5WI4
 
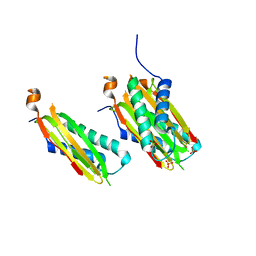 | | CRYSTAL STRUCTURE OF DYNLT1/TCTEX-1 IN COMPLEX WITH ARHGEF2 | | Descriptor: | Dynein light chain Tctex-type 1,Rho guanine nucleotide exchange factor 2, SULFATE ION | | Authors: | Balan, M, Ishiyama, N, Marshall, C.B, Ikura, M. | | Deposit date: | 2017-07-18 | | Release date: | 2017-11-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | MARK3-mediated phosphorylation of ARHGEF2 couples microtubules to the actin cytoskeleton to establish cell polarity.
Sci Signal, 10, 2017
|
|
1X3M
 
 | |
1X3N
 
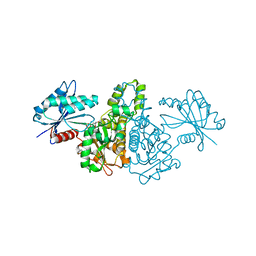
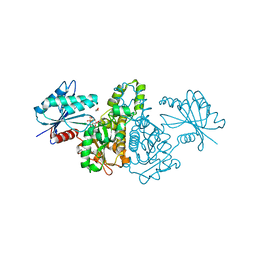 | | Crystal structure of AMPPNP bound Propionate kinase (TdcD) from Salmonella typhimurium | | Descriptor: | 1,2-ETHANEDIOL, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, Propionate kinase | | Authors: | Simanshu, D.K, Savithri, H.S, Murthy, M.R. | | Deposit date: | 2005-05-10 | | Release date: | 2005-09-27 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structures of ADP and AMPPNP-bound Propionate Kinase (TdcD) from Salmonella typhimurium: Comparison with Members of Acetate and Sugar Kinase/Heat Shock Cognate 70/Actin Superfamily.
J.Mol.Biol., 352, 2005
|
|
7E1G
 
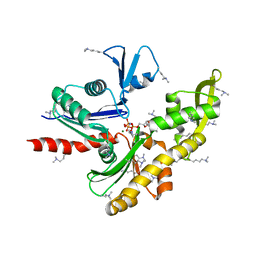 | | Structure of MreB3 from Spiroplasma eriocheiris | | Descriptor: | Cell shape-determining protein MreB, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Takahashi, D, Miyata, M, Imada, K. | | Deposit date: | 2021-02-01 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | ATP-dependent polymerization dynamics of bacterial actin proteins involved in Spiroplasma swimming.
Open Biology, 12, 2022
|
|
7E1C
 
 | | Structure of MreB3 from Spiroplasma eriocheiris | | Descriptor: | ACETATE ION, CALCIUM ION, Cell shape-determining protein MreB | | Authors: | Takahashi, D, Miyata, M, Imada, K. | | Deposit date: | 2021-02-01 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | ATP-dependent polymerization dynamics of bacterial actin proteins involved in Spiroplasma swimming.
Open Biology, 12, 2022
|
|
5G4X
 
 | | The crystal structure of the SHANK3 N-terminus | | Descriptor: | 1,2-ETHANEDIOL, SH3 AND MULTIPLE ANKYRIN REPEAT DOMAINS PROTEIN 3 | | Authors: | Zacharchenko, T, Barsukov, I. | | Deposit date: | 2016-05-17 | | Release date: | 2017-01-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.166 Å) | | Cite: | SHANK proteins limit integrin activation by directly interacting with Rap1 and R-Ras.
Nat. Cell Biol., 19, 2017
|
|
2L4S
 
