2KRG
 
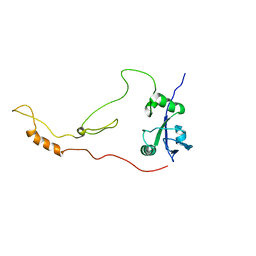 | | Solution Structure of human sodium/ hydrogen exchange regulatory factor 1(150-358) | | Descriptor: | Na(+)/H(+) exchange regulatory cofactor NHE-RF1 | | Authors: | Bhattacharya, S, Dai, Z, Li, J, Baxter, S, Callaway, D.J.E, Cowburn, D, Bu, Z. | | Deposit date: | 2009-12-17 | | Release date: | 2009-12-29 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A conformational switch in the scaffolding protein NHERF1 controls autoinhibition and complex formation.
J.Biol.Chem., 285, 2010
|
|
2M0V
 
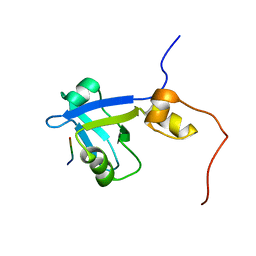 | | Complex structure of C-terminal CFTR peptide and extended PDZ2 domain from NHERF1 | | Descriptor: | C-terminal CFTR peptide, Na(+)/H(+) exchange regulatory cofactor NHE-RF1 | | Authors: | Bhattacharya, S, Ju, J.H, Cowburn, D, Bu, Z. | | Deposit date: | 2012-11-06 | | Release date: | 2013-04-24 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Ligand-Induced Dynamic Changes in Extended PDZ Domains from NHERF1.
J.Mol.Biol., 425, 2013
|
|
2M0T
 
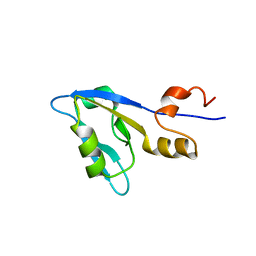 | |
2NXP
 
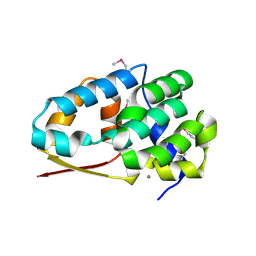 | |
1X6G
 
 | | Solution structures of the SH3 domain of human megakaryocyte-associated tyrosine-protein kinase. | | Descriptor: | Megakaryocyte-associated tyrosine-protein kinase | | Authors: | Sato, M, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-17 | | Release date: | 2005-11-17 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the SH3 domain of human megakaryocyte-associated tyrosine-protein kinase.
To be Published
|
|
1D15
 
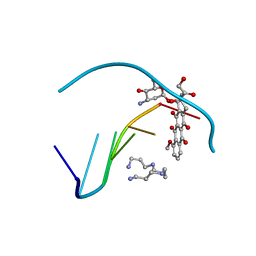 | | TERNARY INTERACTIONS OF SPERMINE WITH DNA: 4'-EPIADRIAMYCIN AND OTHER DNA: ANTHRACYCLINE COMPLEXES | | Descriptor: | 4'-EPIDOXORUBICIN, DNA (5'-D(*CP*GP*AP*TP*CP*G)-3'), SPERMINE | | Authors: | Williams, L.D, Frederick, C.A, Ughetto, G, Rich, A. | | Deposit date: | 1990-07-03 | | Release date: | 1991-07-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Ternary interactions of spermine with DNA: 4'-epiadriamycin and other DNA: anthracycline complexes.
Nucleic Acids Res., 18, 1990
|
|
1D6N
 
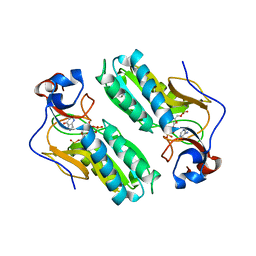 | | TERNARY COMPLEX STRUCTURE OF HUMAN HGPRTASE, PRPP, MG2+, AND THE INHIBITOR HPP REVEALS THE INVOLVEMENT OF THE FLEXIBLE LOOP IN SUBSTRATE BINDING | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, 3H-PYRAZOLO[4,3-D]PYRIMIDIN-7-OL, MAGNESIUM ION, ... | | Authors: | Balendiran, G.K. | | Deposit date: | 1999-10-14 | | Release date: | 1999-12-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Ternary complex structure of human HGPRTase, PRPP, Mg2+, and the inhibitor HPP reveals the involvement of the flexible loop in substrate binding.
Protein Sci., 8, 1999
|
|
2JOD
 
