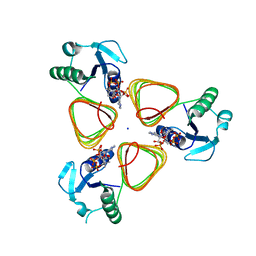 | |
4MCJ
 
 | |
4QOA
 
 | |
4MDB
 
 | | Structure of Mos1 transposase catalytic domain and Raltegravir with Mg | | Descriptor: | MAGNESIUM ION, Mariner Mos1 transposase, N-(4-fluorobenzyl)-5-hydroxy-1-methyl-2-(1-methyl-1-{[(5-methyl-1,3,4-oxadiazol-2-yl)carbonyl]amino}ethyl)-6-oxo-1,6-di hydropyrimidine-4-carboxamide | | Authors: | Richardson, J.M. | | Deposit date: | 2013-08-22 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis of Mos1 Transposase Inhibition by the Anti-retroviral Drug Raltegravir.
Acs Chem.Biol., 9, 2014
|
|
4MHX
 
 | | Crystal Structure of Sulfamidase | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Sidhu, N.S, Uson, I, Schreiber, K, Proepper, K, Becker, S, Gaertner, J, Kraetzner, R, Steinfeld, R, Sheldrick, G.M. | | Deposit date: | 2013-08-30 | | Release date: | 2014-05-14 | | Last modified: | 2021-06-02 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of sulfamidase provides insight into the molecular pathology of mucopolysaccharidosis IIIA.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4MDA
 
 | | Structure of Mos1 transposase catalytic domain and Raltegravir with Mn | | Descriptor: | MANGANESE (II) ION, Mariner Mos1 transposase, N-(4-fluorobenzyl)-5-hydroxy-1-methyl-2-(1-methyl-1-{[(5-methyl-1,3,4-oxadiazol-2-yl)carbonyl]amino}ethyl)-6-oxo-1,6-di hydropyrimidine-4-carboxamide | | Authors: | Richardson, J.M. | | Deposit date: | 2013-08-22 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis of Mos1 Transposase Inhibition by the Anti-retroviral Drug Raltegravir.
Acs Chem.Biol., 9, 2014
|
|
4LW5
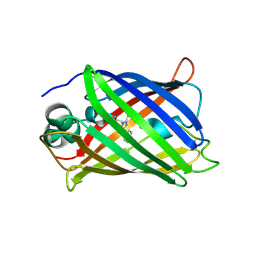 
 | | Crystal structure of all-trans green fluorescent protein | | Descriptor: | Green fluorescent protein | | Authors: | Rosenman, D.J, Huang, Y.-M, Xia, K, Vanroey, P, Colon, W, Bystroff, C. | | Deposit date: | 2013-07-26 | | Release date: | 2014-02-05 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Green-lighting green fluorescent protein: Faster and more efficient folding by eliminating a cis-trans peptide isomerization event.
Protein Sci., 23, 2014
|
|
4QU7
 
 | |
4LXS
 
 | | Structure of the Toll - Spatzle complex, a molecular hub in Drosophila development and innate immunity (glycosylated form) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Stelter, M, Parthier, C, Breithaupt, C, Stubbs, M.T. | | Deposit date: | 2013-07-30 | | Release date: | 2014-04-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of the Toll-Spatzle complex, a molecular hub in Drosophila development and innate immunity.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