 | | Promiscuous Binding at the Crossroads of Numerous Cancer Pathways: Insight from the Binding of GIP with Glutaminase L | | Descriptor: | Tax1-binding protein 3 | | Authors: | Zoetewey, D.L, Ovee, M, Banerjee, M, Bhaskaran, R, Mohanty, S. | | Deposit date: | 2010-10-13 | | Release date: | 2011-04-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Promiscuous binding at the crossroads of numerous cancer pathways: insight from the binding of glutaminase interacting protein with glutaminase L.
Biochemistry, 50, 2011
|
|
2L4T
 
 | | GIP/Glutaminase L peptide complex | | Descriptor: | Glutaminase L peptide, Tax1-binding protein 3 | | Authors: | Zoetewey, D.L, Ovee, M, Banerjee, M, Bhaskaran, R, Mohanty, S. | | Deposit date: | 2010-10-13 | | Release date: | 2011-04-06 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Promiscuous binding at the crossroads of numerous cancer pathways: insight from the binding of glutaminase interacting protein with glutaminase L.
Biochemistry, 50, 2011
|
|
2KXC
 
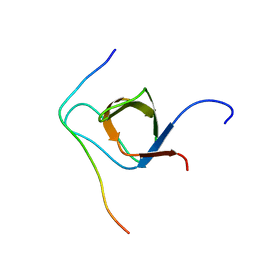 | | 1H, 13C, and 15N Chemical Shift Assignments for IRTKS-SH3 and EspFu-R47 complex | | Descriptor: | Brain-specific angiogenesis inhibitor 1-associated protein 2-like protein 1, EspF-like protein | | Authors: | Aitio, O, Hellman, M, Kazlauskas, A, Vingadassalom, D.F, Leong, J.M, Saksela, K, Permi, P. | | Deposit date: | 2010-04-30 | | Release date: | 2010-11-17 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Recognition of tandem PxxP motifs as a unique Src homology 3-binding mode triggers pathogen-driven actin assembly
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2DAT
 
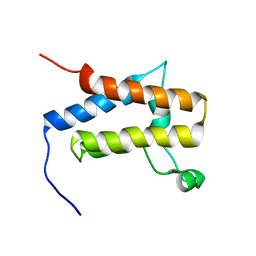 | | Solution structure of the Bromodomain of human SWI/SNF related matrix associated actin dependent regulator of cromatin subfamily A member 2 | | Descriptor: | Possible global transcription activator SNF2L2 | | Authors: | Yoneyama, M, Kigawa, T, Saito, K, Tochio, N, Koshiba, S, Inoue, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-12-14 | | Release date: | 2007-01-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the Bromodomain of human SWI/SNF related matrix associated actin dependent regulator of cromatin subfamily A member 2
To be Published
|
|
7O7J
 
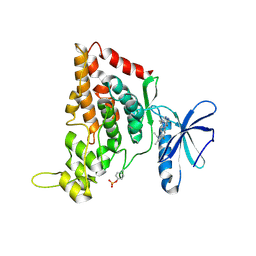 | | Crystal structure of the human HIPK3 kinase domain bound to abemaciclib | | Descriptor: | Homeodomain-interacting protein kinase 3, N-{5-[(4-ethylpiperazin-1-yl)methyl]pyridin-2-yl}-5-fluoro-4-[4-fluoro-2-methyl-1-(propan-2-yl)-1H-benzimidazol-6-yl]py rimidin-2-amine | | Authors: | Kaltheuner, I.H, Anand, K, Geyer, M. | | Deposit date: | 2021-04-13 | | Release date: | 2021-11-24 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Abemaciclib is a potent inhibitor of DYRK1A and HIP kinases involved in transcriptional regulation.
Nat Commun, 12, 2021
|
|
1X99
 
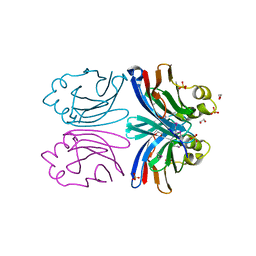 | | X-ray crystal structure of Xerocomus chrysenteron lectin XCL at 1.4 Angstroms resolution, mutated at Q46M, V54M, L58M | | Descriptor: | 1,2-ETHANEDIOL, SULFATE ION, lectin | | Authors: | Birck, C, Damian, L, Marty-Detraves, C, Lougarre, A, Schulze-Briese, C, Koehl, P, Fournier, D, Paquereau, L, Samama, J.P. | | Deposit date: | 2004-08-20 | | Release date: | 2004-12-14 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | A New Lectin Family with Structure Similarity to Actinoporins Revealed by the Crystal Structure of Xerocomus chrysenteron Lectin XCL
J.Mol.Biol., 344, 2004
|
|
1XI0
 
