5LVZ
 
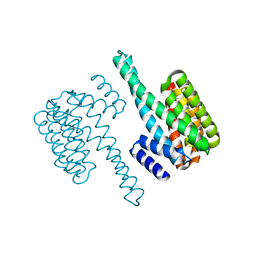 | | Crystal structure of yeast 14-3-3 protein from Lachancea thermotolerans | | Descriptor: | KLTH0G14146p | | Authors: | Klima, M, Boura, E. | | Deposit date: | 2016-09-14 | | Release date: | 2016-10-19 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structures of a yeast 14-3-3 protein from Lachancea thermotolerans in the unliganded form and bound to a human lipid kinase PI4KB-derived peptide reveal high evolutionary conservation.
Acta Crystallogr.,Sect.F, 72, 2016
|
|
2M0P
 
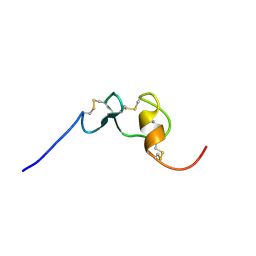 | | Solution structure of the tenth complement type repeat of human megalin | | Descriptor: | CALCIUM ION, Low-density lipoprotein receptor-related protein 2 | | Authors: | Dagil, R, Kragelund, B. | | Deposit date: | 2012-11-01 | | Release date: | 2013-01-09 | | Last modified: | 2024-11-20 | | Method: | SOLUTION NMR | | Cite: | Gentamicin binds to the megalin receptor as a competitive inhibitor using the common ligand binding motif of complement type repeats: insight from the nmr structure of the 10th complement type repeat domain alone and in complex with gentamicin.
J.Biol.Chem., 288, 2013
|
|
4LQ5
 
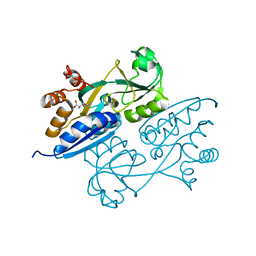 | | Crystal structure of ligand binding domain of CysB, a LysR member from Salmonella typhimurium LT2 in complex with effector ligand, O-acetylserine at 2.8A | | Descriptor: | HTH-type transcriptional regulator CysB, O-ACETYLSERINE | | Authors: | Mittal, M, Singh, A.K, Kumaran, S. | | Deposit date: | 2013-07-17 | | Release date: | 2014-07-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.803 Å) | | Cite: | rystal structure of ligand binding domain of CysB, a LysR member from Salmonella typhimurium LT2 in complex with effector ligand, O-acetylserine at 2.8A
TO BE PUBLISHED
|
|
1H2R
 
 | |
1ZBC
 
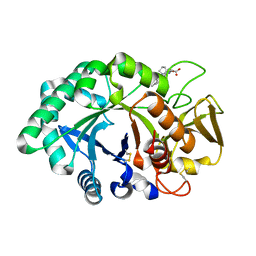 | | Crystal Structure of the porcine signalling protein liganded with the peptide Trp-Pro-Trp (WPW) at 2.3 A resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3 mer peptide, signal processing protein | | Authors: | Srivastava, D.B, Kaur, P, Kumar, J, Somvanshi, R.K, Sharma, S, Dey, S, Singh, T.P. | | Deposit date: | 2005-04-08 | | Release date: | 2005-04-19 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Crystal Structure of the porcine signalling protein liganded with the peptide Trp-Pro-Trp (WPW) at 2.3 A resolution
To be Published
|
|
5LOO
 
