5VDS
 
 | |
8VCG
 
 | | Mutant Src SH2 domain in complex with phosphotyrosine | | Descriptor: | Isoform 3 of Proto-oncogene tyrosine-protein kinase Src, O-PHOSPHOTYROSINE | | Authors: | Hashem, A, Nyovanie, S, Oltean, N, Patskovska, L, Patskovsky, Y, Krogsgaard, M. | | Deposit date: | 2023-12-14 | | Release date: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | Engineered Mutant Src Sh2 Domain
To Be Published
|
|
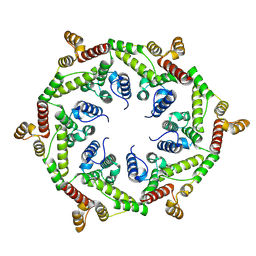
8VW3
 | |
5I9I
 
 | | Crystal structure of LP_PLA2 in complex with Darapladib | | Descriptor: | N-[2-(diethylamino)ethyl]-2-{2-[(4-fluorobenzyl)sulfanyl]-4-oxo-4,5,6,7-tetrahydro-1H-cyclopenta[d]pyrimidin-1-yl}-N-{[ 4'-(trifluoromethyl)biphenyl-4-yl]methyl}acetamide, Platelet-activating factor acetylhydrolase, SULFATE ION | | Authors: | Liu, Q.F, Xu, Y.C. | | Deposit date: | 2016-02-20 | | Release date: | 2016-06-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural and Thermodynamic Characterization of Protein-Ligand Interactions Formed between Lipoprotein-Associated Phospholipase A2 and Inhibitors
J.Med.Chem., 59, 2016
|
|
8W3Q
 
 | |
7MAP
 
 | | Drug Resistant HIV-1 Protease (L10I, V32I, L33F, K45I, M46I, I50V, A71V, V82I, I84V) in Complex with DRV | | Descriptor: | (3R,3AS,6AR)-HEXAHYDROFURO[2,3-B]FURAN-3-YL(1S,2R)-3-[[(4-AMINOPHENYL)SULFONYL](ISOBUTYL)AMINO]-1-BENZYL-2-HYDROXYPROPYLCARBAMATE, Protease | | Authors: | Lockbaum, G.J, Schiffer, C.A. | | Deposit date: | 2021-03-31 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.951 Å) | | Cite: | Drug Resistant HIV-1 Protease (L10I, V32I, L33F, K45I, M46I, I50V, A71V, V82I, I84V) in Complex with DRV
To Be Published
|
|
8VHG
 
 | | Structure of the BMAL1/HIF2A heterodimer in Complex with DNA | | Descriptor: | Basic helix-loop-helix ARNT-like protein 1, Endothelial PAS domain-containing protein 1, Forward strand DNA containing HRE motif, ... | | Authors: | Li, T, Tsai, K.L. | | Deposit date: | 2024-01-01 | | Release date: | 2025-02-12 | | Last modified: | 2025-06-04 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | BMAL1-HIF2A heterodimer modulates circadian variations of myocardial injury.
Nature, 641, 2025
|
|
8VTH
 
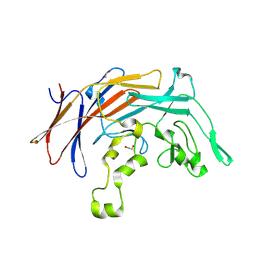 | | DUF2169 domain-containing protein from Vibrio xiamenensis | | Descriptor: | 1,2-ETHANEDIOL, DUF2169 domain-containing protein | | Authors: | Sachar, K, Whitney, J.C, Prehna, G, Kim, Y. | | Deposit date: | 2024-01-26 | | Release date: | 2025-02-12 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | A conserved chaperone protein is required for the formation of a noncanonical type VI secretion system spike tip complex.
J.Biol.Chem., 301, 2025
|
|
5VYP
 
