2JQ1
 | | Phylloseptin-3 | | Descriptor: | Phylloseptin-3 | | Authors: | Resende, J.M, Mendonca Moraes, C, Almeida, F.C.L, Prates, M.V, Cesar, A, Valente, A, Bemquerer, M.P, Pilo-Veloso, D, Bechinger, B. | | Deposit date: | 2007-05-25 | | Release date: | 2008-06-03 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structures of the antimicrobial peptides phylloseptin-1, -2, and -3 and biological activity: The role of charges and hydrogen bonding interactions in stabilizing helix conformations
Peptides, 29, 2008
|
|
2JPY
 
 | | Phylloseptin-2 | | Descriptor: | Phylloseptin-2 protein | | Authors: | Resende, J.M, Mendonca Moraes, C, Almeida, F.C.L, Prates, M.V, Cesar, A, Valente, A, Bemquerer, M.P, Pilo-Veloso, D, Bechinger, B. | | Deposit date: | 2007-05-24 | | Release date: | 2008-06-03 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structures of the antimicrobial peptides phylloseptin-1, -2, and -3 and biological activity: The role of charges and hydrogen bonding interactions in stabilizing helix conformations
Peptides, 29, 2008
|
|
1HHX
 
 | | Solution structure of LNA3:RNA hybrid | | Descriptor: | 5- D(*CP*+TP*GP*AP*+TP*AP*+TP*GP*C) -3, 5- R(*GP*CP*AP*UP*AP*UP*CP*AP*G) -3 | | Authors: | Petersen, M, Bondensgaard, K, Wengel, J, Jacobsen, J.P. | | Deposit date: | 2000-12-29 | | Release date: | 2002-05-30 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Locked Nucleic Acid (Lna) Recognition of RNA: NMR Solution Structures of Lna:RNA Hybrids
J.Am.Chem.Soc., 124, 2002
|
|
1IEZ
 
 | | Solution Structure of 3,4-Dihydroxy-2-Butanone 4-Phosphate Synthase of Riboflavin Biosynthesis | | Descriptor: | 3,4-Dihydroxy-2-Butanone 4-Phosphate Synthase | | Authors: | Kelly, M.J.S, Ball, L.J, Kuhne, R, Bacher, A, Oschkinat, H. | | Deposit date: | 2001-04-11 | | Release date: | 2001-11-07 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The NMR structure of the 47-kDa dimeric enzyme 3,4-dihydroxy-2-butanone-4-phosphate synthase and ligand binding studies reveal the location of the active site.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1HIC
 
 | |
1HHW
 
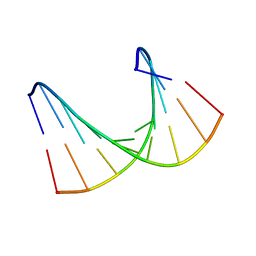 | | Solution structure of LNA1:RNA hybrid | | Descriptor: | 5- D(*CP*TP*GP*AP*+TLNP*AP*TP*GP*C) -3, 5- R(*GP*CP*AP*UP*AP*UP*CP*AP*G) -3 | | Authors: | Petersen, M, Bondensgaard, K, Wengel, J, Jacobsen, J.P. | | Deposit date: | 2000-12-29 | | Release date: | 2002-05-30 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Locked Nucleic Acid (Lna) Recognition of RNA: NMR Solution Structures of Lna:RNA Hybrids
J.Am.Chem.Soc., 124, 2002
|
|
2KIM
 
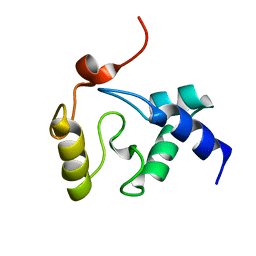 | | 1.7-mm microcryoprobe solution NMR structure of an O6-methylguanine DNA methyltransferase family protein from Vibrio parahaemolyticus. Northeast Structural Genomics Consortium target VpR247. | | Descriptor: | O6-methylguanine-DNA methyltransferase | | Authors: | Aramini, J.M, Belote, R.L, Ciccosanti, C.T, Jiang, M, Rost, B, Nair, R, Swapna, G.V.T, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-05-07 | | Release date: | 2009-07-07 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of an O6-methylguanine DNA methyltransferase family protein from Vibrio parahaemolyticus. Northeast Structural Genomics Consortium target VpR247.
To be Published
|
|
2RRL
 | | Solution structure of the C-terminal domain of the FliK | | Descriptor: | Flagellar hook-length control protein | | Authors: | Mizuno, S, Tate, S, Kobayashi, N, Amida, H. | | Deposit date: | 2011-01-02 | | Release date: | 2011-05-18 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The NMR structure of FliK, the trigger for the switch of substrate specificity in the flagellar type III secretion apparatus
J.Mol.Biol., 409, 2011
|
|
1E7J
 
