1H9K
 
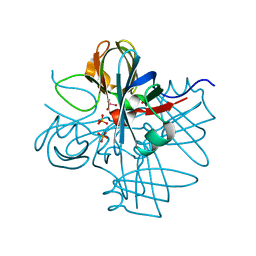 | | Two crystal structures of the cytoplasmic molybdate-binding protein ModG suggest a novel cooperative binding mechanism and provide insights into ligand-binding specificity. Phosphate-grown form with tungstate and phosphate bound | | Descriptor: | MOLYBDENUM-BINDING-PROTEIN, PHOSPHATE ION, TUNGSTATE(VI)ION | | Authors: | Delarbre, L, Stevenson, C.E.M, White, D.J, Mitchenall, L.A, Pau, R.N, Lawson, D.M. | | Deposit date: | 2001-03-13 | | Release date: | 2001-05-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Two Crystal Structures of the Cytoplasmic Molybdate-Binding Protein Modg Suggest a Novel Cooperative Binding Mechanism and Provide Insights Into Ligand-Binding Specificity
J.Mol.Biol., 308, 2001
|
|
1GT7
 
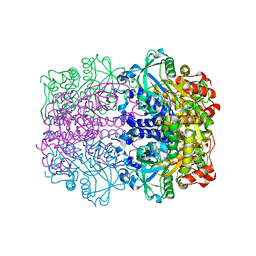 | | L-rhamnulose-1-phosphate aldolase from Escherichia coli | | Descriptor: | PHOSPHOGLYCOLOHYDROXAMIC ACID, RHAMNULOSE-1-PHOSPHATE ALDOLASE, ZINC ION | | Authors: | Kroemer, M, Schulz, G.E. | | Deposit date: | 2002-01-14 | | Release date: | 2002-05-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The Structure of L-Rhamnulose-1-Phosphate Aldolase (Class II) Solved by Low-Resolution Sir Phasing and 20-Fold Ncs Averaging
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1GYT
 
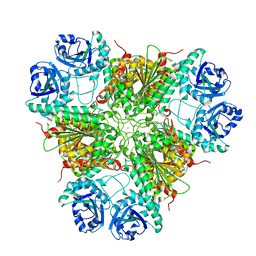 | | E. coli Aminopeptidase A (PepA) | | Descriptor: | CARBONATE ION, CHLORIDE ION, CYTOSOL AMINOPEPTIDASE, ... | | Authors: | Straeter, N. | | Deposit date: | 2002-04-29 | | Release date: | 2002-06-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-Ray Structure of Aminopeptidase a from Escherichia Coli and a Model for the Nucleoprotein Complex in Xer Site-Specific Recombination
Embo J., 18, 1999
|
|
1H2N
 
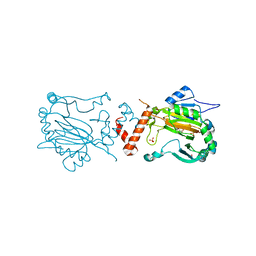 | | Factor Inhibiting HIF-1 alpha | | Descriptor: | 2-OXOGLUTARIC ACID, FE (II) ION, HYPOXIA-INDUCIBLE FACTOR 1-ALPHA INHIBITOR, ... | | Authors: | Elkins, J.M, Hewitson, K.S, McNeill, L.A, Schlemminger, I, Seibel, J.F, Schofield, C.J. | | Deposit date: | 2002-08-12 | | Release date: | 2002-11-28 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Structure of Factor-Inhibiting Hypoxia-Inducible Factor (Hif) Reveals Mechanism of Oxidative Modification of Hif-1Alpha
J.Biol.Chem., 278, 2003
|
|
1GUZ
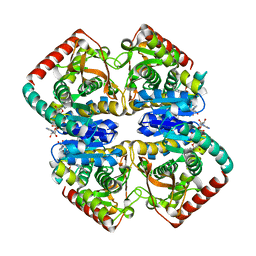 
 | | Structural Basis for Thermophilic Protein Stability: Structures of Thermophilic and Mesophilic Malate Dehydrogenases | | Descriptor: | MALATE DEHYDROGENASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Dalhus, B, Sarinen, M, Sauer, U.H, Eklund, P, Johansson, K, Karlsson, A, Ramaswamy, S, Bjork, A, Synstad, B, Naterstad, K, Sirevag, R, Eklund, H. | | Deposit date: | 2002-02-04 | | Release date: | 2002-02-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Thermophilic Protein Stability: Structures of Thermophilic and Mesophilic Malate Dehydrogenases
J.Mol.Biol., 318, 2002
|
|
1GWG
 
