9INR
 
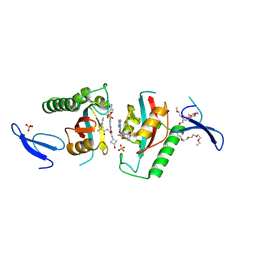 | | Crystal structure of PIN1 in complex with inhibitor C3 | | Descriptor: | 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1, SULFATE ION, ... | | Authors: | Zhang, L.Y. | | Deposit date: | 2024-07-08 | | Release date: | 2024-09-25 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Re-Evaluating PIN1 as a Therapeutic Target in Oncology Using Neutral Inhibitors and PROTACs.
J.Med.Chem., 67, 2024
|
|
9CK3
 
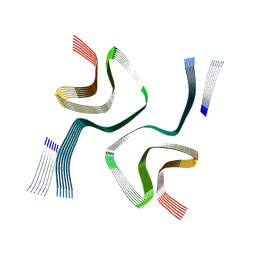 | | Cryo-EM structure of in-vitro alpha-synuclein fibril | | Descriptor: | Alpha-synuclein | | Authors: | Sanchez, J.C, Borcik, C.G, Tonelli, M, Sibert, B, Rienstra, C.M, Wright, E.R. | | Deposit date: | 2024-07-08 | | Release date: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (2.04 Å) | | Cite: | Multimodal High-Resolution Structure Determination Of Alpha-Synuclein Fibrils
To Be Published
|
|
9CJ0
 
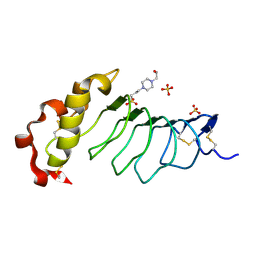 | |
9CJ9
 
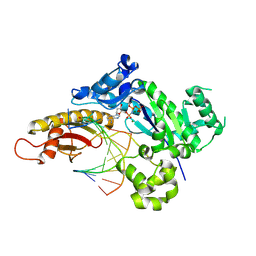 | |
9CIQ
 
 | | Crystal structure of human polymerase eta with incoming threofuranosyl thymine nucleoside triphosphate opposite template dA | | Descriptor: | CALCIUM ION, DI(HYDROXYETHYL)ETHER, DNA (5'-D(*AP*GP*CP*GP*TP*CP*AP*T)-3'), ... | | Authors: | Patra, A, Tomar, R, Stone, M.P, Egli, M. | | Deposit date: | 2024-07-04 | | Release date: | 2024-09-18 | | Last modified: | 2024-09-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | DNA Replication across alpha-l-(3'-2')-Threofuranosyl Nucleotides Mediated by Human DNA Polymerase eta.
Biochemistry, 2024
|
|
9FYP
 
 | | Cryo EM structure of the type 3B polymorph of alpha-synuclein at low pH. | | Descriptor: | Alpha-synuclein, CHLORIDE ION | | Authors: | Frey, L, Qureshi, B.M, Kwiatkowski, W, Rhyner, D, Greenwald, J, Riek, R. | | Deposit date: | 2024-07-03 | | Release date: | 2024-07-17 | | Last modified: | 2024-09-11 | | Method: | ELECTRON MICROSCOPY (2.23 Å) | | Cite: | On the pH-dependence of alpha-synuclein amyloid polymorphism and the role of secondary nucleation in seed-based amyloid propagation.
Elife, 12, 2024
|
|
9CIH
 
 | |
9CHW
 
 | |
9CI9
 
 | |
9FXI
 
 | | Crystal structure of cobalt(II)-substituted double mutant Y115E Y117E human Glutaminyl Cyclase in complex with SEN177 | | Descriptor: | 2-fluoranyl-5-[2-[4-(4-methyl-1,2,4-triazol-3-yl)piperidin-1-yl]pyridin-3-yl]pyridine, COBALT (II) ION, Glutaminyl-peptide cyclotransferase, ... | | Authors: | Tassone, G, Pozzi, C, Mangani, S. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | Metal Ion Binding to Human Glutaminyl Cyclase: A Structural Perspective.
Int J Mol Sci, 25, 2024
|
|
9FXG
 
 | | Crystal structure of double mutant Y115E Y117E human Glutaminyl Cyclase in apo-state | | Descriptor: | 1,2-ETHANEDIOL, Glutaminyl-peptide cyclotransferase, SULFATE ION | | Authors: | Tassone, G, Pozzi, C, Mangani, S. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Metal Ion Binding to Human Glutaminyl Cyclase: A Structural Perspective.
Int J Mol Sci, 25, 2024
|
|
9FXJ
 
 | | Crystal structure of cobalt(II)-substituted double mutant Y115E Y117E human Glutaminyl Cyclase in complex with PBD-150 | | Descriptor: | 1-(3,4-dimethoxyphenyl)-3-[3-(1H-imidazol-1-yl)propyl]thiourea, COBALT (II) ION, Glutaminyl-peptide cyclotransferase, ... | | Authors: | Tassone, G, Pozzi, C, Mangani, S. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | Metal Ion Binding to Human Glutaminyl Cyclase: A Structural Perspective.
Int J Mol Sci, 25, 2024
|
|
9FXH
 
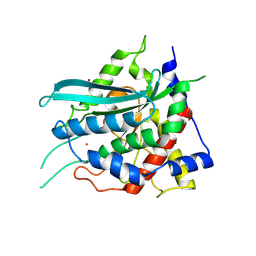 | | Crystal structure of cobalt(II)-substituted double mutant Y115E Y117E human Glutaminyl Cyclase | | Descriptor: | COBALT (II) ION, GLYCEROL, Glutaminyl-peptide cyclotransferase, ... | | Authors: | Tassone, G, Pozzi, C, Mangani, S. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Metal Ion Binding to Human Glutaminyl Cyclase: A Structural Perspective.
Int J Mol Sci, 25, 2024
|
|
9CGZ
 
