4W4U
 
 | |
4W55
 
 | | T4 Lysozyme L99A with n-Propylbenzene Bound | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Endolysin, propylbenzene | | Authors: | Merski, M, Shoichet, B.K, Eidam, O, Fischer, M. | | Deposit date: | 2014-08-16 | | Release date: | 2015-04-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.6401 Å) | | Cite: | Homologous ligands accommodated by discrete conformations of a buried cavity.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4W58
 
 | | T4 Lysozyme L99A with n-Pentylbenzene Bound | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Endolysin, pentylbenzene | | Authors: | Merski, M, Shoichet, B.K, Eidam, O, Fischer, M. | | Deposit date: | 2014-08-16 | | Release date: | 2015-04-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Homologous ligands accommodated by discrete conformations of a buried cavity.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4UU3
 
 | | Ferulic acid decarboxylase from Enterobacter sp. | | Descriptor: | FERULIC ACID DECARBOXYLASE | | Authors: | Hromic, A, Pavkov-Keller, T, Steinkellner, G, Lyskowski, A, Wuensch, C, Gross, J, Fuchs, M, Fauland, K, Glueck, S.M, Faber, K, Gruber, K. | | Deposit date: | 2014-07-24 | | Release date: | 2015-06-10 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Regioselective Enzymatic Beta-Carboxylation of Para-Hydroxy-Styrene Derivatives Catalyzed by Phenolic Acid Decarboxylases.
Adv. Synth. Catal., 357, 2015
|
|
4UQO
 
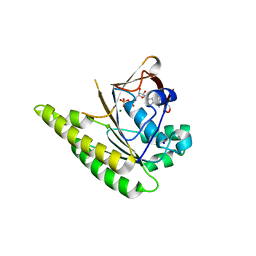 | | RADA C-TERMINAL ATPASE DOMAIN FROM PYROCOCCUS FURIOSUS BOUND TO ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA REPAIR AND RECOMBINATION PROTEIN RADA, MAGNESIUM ION, ... | | Authors: | Marsh, M.E, Ehebauer, M.T, Scott, D, Abell, C, Blundell, T.L, Hyvonen, M. | | Deposit date: | 2014-06-24 | | Release date: | 2015-01-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | ATP Half-Sites in Rada and Rad51 Recombinases Bind Nucleotides
FEBS Open Bio, 6, 2016
|
|
4V3F
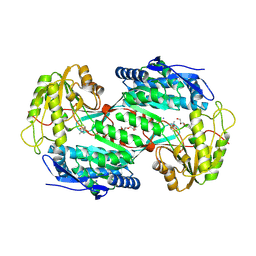 
 | |
4UUM
 
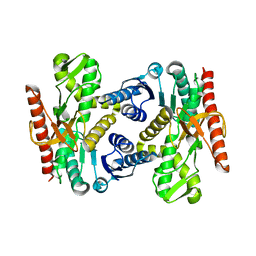 | |
2MKG
 
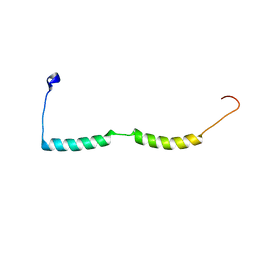 | |
2MKF
 
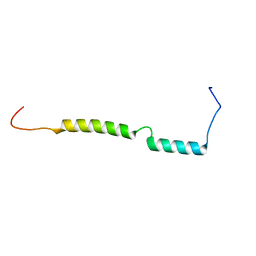 | |
2M9M
 
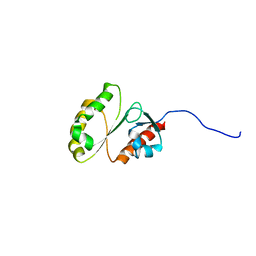 | | Solution Structure of ERCC4 domain of human FAAP24 | | Descriptor: | Fanconi anemia-associated protein of 24 kDa | | Authors: | Wu, F, Han, X, Shi, C, Gong, W, Tian, C. | | Deposit date: | 2013-06-18 | | Release date: | 2013-09-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure analysis of FAAP24 reveals single-stranded DNA-binding activity and domain functions in DNA damage response.
Cell Res., 23, 2013
|
|
2MBG
 