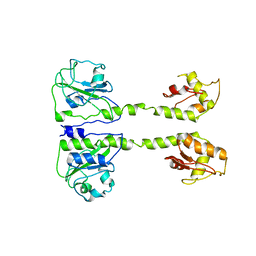 | |
1O9I
 
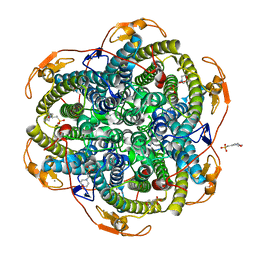 | | Crystal structure of the Y42F mutant of manganese catalase from Lactobacillus plantarum at 1.33A resolution | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, MANGANESE (III) ION, ... | | Authors: | Barynin, V.V, Whittaker, M.M, Whittaker, J.W. | | Deposit date: | 2002-12-13 | | Release date: | 2003-12-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Outer Sphere Mutagenesis of Lactobacillus Plantarum Manganese Catalase Disrupts the Cluster Core. Mechanistic Implications.
Eur.J.Biochem., 270, 2003
|
|
2J4K
 | | Crystal structure of uridylate kinase from Sulfolobus solfataricus in complex with UMP to 2.2 Angstrom resolution | | Descriptor: | CADMIUM ION, MAGNESIUM ION, URIDINE-5'-MONOPHOSPHATE, ... | | Authors: | Jensen, K.S, Johansson, E, Jensen, K.F. | | Deposit date: | 2006-09-01 | | Release date: | 2007-02-27 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and Enzymatic Investigation of the Sulfolobus Solfataricus Uridylate Kinase Shows Competitive Utp Inhibition and the Lack of GTP Stimulation
Biochemistry, 46, 2007
|
|
2FVY
 | | High Resolution Glucose Bound Crystal Structure of GGBP | | Descriptor: | ACETATE ION, CALCIUM ION, CARBON DIOXIDE, ... | | Authors: | Borrok, M.J, Kiessling, L.L, Forest, K.T. | | Deposit date: | 2006-01-31 | | Release date: | 2007-02-06 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.92 Å) | | Cite: | Conformational changes of glucose/galactose-binding protein illuminated by open, unliganded, and ultra-high-resolution ligand-bound structures.
Protein Sci., 16, 2007
|
|
7YXZ
 
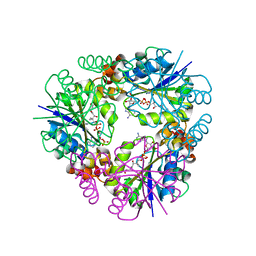 | |
7TXE
 
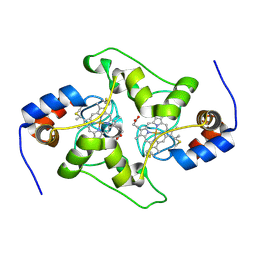 | | Plasmodium falciparum Cyt c2 DSD | | Descriptor: | Cytochrome c2, HEME C | | Authors: | Hill, C.P, Wienkers, H.J, Whitby, F.G. | | Deposit date: | 2022-02-08 | | Release date: | 2023-05-10 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Direct tests of cytochrome c and c1 functions in the electron transport chain of malaria parasites
Proc Natl Acad Sci U S A, 120, 2023
|
|
1ZDS
 
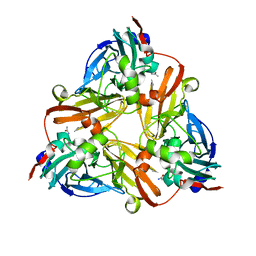 | | Crystal Structure of Met150Gly AfNiR with Acetamide Bound | | Descriptor: | ACETAMIDE, COPPER (II) ION, Copper-containing nitrite reductase | | Authors: | Wijma, H.J, MacPherson, I.S, Alexandre, M, Diederix, R.E.M, Canters, G.W, Murphy, M.E.P, Verbeet, M.P. | | Deposit date: | 2005-04-14 | | Release date: | 2006-03-28 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | A rearranging ligand enables allosteric control of catalytic activity in copper-containing nitrite reductase.
J.Mol.Biol., 358, 2006
|
|
5FJV
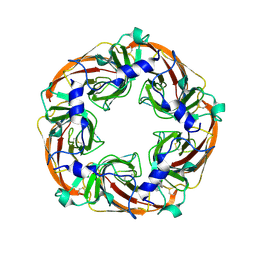 
 | | Crystal structure of the extracellular domain of alpha2 nicotinic acetylcholine receptor in pentameric assembly | | Descriptor: | EPIBATIDINE, NEURONAL ACETYLCHOLINE RECEPTOR SUBUNIT ALPHA-2 | | Authors: | Giastas, P, Kouvatsos, N, Chroni-Tzartou, D, Tzartos, S.J. | | Deposit date: | 2015-10-13 | | Release date: | 2016-08-10 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal Structure of a Human Neuronal Nachr Extracellular Domain in Pentameric Assembly: Ligand-Bound Alpah2 Homopentamer.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
4CAK
 