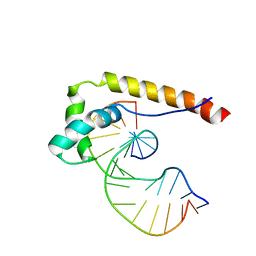 | | HMG-D complexed to a bulge DNA | | Descriptor: | DNA (5'-D(*CP*GP*AP*TP*AP*TP*TP*AP*AP*GP*AP*GP*CP*C)-3'), DNA (5'-D(*GP*GP*CP*TP*CP*AP*AP*TP*AP*TP*CP*G)-3'), HIGH MOBILITY GROUP PROTEIN D | | Authors: | Cerdan, R, Payet, D, Yang, J.-C, Travers, A.A, Neuhaus, D. | | Deposit date: | 2000-08-29 | | Release date: | 2001-03-05 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Hmg-D Complexed to a Bulge DNA: An NMR Model
Protein Sci., 10, 2001
|
|
2WCC
 
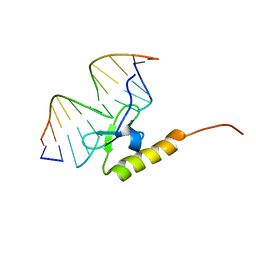 | | phage lambda IntDBD1-64 complex with p prime 2 DNA | | Descriptor: | DNA (5'-D(*DCP*DGP*DAP*DGP*DTP*DCP*DAP *DAP*DAP*DAP*DTP*DC)-3'), DNA (5'-D(*DGP*DAP*DTP*DTP*DTP*DTP*DGP *DAP*DCP*DTP*DGP*DC)-3'), INTEGRASE | | Authors: | Fadeev, E.A, Sam, M.D, Clubb, R.T. | | Deposit date: | 2009-03-11 | | Release date: | 2009-04-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Amino-Terminal Domain of the Lambda Integrase Protein in Complex with DNA: Immobilization of a Flexible Tail Facilitates Beta- Sheet Recognition of the Major Groove.
J.Mol.Biol., 388, 2009
|
|
2WQQ
 
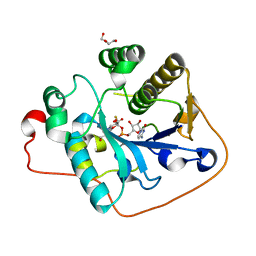 | | Crystallographic analysis of monomeric CstII | | Descriptor: | ALPHA-2,3-/2,8-SIALYLTRANSFERASE, CYTIDINE-5'-MONOPHOSPHATE-3-FLUORO-N-ACETYL-NEURAMINIC ACID, DI(HYDROXYETHYL)ETHER | | Authors: | Chan, P.H.W, Lairson, L.L, Lee, H.J, Wakarchuk, W.W, Strynadka, N.C.J, Withers, S.G, McIntosh, L.P. | | Deposit date: | 2009-08-25 | | Release date: | 2009-10-27 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | NMR Spectroscopic Characterization of the Sialyltransferase Cstii from Camplyobacter Jejuni: Histidine 188 is the General Base.
Biochemistry, 48, 2009
|
|
1UMS
 