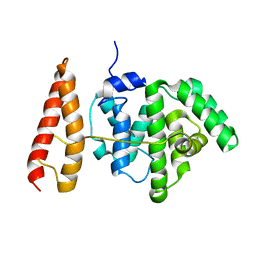 | | Rlip76 (gap-gbd) | | Descriptor: | RalA-binding protein 1 | | Authors: | Rajasekar, K.V, Campbell, L.J, Nietlispach, D, Owen, D, Mott, H.R. | | Deposit date: | 2013-07-30 | | Release date: | 2013-12-04 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Structure of the RLIP76 RhoGAP-Ral Binding Domain Dyad: Fixed Position of the Domains Leads to Dual Engagement of Small G Proteins at the Membrane.
Structure, 21, 2013
|
|
2M9N
 
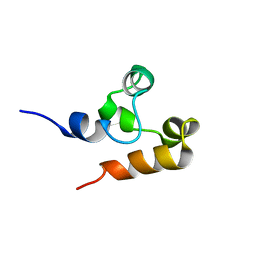 | | Solution Structure of (HhH)2 domain of human FAAP24 | | Descriptor: | Fanconi anemia-associated protein of 24 kDa | | Authors: | Wu, F, Han, X, Shi, C, Gong, W, Tian, C. | | Deposit date: | 2013-06-18 | | Release date: | 2013-09-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure analysis of FAAP24 reveals single-stranded DNA-binding activity and domain functions in DNA damage response.
Cell Res., 23, 2013
|
|
2MUU
 
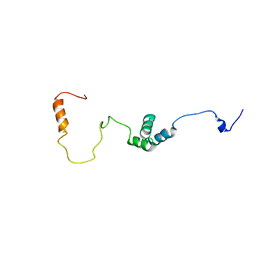 | |
7BGF
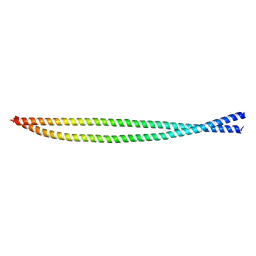 
 | |
7D2G
 
 | | Coiled-coil structure of liprin-alpha2_H2delC | | Descriptor: | GLYCEROL, Liprin-alpha-2 | | Authors: | Liang, M, Wei, Z. | | Deposit date: | 2020-09-16 | | Release date: | 2021-04-07 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Oligomerized liprin-alpha promotes phase separation of ELKS for compartmentalization of presynaptic active zone proteins.
Cell Rep, 34, 2021
|
|
7D2E
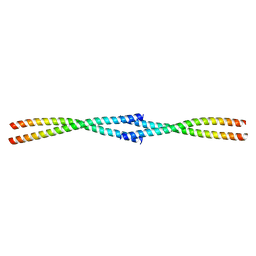 
 | | Tetrameric coiled-coil structure of liprin-alpha2_H3 | | Descriptor: | Liprin-alpha-2 | | Authors: | Liang, M, Wei, Z. | | Deposit date: | 2020-09-16 | | Release date: | 2021-04-07 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Oligomerized liprin-alpha promotes phase separation of ELKS for compartmentalization of presynaptic active zone proteins.
Cell Rep, 34, 2021
|
|
7D2H
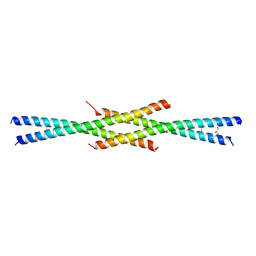 
 | | Tetrameric coiled-coil structure of liprin-alpha2_H2 | | Descriptor: | GLYCEROL, Liprin-alpha-2 | | Authors: | Liang, M, Wei, Z. | | Deposit date: | 2020-09-16 | | Release date: | 2021-04-07 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Oligomerized liprin-alpha promotes phase separation of ELKS for compartmentalization of presynaptic active zone proteins.
Cell Rep, 34, 2021
|
|
2OLM
 