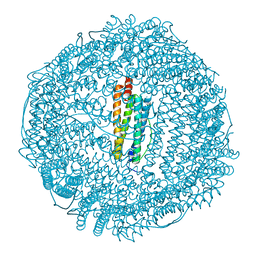 | | Tri-iodide derivative of apoferritin | | Descriptor: | CADMIUM ION, FERRITIN LIGHT CHAIN, IODIDE ION | | Authors: | Evans, G, Bricogne, G. | | Deposit date: | 2002-03-15 | | Release date: | 2002-06-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Triiodide Derivatization and Combinatorial Counter-Ion Replacement: Two Methods for Enhancing Phasing Signal Using Laboratory Cu Kalpha X-Ray Equipment
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1HF8
 
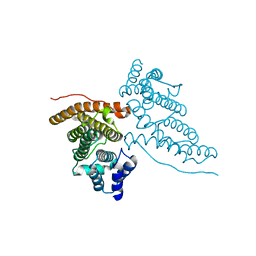 | |
1HC1
 
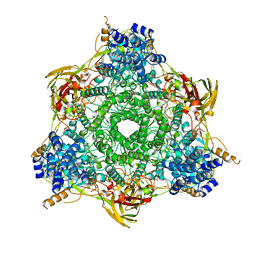 | |
1HLQ
 
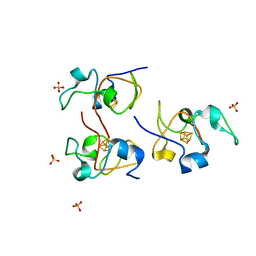 | | CRYSTAL STRUCTURE OF RHODOFERAX FERMENTANS HIGH POTENTIAL IRON-SULFUR PROTEIN REFINED TO 1.45 A | | Descriptor: | HIGH-POTENTIAL IRON-SULFUR PROTEIN, IRON/SULFUR CLUSTER, SULFATE ION | | Authors: | Gonzalez, A, Ciurli, S, Benini, S. | | Deposit date: | 2000-12-01 | | Release date: | 2003-06-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structure of Rhodoferax fermentans high-potential iron-sulfur protein solved by MAD.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1H6V
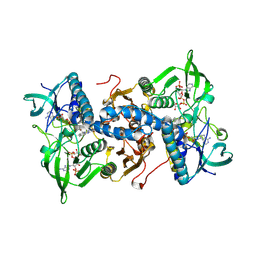 
 | | Mammalian thioredoxin reductase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, THIOREDOXIN REDUCTASE | | Authors: | Sandalova, T, Zhong, L, Lindqvist, Y, Holmgren, A, Schneider, G. | | Deposit date: | 2001-06-27 | | Release date: | 2001-08-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Three-Dimensional Structure of a Mammalian Thioredoxin Reductase: Implication for Mechanism and Evolution of a Selenocysteine Dependent Enzyme
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1GUL
 
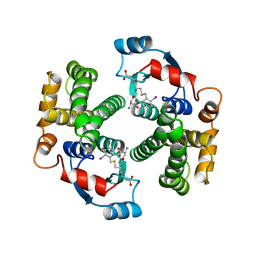 | | HUMAN GLUTATHIONE TRANSFERASE A4-4 COMPLEX WITH IODOBENZYL GLUTATHIONE | | Descriptor: | GAMMA-GLUTAMYL[S-(2-IODOBENZYL)CYSTEINYL]GLYCINE, Glutathione S-transferase A4 | | Authors: | Bruns, C.M, Hubatsch, I, Ridderstrom, M, Mannervik, B, Tainer, J.A. | | Deposit date: | 1998-06-10 | | Release date: | 1999-01-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Human glutathione transferase A4-4 crystal structures and mutagenesis reveal the basis of high catalytic efficiency with toxic lipid peroxidation products
J.Mol.Biol., 288, 1999
|
|
1GWD
 
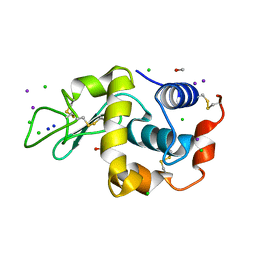 | | Tri-iodide derivative of hen egg-white lysozyme | | Descriptor: | 1,2-ETHANEDIOL, CARBON MONOXIDE, CHLORIDE ION, ... | | Authors: | Evans, G, Bricogne, G. | | Deposit date: | 2002-03-14 | | Release date: | 2002-06-06 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Triiodide Derivatization and Combinatorial Counter-Ion Replacement: Two Methods for Enhancing Phasing Signal Using Laboratory Cu Kalpha X-Ray Equipment
Acta Crystallogr.,Sect.D, 58, 2002
|
|
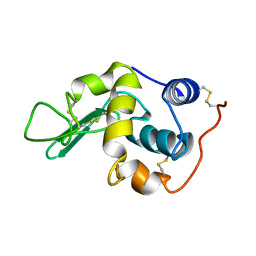
1GXV
 