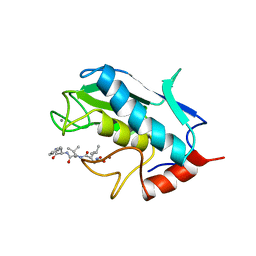 | | STROMELYSIN-1 CATALYTIC DOMAIN WITH HYDROPHOBIC INHIBITOR BOUND, PH 7.0, 32OC, 20 MM CACL2, 15% ACETONITRILE; NMR ENSEMBLE OF 20 STRUCTURES | | Descriptor: | CALCIUM ION, N-{(2R)-2-[2-(hydroxyamino)-2-oxoethyl]-4-methylpentanoyl}-L-leucyl-L-phenylalaninamide, STROMELYSIN-1, ... | | Authors: | Van Doren, S.R, Kurochkin, A.V, Hu, W, Zuiderweg, E.R.P. | | Deposit date: | 1995-10-31 | | Release date: | 1996-03-08 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the catalytic domain of human stromelysin complexed with a hydrophobic inhibitor.
Protein Sci., 4, 1995
|
|
1V6R
 | | Solution Structure of Endothelin-1 with its C-terminal Folding | | Descriptor: | Endothelin-1 | | Authors: | Takashima, H, Mimura, N, Ohkubo, T, Yoshida, T, Tamaoki, H, Kobayashi, Y. | | Deposit date: | 2003-12-03 | | Release date: | 2004-03-16 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Distributed Computing and NMR Constraint-Based High-Resolution Structure
Determination: Applied for Bioactive Peptide Endothelin-1 To Determine C-Terminal
Folding
J.Am.Chem.Soc., 126, 2004
|
|
2KL6
 | | Solution NMR structure of the CARDB domain of PF1109 from Pyrococcus furiosus. Northeast Structural Genomics Consortium target PfR193A | | Descriptor: | Uncharacterized protein | | Authors: | Aramini, J.M, Lee, D, Ciccosanti, C, Hamilton, K, Nair, R, Rost, B, Acton, T.B, Xiao, R, Swapna, G.V.T, Everett, J.K, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-30 | | Release date: | 2009-08-25 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the CARDB domain of PF1109 from Pyrococcus furiosus. Northeast Structural Genomics Consortium target PfR193A
To be Published
|
|
2K5R
 | | Solution NMR Structure of XF2673 from Xylella fastidiosa. Northeast Structural Genomics Consortium Target XfR39 | | Descriptor: | uncharacterized protein XF2673 | | Authors: | Tang, Y, Wang, H, Jiang, M, Maglaqui, M, Xiao, R, Liu, J, Baran, M.C, Swapna, G, Acton, T.B, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-30 | | Release date: | 2008-08-26 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of XF2673 from Xylella fastidiosa. Northeast Structural Genomics
Consortium Target XfR39
To be Published
|
|
2K5V
 
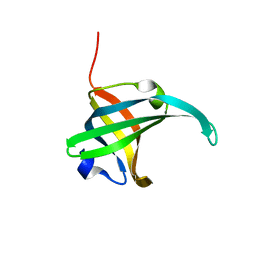 | | SOLUTION NMR STRUCTURE OF the second OB-fold domain of replication protein A from Methanococcus maripaludis. NORTHEAST STRUCTURAL GENOMICS TARGET MrR110B. | | Descriptor: | Replication protein A | | Authors: | Aramini, J.M, Maglaqui, M, Jiang, M, Ciccosanti, C, Xiao, R, Nair, R, Everett, J.K, Swapna, G.VT, Acton, T.B, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-30 | | Release date: | 2008-08-19 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | SOLUTION NMR STRUCTURE OF the second OB-fold domain of replication protein A from Methanococcus maripaludis. NORTHEAST STRUCTURAL GENOMICS TARGET MrR110B.
To be Published
|
|
2KKZ
 | | Solution NMR structure of the monomeric W187R mutant of A/Udorn NS1 effector domain. Northeast Structural Genomics target OR8C[W187R]. | | Descriptor: | Non-structural protein NS1 | | Authors: | Aramini, J.M, Ma, L, Lee, H, Zhao, L, Cunningham, K, Ciccosanti, C, Janjua, H, Fang, Y, Xiao, R, Krug, R.M, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-29 | | Release date: | 2009-07-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the monomeric W187R mutant of A/Udorn NS1 effector domain. Northeast Structural Genomics target OR8C[W187R].
To be Published
|
|
2KPN
 | | Solution NMR structure of a Bacterial Ig-like (Big_3) domain from Bacillus cereus. Northeast Structural Genomics Consortium target BcR147A | | Descriptor: | Bacillolysin, CALCIUM ION | | Authors: | Aramini, J.M, Wang, D, Ciccosanti, C.T, Janjua, H, Rost, B, Acton, T.B, Xiao, R, Swapna, G.V.T, Everett, J.K, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-10-16 | | Release date: | 2010-01-12 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of a Bacterial Ig-like (Big_3) domain from Bacillus cereus. Northeast Structural Genomics Consortium target BcR147A
To be Published
|
|
1HDN
 | |
1JT8
 