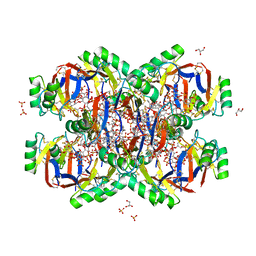 | | Crystal structure of the Plant Defensin NsD7 bound to PIP2 | | Descriptor: | Defensin NsD7, GLYCEROL, SULFATE ION, ... | | Authors: | Jarva, M, Lay, F.T, Hulett, M, Kvansakul, M. | | Deposit date: | 2017-05-25 | | Release date: | 2017-08-02 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of the defensin NsD7 in complex with PIP2 reveals that defensin : lipid oligomer topologies are dependent on lipid type.
FEBS Lett., 591, 2017
|
|
8VWG
 
 | |
6VUE
 
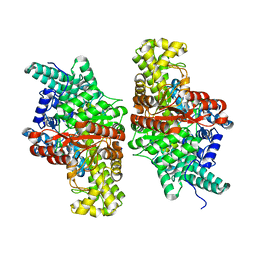 | | wild-type choline TMA lyase in complex with 1-methyl-1,2,3,6-tetrahydropyridin-3-ol | | Descriptor: | (3S)-1-methyl-1,2,3,6-tetrahydropyridin-3-ol, Choline trimethylamine-lyase, SODIUM ION | | Authors: | Ortega, M.A, Drennan, C.L. | | Deposit date: | 2020-02-15 | | Release date: | 2020-11-04 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Discovery of a Cyclic Choline Analog That Inhibits Anaerobic Choline Metabolism by Human Gut Bacteria.
Acs Med.Chem.Lett., 11, 2020
|
|
5IGE
 
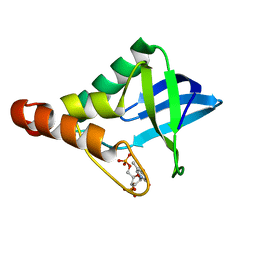 | |
7MBH
 
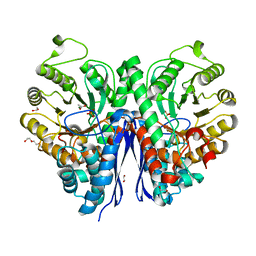 | | Structure of Human Enolase 2 in complex with phosphoserine | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Gamma-enolase, ... | | Authors: | Leonard, P.G, Hicks, K.G, Rutter, J. | | Deposit date: | 2021-03-31 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Protein-metabolite interactomics of carbohydrate metabolism reveal regulation of lactate dehydrogenase.
Science, 379, 2023
|
|
7MAQ
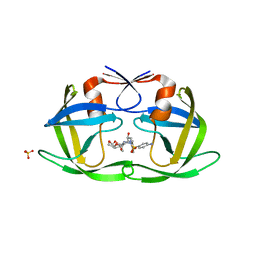 
 | | Drug Resistant HIV-1 Protease (L10F, V32I, L33F, K45I, A71V, V82I, I84V) in Complex with DRV | | Descriptor: | (3R,3AS,6AR)-HEXAHYDROFURO[2,3-B]FURAN-3-YL(1S,2R)-3-[[(4-AMINOPHENYL)SULFONYL](ISOBUTYL)AMINO]-1-BENZYL-2-HYDROXYPROPYLCARBAMATE, Protease, SULFATE ION | | Authors: | Lockbaum, G.J, Schiffer, C.A. | | Deposit date: | 2021-03-31 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Drug Resistant HIV-1 Protease (L10F, V32I, L33F, K45I, A71V, V82I, I84V) in Complex with DRV
To Be Published
|
|
6W8L
 