 | |
1GZ5
 
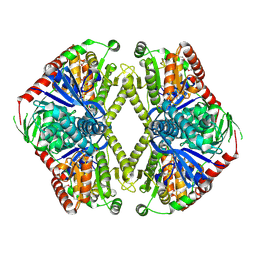 | | Trehalose-6-phosphate synthase. OtsA | | Descriptor: | 6-O-phosphono-alpha-D-glucopyranose, ALPHA-TREHALOSE-PHOSPHATE SYNTHASE, IMIDAZOLE, ... | | Authors: | Gibson, R.P, Turkenburg, J.P, Davies, G.J. | | Deposit date: | 2002-05-15 | | Release date: | 2003-02-07 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Insights Into Trehalose Synthesis Provided by the Structure of the Retaining Glycosyltransferase Otsa
Chem.Biol., 9, 2002
|
|
1H6G
 
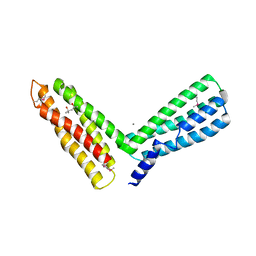 | | alpha-catenin M-domain | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ALPHA-1 CATENIN, CALCIUM ION, ... | | Authors: | Yang, J, Dokurno, P, Tonks, N.K, Barford, D. | | Deposit date: | 2001-06-14 | | Release date: | 2001-08-07 | | Last modified: | 2016-02-10 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of the M-Fragment of Alpha-Catenin: Implications for Modulation of Cell Adhesion.
Embo J., 20, 2001
|
|
1HE1
 
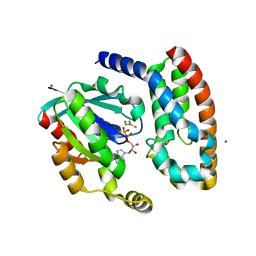 | | Crystal structure of the complex between the GAP domain of the Pseudomonas aeruginosa ExoS toxin and human Rac | | Descriptor: | ALUMINUM FLUORIDE, EXOENZYME S, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Wurtele, M, Wolf, E, Pederson, K.J, Buchwald, G, Ahmadian, M.R, Barbieri, J.T, Wittinghofer, A. | | Deposit date: | 2000-11-18 | | Release date: | 2001-01-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | How the Pseudomonas Aeruginosa Exos Toxin Downregulates Rac
Nat.Struct.Biol., 8, 2001
|
|
1HAI
 
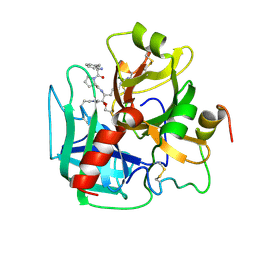 | | THE ISOMORPHOUS STRUCTURES OF PRETHROMBIN2, HIRUGEN-AND PPACK-THROMBIN: CHANGES ACCOMPANYING ACTIVATION AND EXOSITE BINDING TO THROMBIN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ALPHA-THROMBIN (LARGE SUBUNIT), ALPHA-THROMBIN (SMALL SUBUNIT), ... | | Authors: | Tulinsky, A, Vijayalakshmi, J. | | Deposit date: | 1994-06-27 | | Release date: | 1994-12-20 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The isomorphous structures of prethrombin2, hirugen-, and PPACK-thrombin: changes accompanying activation and exosite binding to thrombin.
Protein Sci., 3, 1994
|
|
1HB9
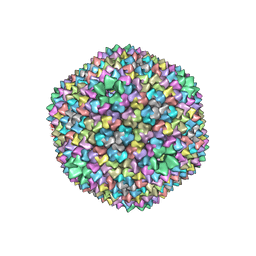 
 | | quasi-atomic resolution model of bacteriophage PRD1 wild type virion, obtained by combined cryo-EM and X-ray crystallography. | | Descriptor: | BACTERIOPHAGE PRD1 | | Authors: | San Martin, C, Burnett, R.M, De Haas, F, Heinkel, R, Rutten, T, Fuller, S.D, Butcher, S.J, Bamford, D.H. | | Deposit date: | 2001-04-13 | | Release date: | 2001-12-05 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (25 Å) | | Cite: | Combined Em/X-Ray Imaging Yields a Quasi-Atomic Model of the Adenovirus-Related Bacteriophage Prd1 and Shows Key Capsid and Membrane Interactions
Structure, 9, 2001
|
|
1HGY
 