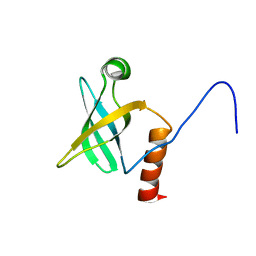 | | ARCHAEAL INITIATION FACTOR-1A, AIF-1A | | Descriptor: | PROBABLE TRANSLATION INITIATION FACTOR 1A | | Authors: | Hoffman, D.W, Li, W. | | Deposit date: | 2001-08-20 | | Release date: | 2001-09-05 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure and dynamics of translation initiation factor aIF-1A from the archaeon Methanococcus jannaschii determined by NMR spectroscopy
Protein Sci., 10, 2001
|
|
1MSG
 
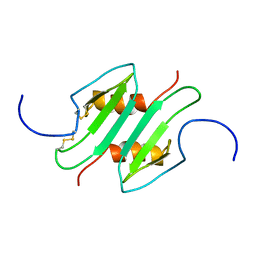 | |
1KOD
 
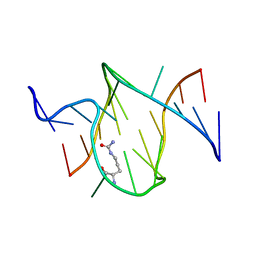 | | RNA APTAMER COMPLEXED WITH CITRULLINE, NMR | | Descriptor: | CITRULLINE, RNA (5'-R(P*AP*CP*GP*GP*UP*UP*AP*GP*GP*UP*CP*GP*CP*U)-3'), RNA (5'-R(P*AP*GP*AP*AP*GP*GP*AP*GP*UP*GP*U)-3') | | Authors: | Yang, Y.S, Kochoyan, M, Burgstaller, P, Westhof, E, Famulok, M. | | Deposit date: | 1996-03-28 | | Release date: | 1996-11-08 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis of ligand discrimination by two related RNA aptamers resolved by NMR spectroscopy.
Science, 272, 1996
|
|
1NEA
 | | THREE-DIMENSIONAL SOLUTION STRUCTURE OF A CURAREMIMETIC TOXIN FROM NAJA NIGRICOLLIS VENOM: A PROTON NMR AND MOLECULAR MODELING STUDY | | Descriptor: | TOXIN ALPHA | | Authors: | Zinn-Justin, S, Roumestand, C, Gilquin, B, Bontems, F, Menez, A, Toma, F. | | Deposit date: | 1992-09-22 | | Release date: | 1993-10-31 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional solution structure of a curaremimetic toxin from Naja nigricollis venom: a proton NMR and molecular modeling study.
Biochemistry, 31, 1992
|
|
1KOC
 
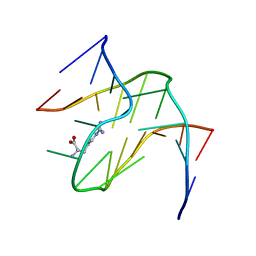 | | RNA APTAMER COMPLEXED WITH ARGININE, NMR | | Descriptor: | ARGININE, RNA (5'-R(P*AP*CP*AP*GP*GP*UP*AP*GP*GP*UP*CP*GP*CP*U)-3'), RNA (5'-R(P*AP*GP*AP*AP*GP*GP*AP*GP*CP*GP*U)-3') | | Authors: | Yang, Y.S, Kochoyan, M, Burgstaller, P, Westhof, E, Famulok, M. | | Deposit date: | 1996-03-28 | | Release date: | 1996-08-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis of ligand discrimination by two related RNA aptamers resolved by NMR spectroscopy.
Science, 272, 1996
|
|
1MSH
 | |