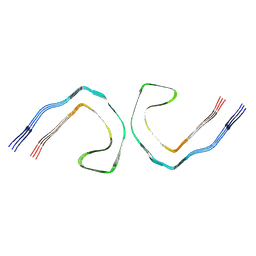 | | Alzheimer's Disease Seeded Mixed 0N4R and 0N3R Tau Fibrils | | Descriptor: | Isoform Fetal-tau of Microtubule-associated protein tau | | Authors: | Duan, P, Dregni, A.J, Xu, H, Changolkar, L, Lee, V.M.-Y, Hong, M. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-11 | | Last modified: | 2024-10-02 | | Method: | ELECTRON MICROSCOPY (2.69 Å) | | Cite: | Alzheimer's disease seeded tau forms paired helical filaments yet lacks seeding potential.
J.Biol.Chem., 300, 2024
|
|
9CGX
 | | Alzheimer's Disease Seeded 0N3R Tau Fibrils | | Descriptor: | Isoform Fetal-tau of Microtubule-associated protein tau | | Authors: | Duan, P, Dregni, A.J, Xu, H, Changolkar, L, Lee, V.M.-Y, Hong, M. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-11 | | Last modified: | 2024-10-02 | | Method: | ELECTRON MICROSCOPY (2.97 Å) | | Cite: | Alzheimer's disease seeded tau forms paired helical filaments yet lacks seeding potential.
J.Biol.Chem., 300, 2024
|
|
9CGO
 
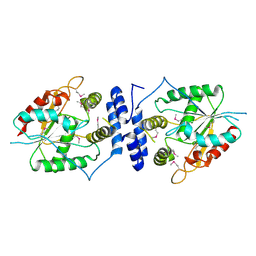 | | Tylosin thioesterase domain (TylG5 TE) | | Descriptor: | Tylactone synthase module 7 | | Authors: | Smith, J.L, Choudhary, V. | | Deposit date: | 2024-06-30 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Substrate Trapping in Polyketide Synthase Thioesterase Domains: Structural Basis for Macrolactone Formation
Acs Catalysis, 14, 2024
|
|
9CGN
 | | Pikromycin Thioesterase with heptaketide adduct | | Descriptor: | (2S,4R,5S,6S,8R,12R,13R)-5,13-dihydroxy-2,4,6,8,12-pentamethyl-3,9-dioxopentadecanal, Narbonolide/10-deoxymethynolide synthase PikA4, module 6 | | Authors: | Smith, J.L, Choudhary, V. | | Deposit date: | 2024-06-30 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Substrate Trapping in Polyketide Synthase Thioesterase Domains: Structural Basis for Macrolactone Formation
Acs Catalysis, 14, 2024
|
|
9FWG
 | | LSD1/CoREST bound to bomedemstat | | Descriptor: | Bomedemstat FAD adduct, Lysine-specific histone demethylase 1A, REST corepressor 1 | | Authors: | Speranzini, V, Mattevi, A. | | Deposit date: | 2024-06-30 | | Release date: | 2024-07-10 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Characterization of structural, biochemical, pharmacokinetic, and pharmacodynamic properties of the LSD1 inhibitor bomedemstat in preclinical models.
Prostate, 84, 2024
|
|
9CGI
 
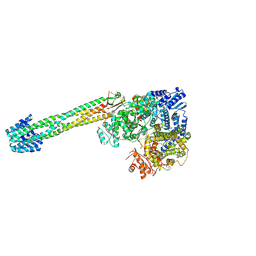 | |
9CGL
 | | Pikromycin Thioesterase Doubly Protected DAP | | Descriptor: | 2-{[(1R)-1-(6-nitro-2H-1,3-benzodioxol-5-yl)ethyl]sulfanyl}ethyl formate, Narbonolide/10-deoxymethynolide synthase PikA4, module 6 | | Authors: | McCullough, T.M, Smith, J.L. | | Deposit date: | 2024-06-29 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Substrate Trapping in Polyketide Synthase Thioesterase Domains: Structural Basis for Macrolactone Formation
Acs Catalysis, 14, 2024
|
|
9CG5
 
 | |
9CG6
 | |
9CG7
 
 | |
9CER
 
 | | Guillardia theta Fanzor (GtFz) State 1 | | Descriptor: | Maltose/maltodextrin-binding periplasmic protein,Guillardia theta Fanzor1, RNA (142-MER), ZINC ION | | Authors: | Xu, P, Saito, M, Zhang, F. | | Deposit date: | 2024-06-27 | | Release date: | 2024-09-11 | | Last modified: | 2024-10-02 | | Method: | ELECTRON MICROSCOPY (4.7 Å) | | Cite: | Structural insights into the diversity and DNA cleavage mechanism of Fanzor.
Cell, 187, 2024
|
|
9CEX
 
 | | Spizellomyces punctatus Fanzor (SpuFz) State 4 | | Descriptor: | DNA (29-MER), DNA (5'-D(*(MG)*(MG)P*AP*AP*AP*AP*AP*AP*AP*AP*AP*AP*AP*AP*AP*AP*AP*A)-3'), DNA (5'-D(P*CP*GP*GP*TP*AP*CP*CP*CP*GP*GP*GP*CP*AP*TP*A)-3'), ... | | Authors: | Xu, P, Saito, M, Zhang, F. | | Deposit date: | 2024-06-27 | | Release date: | 2024-09-11 | | Last modified: | 2024-10-02 | | Method: | ELECTRON MICROSCOPY (3.27 Å) | | Cite: | Structural insights into the diversity and DNA cleavage mechanism of Fanzor.
Cell, 187, 2024
|
|