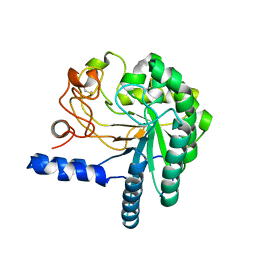 | | CEL6A D221A mutant | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CELLOBIOHYDROLASE CEL6A (FORMERLY CALLED CBH II), alpha-D-glucopyranose, ... | | Authors: | Zou, J.-Y, Kleywegt, G.J, Jones, T.A. | | Deposit date: | 2000-12-15 | | Release date: | 2002-01-15 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Active Site of Cellobiohydrolase Cel6A from Trichoderma Reesei: The Roles of Aspartic Acids D221 and D175
J.Am.Chem.Soc., 124, 2002
|
|
1H29
 
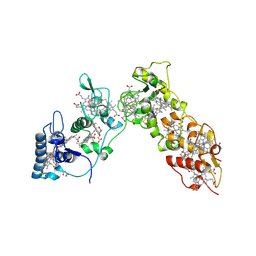 | | Sulfate respiration in Desulfovibrio vulgaris Hildenborough: Structure of the 16-heme Cytochrome c HmcA at 2.5 A resolution and a view of its role in transmembrane electron transfer | | Descriptor: | HEME C, HIGH-MOLECULAR-WEIGHT CYTOCHROME C | | Authors: | Matias, P.M, Coelho, A.V, Valente, F.M.A, Placido, D, Legall, J, Xavier, A.V, Pereira, I.A.C, Carrondo, M.A. | | Deposit date: | 2002-08-01 | | Release date: | 2002-10-02 | | Last modified: | 2019-05-15 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Sulfate Respiration in Desulfovibrio Vulgaris Hildenborough: Structure of the 16-Heme Cytochrome C Hmca at 2.5 A Resolution and a View of its Role in Transmembrane Electron Transfer
J.Biol.Chem., 277, 2002
|
|
1GXX
 
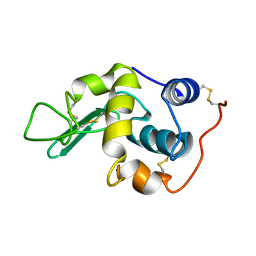 | |
1GZM
 
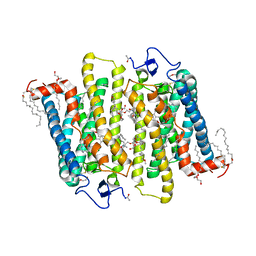 | | Structure of Bovine Rhodopsin in a Trigonal Crystal Form | | Descriptor: | (HYDROXYETHYLOXY)TRI(ETHYLOXY)OCTANE, DI-PALMITOYL-3-SN-PHOSPHATIDYLETHANOLAMINE, LAURYL DIMETHYLAMINE-N-OXIDE, ... | | Authors: | Li, J. | | Deposit date: | 2002-05-24 | | Release date: | 2003-11-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structure of Bovine Rhodopsin in a Trigonal Crystal Form
J.Mol.Biol., 343, 2004
|
|
1H15
 
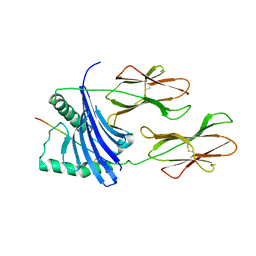 | | X-ray crystal structure of HLA-DRA1*0101/DRB5*0101 complexed with a peptide from Epstein Barr Virus DNA polymerase | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DNA POLYMERASE, ... | | Authors: | Lang, H, Jacobsen, H, Ikemizu, S, Andersson, C, Harlos, K, Madsen, L, Hjorth, P, Sondergaard, L, Svejgaard, A, Wucherpfennig, K, Stuart, D.I, Bell, J.I, Jones, E.Y, Fugger, L. | | Deposit date: | 2002-07-02 | | Release date: | 2002-10-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | A Functional and Structural Basis for Tcr Cross-Reactivity in Multiple Sclerosis
Nat.Immunol., 3, 2002
|
|
1GO1
 
 | |
1H7W
 
 | | Dihydropyrimidine dehydrogenase (DPD) from pig | | Descriptor: | DIHYDROPYRIMIDINE DEHYDROGENASE, FLAVIN MONONUCLEOTIDE, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Dobritzsch, D, Schneider, G, Schnackerz, K.D, Lindqvist, Y. | | Deposit date: | 2001-01-19 | | Release date: | 2001-02-23 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of dihydropyrimidine dehydrogenase, a major determinant of the pharmacokinetics of the anti-cancer drug 5-fluorouracil.
EMBO J., 20, 2001
|
|