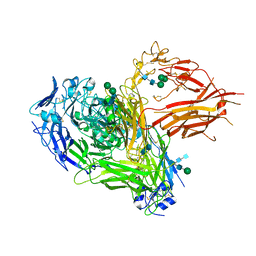 | | Three-dimensional reconstruction of intact human integrin alphaIIbbeta3 in a phospholipid bilayer nanodisc | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Integrin alpha-IIb, ... | | Authors: | Choi, W.S, Rice, W.J, Stokes, D.L, Coller, B.S. | | Deposit date: | 2013-10-08 | | Release date: | 2013-10-30 | | Last modified: | 2025-04-09 | | Method: | ELECTRON MICROSCOPY (20.5 Å) | | Cite: | Three-Dimensional Reconstruction of Intact Human Integrin Alphaiibbeta3; New Implications for Activation-Dependent Ligand Binding.
Blood, 122, 2013
|
|
5LNH
 
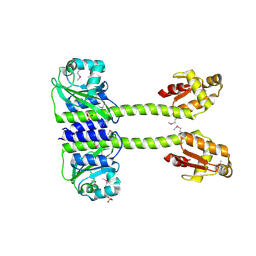 | | Structure of full length Unliganded CodY from Bacillus subtilis | | Descriptor: | GTP-sensing transcriptional pleiotropic repressor CodY, SULFATE ION | | Authors: | Wilkinson, A.J, Levdikov, V.M, Blagova, E.V. | | Deposit date: | 2016-08-04 | | Release date: | 2017-01-11 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of the Branched-chain Amino Acid and GTP-sensing Global Regulator, CodY, from Bacillus subtilis.
J. Biol. Chem., 292, 2017
|
|
2JN3
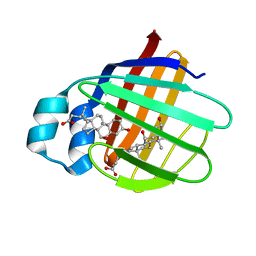 
 | | NMR structure of cl-BABP complexed to chenodeoxycholic acid | | Descriptor: | CHENODEOXYCHOLIC ACID, Fatty acid-binding protein, liver | | Authors: | Eliseo, T, Ragona, L, Catalano, M, Assfalf, M, Paci, M, Zetta, L, Molinari, H, Cicero, D.O. | | Deposit date: | 2006-12-22 | | Release date: | 2007-07-03 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | Structural and dynamic determinants of ligand binding in the ternary complex of chicken liver bile acid binding protein with two bile salts revealed by NMR
Biochemistry, 46, 2007
|
|
6H1N
 
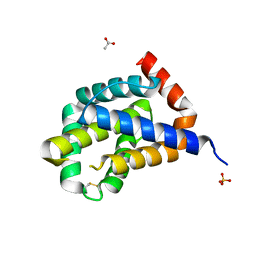 | |
1FI7
 
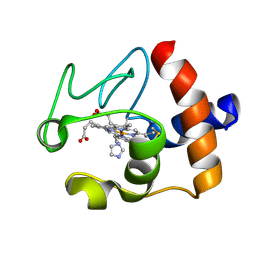 | | Solution structure of the imidazole complex of cytochrome C | | Descriptor: | CYTOCHROME C, HEME C, IMIDAZOLE | | Authors: | Banci, L, Bertini, I, Liu, G, Lu, J, Reddig, T, Tang, W, Wu, Y, Zhu, D. | | Deposit date: | 2000-08-03 | | Release date: | 2000-08-23 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | Effects of extrinsic imidazole ligation on the molecular and electronic structure of cytochrome c
J.Biol.Inorg.Chem., 6, 2001
|
|
4OAL
 
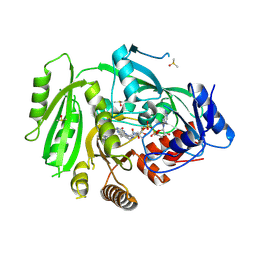 | | Crystal structure of maize cytokinin oxidase/dehydrogenase 4 (ZmCKO4) in complex with phenylurea inhibitor CPPU in alternative spacegroup | | Descriptor: | 1-(2-chloropyridin-4-yl)-3-phenylurea, Cytokinin dehydrogenase 4, DIMETHYL SULFOXIDE, ... | | Authors: | Kopecny, D, Morera, S, Vigouroux, A, Koncitikova, R. | | Deposit date: | 2014-01-05 | | Release date: | 2015-04-01 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Kinetic and structural investigation of the cytokinin oxidase/dehydrogenase active site.
Febs J., 283, 2016
|
|
5A15
 