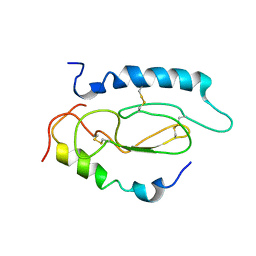 | | Pac1-Rshort N-terminal EC domain Pacap(6-38) complex | | Descriptor: | Pituitary adenylate cyclase-activating polypeptide, Pituitary adenylate cyclase-activating polypeptide type I receptor | | Authors: | Olejniczak, E.T, Sun, C, Song, D, Davis-Taber, R.A, Barrett, L.W, Scott, V.E, Richardson, P.L, Pereda-lopez, A, Uchic, M.E, Solomon, L.R, Lake, M.R, Walter, K.A, Hajduk, P.J. | | Deposit date: | 2007-03-07 | | Release date: | 2007-05-22 | | Last modified: | 2024-11-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure and mutational analysis of pituitary adenylate cyclase-activating polypeptide binding to the extracellular domain of PAC1-RS.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2F9G
 
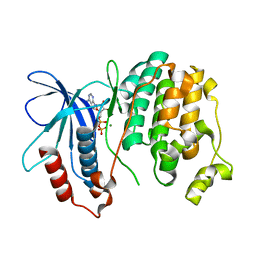 | | Crystal structure of Fus3 phosphorylated on Tyr182 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Mitogen-activated protein kinase FUS3 | | Authors: | Bhattacharyya, R.P, Remenyi, A, Good, M.C, Bashor, C.J, Falick, A.M, Lim, W.A. | | Deposit date: | 2005-12-05 | | Release date: | 2006-11-14 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Ste5 scaffold allosterically modulates signaling output of the yeast mating pathway.
Science, 311, 2006
|
|
2FA2
 
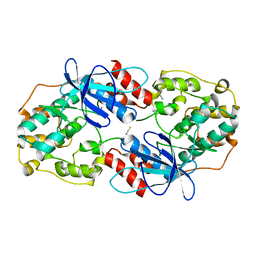 | | Crystal structure of Fus3 without a peptide from Ste5 | | Descriptor: | Mitogen-activated protein kinase FUS3, THIOCYANATE ION | | Authors: | Bhattacharyya, R.P, Remenyi, A, Good, M.C, Bashor, C.J, Falick, A.M, Lim, W.A. | | Deposit date: | 2005-12-06 | | Release date: | 2006-03-28 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | The Ste5 scaffold allosterically modulates signaling output of the yeast mating pathway
Science, 311, 2006
|
|
1I11
 
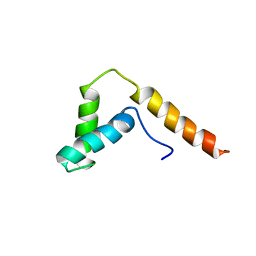 | | SOLUTION STRUCTURE OF THE DNA BINDING DOMAIN, SOX-5 HMG BOX FROM MOUSE | | Descriptor: | TRANSCRIPTION FACTOR SOX-5 | | Authors: | Cary, P.D, Read, C.M, Davis, B, Driscoll, P.C, Crane-Robinson, C. | | Deposit date: | 2001-01-30 | | Release date: | 2001-02-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure and backbone dynamics of the DNA-binding domain of mouse Sox-5.
Protein Sci., 10, 2001
|
|
2MLF
 
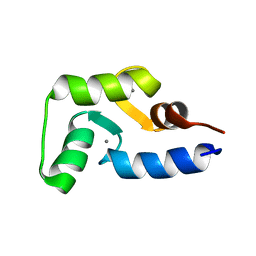 | | NMR structure of the dilated cardiomyopathy mutation G159D in troponin C bound to the anchoring region of troponin I | | Descriptor: | CALCIUM ION, Troponin C, slow skeletal and cardiac muscles | | Authors: | Baryshnikova, O.K, Robertson, I.M, Mercier, P, Sykes, B.D. | | Deposit date: | 2014-02-26 | | Release date: | 2014-03-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The dilated cardiomyopathy G159D mutation in cardiac troponin C weakens the anchoring interaction with troponin I.
Biochemistry, 47, 2008
|
|
1JKU
 