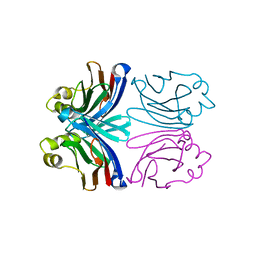 | | X-ray crystal structure of wild-type Xerocomus chrysenteron lectin XCL | | Descriptor: | lectin | | Authors: | Birck, C, Damian, L, Marty-Detraves, C, Lougarre, A, Schulze-Briese, C, Koehl, P, Fournier, D, Paquereau, L, Samama, J.P. | | Deposit date: | 2004-09-21 | | Release date: | 2005-08-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A new lectin family with structure similarity to actinoporins revealed by the crystal structure of Xerocomus chrysenteron lectin XCL
J.Mol.Biol., 344, 2004
|
|
7P47
 
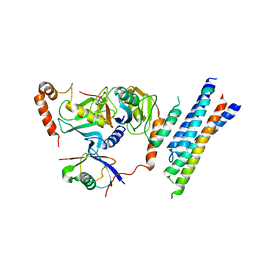 | | Structure of the E3 ligase Smc5/Nse2 in complex with Ubc9-SUMO thioester mimetic | | Descriptor: | E3 SUMO-protein ligase MMS21, SUMO-conjugating enzyme UBC9, Structural maintenance of chromosomes protein 5, ... | | Authors: | Lascorz, J, Varejao, N, Reverter, D. | | Deposit date: | 2021-07-09 | | Release date: | 2021-11-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.314 Å) | | Cite: | Structural basis for the E3 ligase activity enhancement of yeast Nse2 by SUMO-interacting motifs.
Nat Commun, 12, 2021
|
|
5W9A
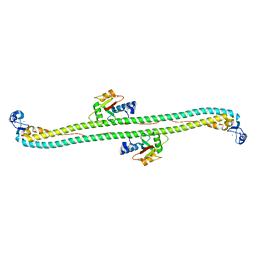 
 | | The structure of the Trim5alpha Bbox- coiled coil in complex LC3B | | Descriptor: | Microtubule-associated proteins 1A/1B light chain 3B, Tripartite motif-containing protein 5, ZINC ION | | Authors: | Keown, J.R, Goldstone, D.C. | | Deposit date: | 2017-06-22 | | Release date: | 2018-10-10 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.74 Å) | | Cite: | A helical LC3-interacting region mediates the interaction between the retroviral restriction factor Trim5 alpha and mammalian autophagy-related ATG8 proteins.
J. Biol. Chem., 293, 2018
|
|
7LOI
 
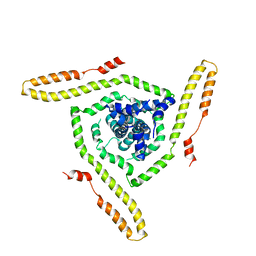 | | Model of the HIV-1 gp41 membrane-proximal external region, transmembrane domain and cytoplasmic tail | | Descriptor: | Transmembrane protein gp41 | | Authors: | Piai, A, Fu, Q, Sharp, A.K, Bighi, B, Brown, A.M, Chou, J.J. | | Deposit date: | 2021-02-10 | | Release date: | 2021-04-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR Model of the Entire Membrane-Interacting Region of the HIV-1 Fusion Protein and Its Perturbation of Membrane Morphology.
J.Am.Chem.Soc., 143, 2021
|
|
2FMU
 