 | | ArfGap domain of HIV-1 Rev binding protein | | Descriptor: | GLYCEROL, Nucleoporin-like protein RIP, SULFATE ION, ... | | Authors: | Tong, Y, Tempel, W, Shen, L, Dimov, S, Landry, R, Arrowsmith, C.H, Edwards, A.M, Sundstrom, M, Weigelt, J, Bochkarev, A, Park, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-01-19 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | ArfGap domain of HIV-1 Rev binding protein
To be Published
|
|
7EVR
 
 | | Crystal structure of hnRNP L RRM2 in complex with SETD2 | | Descriptor: | Heterogeneous nuclear ribonucleoprotein L, SHI domain from Histone-lysine N-methyltransferase SETD2 | | Authors: | Li, F.D, Wang, S.M. | | Deposit date: | 2021-05-22 | | Release date: | 2021-10-20 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis of the interaction between SETD2 methyltransferase and hnRNP L paralogs for governing co-transcriptional splicing.
Nat Commun, 12, 2021
|
|
7F0U
 
 | |
7EVS
 
 | | Crystal structure of hnRNP LL RRM2 in complex with SETD2 | | Descriptor: | Heterogeneous nuclear ribonucleoprotein L-like, SHI domain from Histone-lysine N-methyltransferase SETD2, SULFATE ION | | Authors: | Li, F.D, Wang, S.M. | | Deposit date: | 2021-05-22 | | Release date: | 2021-10-20 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural basis of the interaction between SETD2 methyltransferase and hnRNP L paralogs for governing co-transcriptional splicing.
Nat Commun, 12, 2021
|
|
7D42
 
 | |
2O4J
 
 | | Crystal Structure of Rat Vitamin D Receptor Ligand Binding Domain Complexed with VitIII 17-20Z and the NR2 Box of DRIP 205 | | Descriptor: | (1R,3R,7E,17Z)-17-(5-hydroxy-1,5-dimethylhexylidene)-2-methylene-9,10-secoestra-5,7-diene-1,3-diol, Peroxisome proliferator-activated receptor-binding protein, Vitamin D3 receptor | | Authors: | Vanhooke, J.L, Benning, M.M, DeLuca, H.F. | | Deposit date: | 2006-12-04 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | New analogs of 2-methylene-19-nor-(20S)-1,25-dihydroxyvitamin D(3) with conformationally restricted side chains: Evaluation of biological activity and structural determination of VDR-bound conformations.
Arch.Biochem.Biophys., 460, 2007
|
|
2O4R
 
 | | Crystal Structure of Rat Vitamin D Receptor Ligand Binding Domain Complexed with VitIII 17-20E and the NR2 Box of DRIP 205 | | Descriptor: | (1R,3R,7E,17E)-17-(5-hydroxy-1,5-dimethylhexylidene)-2-methylene-9,10-secoestra-5,7-diene-1,3-diol, Peroxisome proliferator-activated receptor-binding protein, Vitamin D3 receptor | | Authors: | Vanhooke, J.L, Benning, M.M, DeLuca, H.F. | | Deposit date: | 2006-12-04 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | New analogs of 2-methylene-19-nor-(20S)-1,25-dihydroxyvitamin D(3) with conformationally restricted side chains: Evaluation of biological activity and structural determination of VDR-bound conformations.
Arch.Biochem.Biophys., 460, 2007
|
|
6CPJ
 
 | | Solution structure of SH3 domain from Shank2 | | Descriptor: | SH3 and multiple ankyrin repeat domains protein 2 | | Authors: | Ishida, H, Vogel, H.J. | | Deposit date: | 2018-03-13 | | Release date: | 2018-08-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the SH3 domains from Shank scaffold proteins and their interactions with Cav1.3 calcium channels.
FEBS Lett., 592, 2018
|
|