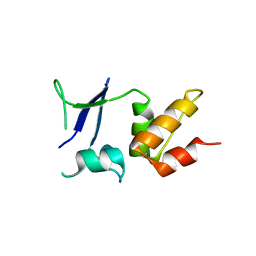 | | Crystal structure of the BTB domain of human KCTD16 | | Descriptor: | BTB/POZ DOMAIN-CONTAINING PROTEIN KCTD16 | | Authors: | Pinkas, D.M, Sanvitale, C.E, Solcan, N, Goubin, S, Canning, P, Dixon Clarke, S.E, Talon, R, Wiggers, H.J, Fitzpatrick, F, Tallant, C, Kopec, J, Chalk, R, Doutch, J, Krojer, T, Burgess-Brown, N.A, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Bullock, A. | | Deposit date: | 2015-04-28 | | Release date: | 2015-11-04 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.76 Å) | | Cite: | Structural complexity in the KCTD family of Cullin3-dependent E3 ubiquitin ligases.
Biochem. J., 474, 2017
|
|
4BSJ
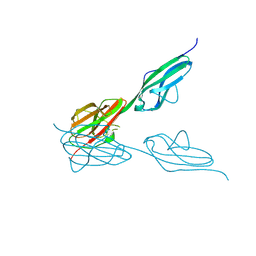 
 | | Crystal structure of VEGFR-3 extracellular domains D4-5 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR 3 | | Authors: | Leppanen, V.-M, Tvorogov, D, Kisko, K, Prota, A.E, Jeltsch, M, Anisimov, A, Markovic-Mueller, S, Stuttfeld, E, Goldie, K.N, Ballmer-Hofer, K, Alitalo, K. | | Deposit date: | 2013-06-10 | | Release date: | 2013-07-31 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and Mechanistic Insights Into Vegfr-3 Ligand Binding and Activation
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
1ZEF
 
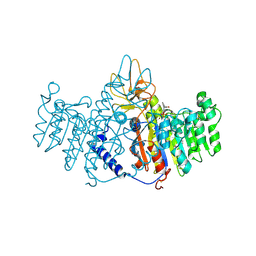 | | structure of alkaline phosphatase from human placenta in complex with its uncompetitive inhibitor L-Phe | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Alkaline phosphatase, CALCIUM ION, ... | | Authors: | Llinas, P, Stura, E.A, Menez, A, Kiss, Z, Stigbrand, T, Millan, J.L, Le Du, M.H. | | Deposit date: | 2005-04-18 | | Release date: | 2005-06-28 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Studies of Human Placental Alkaline Phosphatase in Complex with Functional Ligands.
J.Mol.Biol., 350, 2005
|
|
4EPC
 
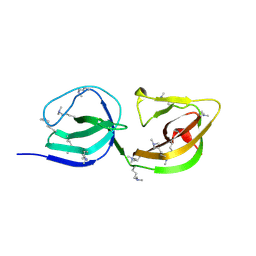 | |
2Y4O
 
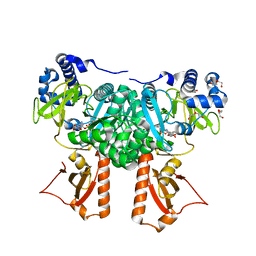 | | Crystal Structure of PaaK2 in complex with phenylacetyl adenylate | | Descriptor: | 5'-O-[HYDROXY(PHENYLACETYL)PHOSPHORYL]ADENOSINE, BETA-MERCAPTOETHANOL, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Law, A, Boulanger, M.J. | | Deposit date: | 2011-01-07 | | Release date: | 2011-03-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Defining a Structural and Kinetic Rationale for Paralogous Copies of Phenylacetate-Coa Ligases from the Cystic Fibrosis Pathogen Burkholderia Cenocepacia J2315.
J.Biol.Chem., 286, 2011
|
|
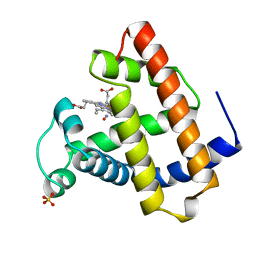
1JDO
 
 | |