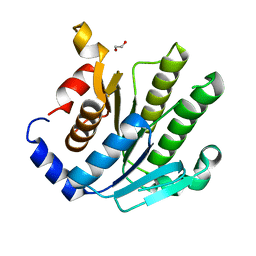 | |
5DZ6
 
 | | Acyl transferase from Bacillaene PKS | | Descriptor: | 1,2-ETHANEDIOL, BROMIDE ION, CHLORIDE ION, ... | | Authors: | Till, M, Race, P.R. | | Deposit date: | 2015-09-25 | | Release date: | 2016-10-05 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | Architectural hierarchy of trans-acting enoyl reductases from polyunsaturated fatty acid and trans-acyltransferase polyketide synthases
To Be Published
|
|
4UVQ
 
 | | PatG Domain of Unknown Function | | Descriptor: | THIAZOLINE OXIDASE/SUBTILISIN-LIKE PROTEASE, ZINC ION | | Authors: | Mann, G, Koehnke, J, Bent, A.F, Graham, R, Schwarz-Linek, U, Naismith, J.H. | | Deposit date: | 2014-08-07 | | Release date: | 2014-09-17 | | Last modified: | 2014-12-17 | | Method: | X-RAY DIFFRACTION (1.724 Å) | | Cite: | The Structure of the Cyanobactin Domain of Unknown Function from Patg in the Patellamide Gene Cluster
Acta Crystallogr.,Sect.F, 70, 2014
|
|
4EIH
 
 | | Crystal structure of Arg SH2 domain | | Descriptor: | Abelson tyrosine-protein kinase 2, CHLORIDE ION | | Authors: | Liu, W, MacGrath, S.M, Koleske, A.J, Boggon, T.J. | | Deposit date: | 2012-04-05 | | Release date: | 2013-04-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Two Amino Acid Residues Confer Different Binding Affinities of Abelson Family Kinase Src Homology 2 Domains for Phosphorylated Cortactin.
J.Biol.Chem., 289, 2014
|
|
5W2E
 
 | | HCV NS5B RNA-dependent RNA polymerase in complex with non-nucleoside inhibitor MK-8876 | | Descriptor: | 2-(4-fluorophenyl)-5-(11-fluoro-6H-pyrido[2',3':5,6][1,3]oxazino[3,4-a]indol-2-yl)-N-methyl-6-[methyl(methylsulfonyl)amino]-1-benzofuran-3-carboxamide, Genome polyprotein | | Authors: | Lesburg, C.A, Ummat, A. | | Deposit date: | 2017-06-06 | | Release date: | 2017-08-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Development of a New Structural Class of Broadly Acting HCV Non-Nucleoside Inhibitors Leading to the Discovery of MK-8876.
ChemMedChem, 12, 2017
|
|
5VT2
 
 | | Crystal structure of growth differentiation factor | | Descriptor: | 1,2-ETHANEDIOL, Growth/differentiation factor 15 | | Authors: | Min, X, Wang, Z. | | Deposit date: | 2017-05-15 | | Release date: | 2017-11-01 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Long-acting MIC-1/GDF15 molecules to treat obesity: Evidence from mice to monkeys.
Sci Transl Med, 9, 2017
|
|
6SVL
 
 | |
6SVK
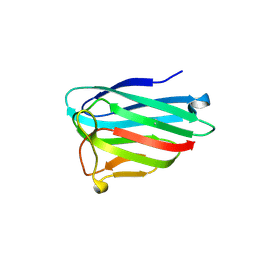 
 | | human Myeloid-derived growth factor (MYDGF) | | Descriptor: | MALONATE ION, Myeloid-derived growth factor | | Authors: | Ebenhoch, R, Nar, H. | | Deposit date: | 2019-09-18 | | Release date: | 2019-11-27 | | Last modified: | 2019-12-04 | | Method: | X-RAY DIFFRACTION (1.604 Å) | | Cite: | Crystal structure and receptor-interacting residues of MYDGF - a protein mediating ischemic tissue repair.
Nat Commun, 10, 2019
|
|
5D67
 
 | |