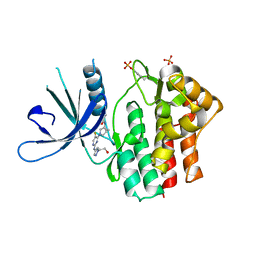 | | Crystal structure of JAK1 kinase with compound 10 | | Descriptor: | N-[(1S,5R)-3-(5-fluoro-2-{[1-(2-hydroxyethyl)-1H-pyrazol-4-yl]amino}pyrimidin-4-yl)-3-azabicyclo[3.1.0]hexan-1-yl]cyclopropanecarboxamide, Tyrosine-protein kinase JAK1 | | Authors: | Vajdos, F.F. | | Deposit date: | 2020-03-20 | | Release date: | 2020-04-08 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Design and optimization of a series of 4-(3-azabicyclo[3.1.0]hexan-3-yl)pyrimidin-2-amines: Dual inhibitors of TYK2 and JAK1.
Bioorg.Med.Chem., 28, 2020
|
|
8VVZ
 
 | |
8VVW
 
 | |
6WBD
 
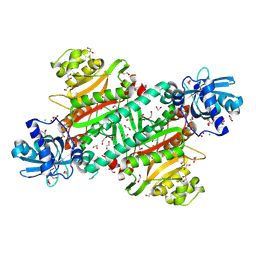 | |
5IGP
 
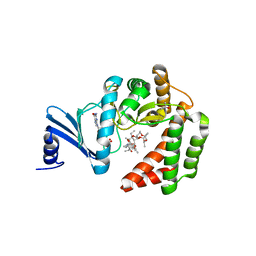 | |
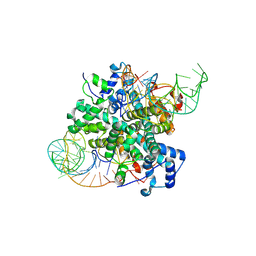
8VZ6
 
 | |
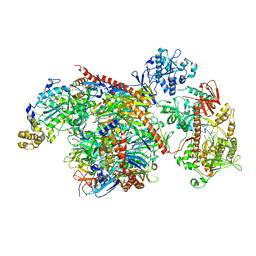
5VZJ
 
 | |
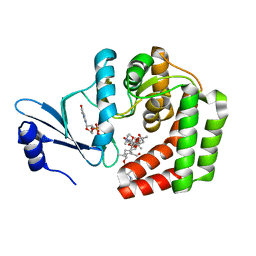
5IGZ
 
 | |
7MA5
 
 | | HIV-1 Protease (I84V) in Complex with UMass4 | | Descriptor: | (3R,3aS,6aR)-hexahydrofuro[2,3-b]furan-3-yl [(1S,2R)-3-{(1,3-benzodioxol-5-ylsulfonyl)[(2S)-2-methylbutyl]amino}-1-benzyl-2-hydroxypropyl]carbamate, Protease, SULFATE ION | | Authors: | Lockbaum, G.J, Schiffer, C.A. | | Deposit date: | 2021-03-31 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.977 Å) | | Cite: | HIV-1 Protease (I84V) in Complex with UMass4
To Be Published
|
|
8VYA
 
 | | SARS-CoV-2 Omicron Variant Spike Glycoprotein Fusion Core (Q954H) | | Descriptor: | SARS-CoV-2 Omicron variant spike glycoprotein C-terminal heptad repeat domain, SARS-CoV-2 Omicron variant spike glycoprotein N-terminal heptad repeat domain (Q954H) | | Authors: | Outlaw, V.K, Apurba, R, Vithanage, N. | | Deposit date: | 2024-02-07 | | Release date: | 2025-02-19 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | SARS-CoV-2 Omicron Variant Spike Glycoprotein Mutation Q954H Enhances Fusion Core Stability.
Acs Chem.Biol., 20, 2025
|
|
5IAV
 
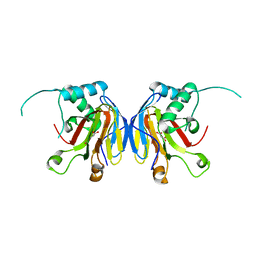 | |