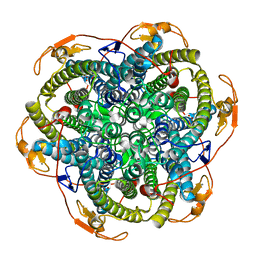 | | Crystal Structure of Manganese Catalase from Lactobacillus plantarum | | Descriptor: | CALCIUM ION, HYDROXIDE ION, MANGANESE (III) ION, ... | | Authors: | Barynin, V.V, Whittaker, M.M, Antonyuk, S.V, Lamzin, V.S, Harrison, P.M, Artymiuk, P.J, Whittaker, J.W. | | Deposit date: | 2001-07-13 | | Release date: | 2002-07-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal structure of manganese catalase from Lactobacillus plantarum.
Structure, 9, 2001
|
|
2K0U
 
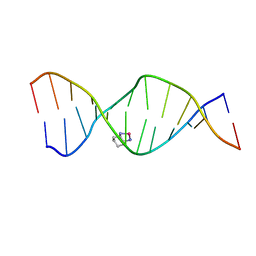 | | High Resolution Solution NMR Structures of Oxaliplatin-DNA Adduct | | Descriptor: | CYCLOHEXANE-1(R),2(R)-DIAMINE-PLATINUM(II), DNA (5'-D(*DCP*DCP*DTP*DCP*DTP*DGP*DGP*DTP*DCP*DTP*DCP*DC)-3'), DNA (5'-D(*DGP*DGP*DAP*DGP*DAP*DCP*DCP*DAP*DGP*DAP*DGP*DG)-3') | | Authors: | Bhattacharyya, D, King, C.L, Chaney, S.G, Campbell, S.L. | | Deposit date: | 2008-02-15 | | Release date: | 2009-02-03 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Flanking Bases Influence the Nature of DNA Distortion by Platinum 1,2-Intrastrand (GG) Cross-Links.
Plos One, 6, 2011
|
|
2K0V
 
 | | High Resolution Solution NMR Structures of Undamaged DNA Dodecamer Duplex | | Descriptor: | DNA (5'-D(*DCP*DCP*DTP*DCP*DTP*DGP*DGP*DTP*DCP*DTP*DCP*DC)-3'), DNA (5'-D(*DGP*DGP*DAP*DGP*DAP*DCP*DCP*DAP*DGP*DAP*DGP*DG)-3') | | Authors: | Bhattacharyya, D, King, C.L, Chaney, S.G, Campbell, S.L. | | Deposit date: | 2008-02-15 | | Release date: | 2009-02-03 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Flanking Bases Influence the Nature of DNA Distortion by Platinum 1,2-Intrastrand (GG) Cross-Links.
Plos One, 6, 2011
|
|
2K0T
 
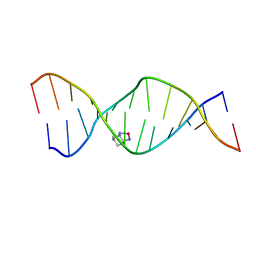 | | High Resolution Solution NMR Structures of Oxaliplatin-DNA Adduct | | Descriptor: | CYCLOHEXANE-1(R),2(R)-DIAMINE-PLATINUM(II), DNA (5'-D(*DCP*DCP*DTP*DCP*DTP*DGP*DGP*DTP*DCP*DTP*DCP*DC)-3'), DNA (5'-D(*DGP*DGP*DAP*DGP*DAP*DCP*DCP*DAP*DGP*DAP*DGP*DG)-3') | | Authors: | Bhattacharyya, D, King, C.L, Chaney, S.G, Campbell, S.L. | | Deposit date: | 2008-02-14 | | Release date: | 2009-02-03 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Flanking Bases Influence the Nature of DNA Distortion by Platinum 1,2-Intrastrand (GG) Cross-Links.
Plos One, 6, 2011
|
|
1JKV
 
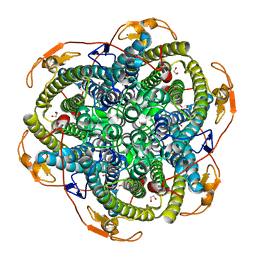 | | Crystal Structure of Manganese Catalase from Lactobacillus plantarum complexed with azide | | Descriptor: | 1,2-ETHANEDIOL, AZIDE ION, CALCIUM ION, ... | | Authors: | Barynin, V.V, Whittaker, M.M, Antonyuk, S.V, Lamzin, V.S, Harrison, P.M, Artymiuk, P.J, Whittaker, J.W. | | Deposit date: | 2001-07-13 | | Release date: | 2002-07-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Crystal structure of manganese catalase from Lactobacillus plantarum.
Structure, 9, 2001
|
|
1E7C
 
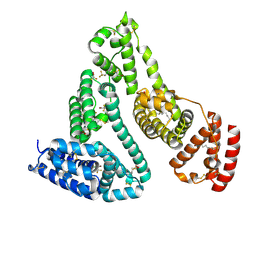 | | HUMAN SERUM ALBUMIN COMPLEXED WITH MYRISTIC ACID and the general anesthetic halothane | | Descriptor: | 2-BROMO-2-CHLORO-1,1,1-TRIFLUOROETHANE, MYRISTIC ACID, SERUM ALBUMIN | | Authors: | Bhattacharya, A.A, Curry, S, Franks, N.P. | | Deposit date: | 2000-08-26 | | Release date: | 2001-01-17 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Binding of the General Anesthetics Propofol and Halothane to Human Serum Albumin. High Resolution Crystal Structures
J.Biol.Chem., 275, 2000
|
|
1E7A
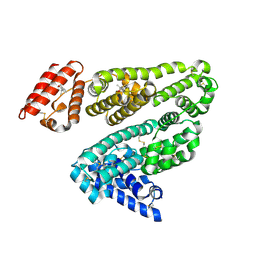 
 | |
1E78
 
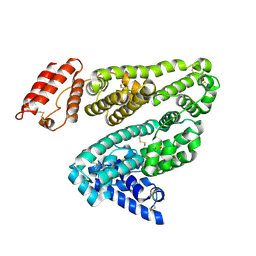 | |
1E7B
 
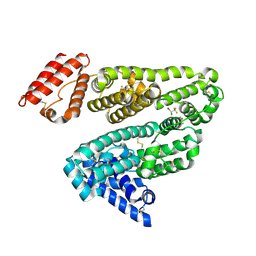 | | Crystal structure of human serum albumin complexed with the general anesthetic halothane | | Descriptor: | 2-BROMO-2-CHLORO-1,1,1-TRIFLUOROETHANE, SERUM ALBUMIN | | Authors: | Bhattacharya, A.A, Curry, S, Franks, N.P. | | Deposit date: | 2000-08-26 | | Release date: | 2001-01-17 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Binding of the General Anesthetics Propofol and Halothane to Human Serum Albumin. High Resolution Crystal Structures
J.Biol.Chem., 275, 2000
|
|
9LPB
 
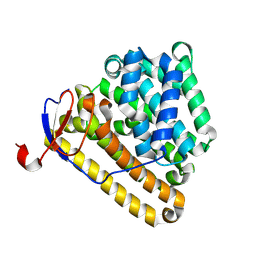 | | Terpene cyclase AriE mutant-D128A | | Descriptor: | Terpene synthase | | Authors: | Li, F.R, Yang, Q, He, J.Y, Sun, X.R, Pan, X.M, Xu, H.M, Rudolf, J.D, Dong, L.B. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-07 | | Last modified: | 2025-06-11 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Crystal Structure and Catalytic Mechanism of the cis-Eunicellane Cyclase AriE.
Chemistry, 31, 2025
|
|
3DHY
 
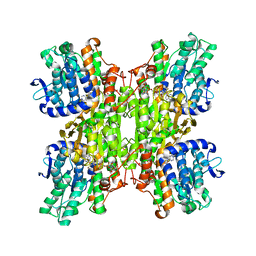 | | Crystal Structures of Mycobacterium tuberculosis S-Adenosyl-L-Homocysteine Hydrolase in Ternary Complex with Substrate and Inhibitors | | Descriptor: | 5'-S-ethyl-5'-thioadenosine, Adenosylhomocysteinase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Shetty, N.D, Ioerger, T.R, Gokulan, K, Reddy, M.C.M, Owen, J.L, Sacchettini, J.C, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2008-06-19 | | Release date: | 2008-09-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of Mycobacterium tuberculosis S-adenosyl-L-homocysteine hydrolase in ternary complex with substrate and inhibitors.
Protein Sci., 17, 2008
|
|
7CK5
 
 | | Solution structure of 28 amino acid polypeptide (354-381) in Plantago asiatica mosaic virus replicase bound to SDS micelle | | Descriptor: | PlAMV replicase peptide from RNA-dependent RNA polymerase | | Authors: | Komatsu, K, Sasaki, N, Yoshida, T, Suzuki, K, Masujima, Y, Hashimoto, M, Watanabe, S, Tochio, N, Kigawa, T, Yamaji, Y, Oshima, K, Namba, S, Nelson, R, Arie, T. | | Deposit date: | 2020-07-15 | | Release date: | 2021-07-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Identification of a Proline-Kinked Amphipathic alpha-Helix Downstream from the Methyltransferase Domain of a Potexvirus Replicase and Its Role in Virus Replication and Perinuclear Complex Formation.
J.Virol., 95, 2021
|
|
6DFR
 
 | |
