5J2C
 
 | |
5J2J
 
 | |
5J2D
 
 | |
5J29
 
 | |
5J2G
 
 | |
5J2E
 
 | |
5J2I
 
 | |
5J2H
 
 | |
5J2B
 
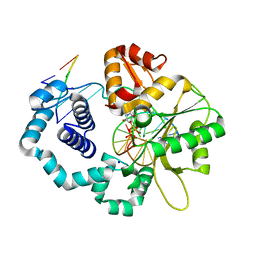 | |
8IR3
 
 | |
5J2A
 
 | |
5J2F
 
 | |
6YBT
 
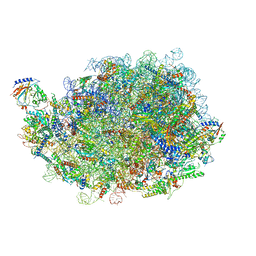
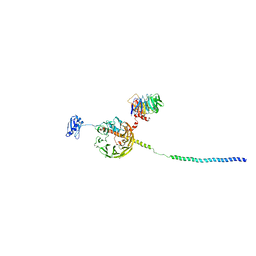 | | Structure of a human 48S translational initiation complex - eIF3bgi | | Descriptor: | Eukaryotic translation initiation factor 3 subunit A, Eukaryotic translation initiation factor 3 subunit B, Eukaryotic translation initiation factor 3 subunit I | | Authors: | Brito Querido, J, Sokabe, M, Kraatz, S, Gordiyenko, Y, Skehel, M, Fraser, C, Ramakrishnan, V. | | Deposit date: | 2020-03-17 | | Release date: | 2020-09-16 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (6 Å) | | Cite: | Structure of a human 48Stranslational initiation complex.
Science, 369, 2020
|
|
8BI7
 
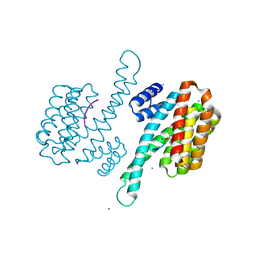 | | Binary structure of 14-3-3s and PKR phosphopeptide | | Descriptor: | 14-3-3 protein sigma, MAGNESIUM ION, PKR phosphopeptide | | Authors: | Somsen, B.A, Ottmann, C. | | Deposit date: | 2022-11-01 | | Release date: | 2023-03-29 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Reversible Dual-Covalent Molecular Locking of the 14-3-3/ERR gamma Protein-Protein Interaction as a Molecular Glue Drug Discovery Approach.
J.Am.Chem.Soc., 145, 2023
|
|
4MHT
 
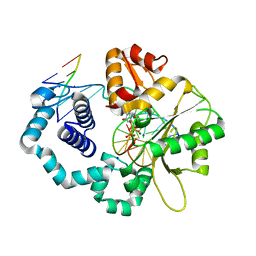
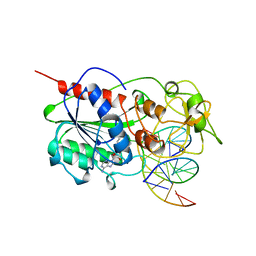 | | TERNARY STRUCTURE OF HHAI METHYLTRANSFERASE WITH NATIVE DNA AND ADOHCY | | Descriptor: | DNA (5'-D(*GP*AP*TP*AP*GP*(5CM)P*GP*CP*TP*AP*TP*C)-3'), DNA (5'-D(*TP*GP*AP*TP*AP*GP*(5CM)P*GP*CP*TP*AP*TP*C)-3'), PROTEIN (HHAI METHYLTRANSFERASE (E.C.2.1.1.73)), ... | | Authors: | Cheng, X. | | Deposit date: | 1996-07-24 | | Release date: | 1997-01-27 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Enzymatic C5-cytosine methylation of DNA: mechanistic implications of new crystal structures for HhaL methyltransferase-DNA-AdoHcy complexes.
J.Mol.Biol., 261, 1996
|
|
4M8O
 
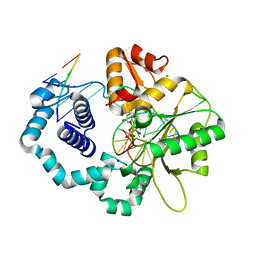
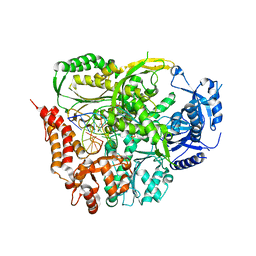 | | TERNARY COMPLEX OF DNA POLYMERASE EPSILON WITH AN INCOMING dATP | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, 2-AMINOETHANESULFONIC ACID, CHLORIDE ION, ... | | Authors: | Sauer-Eriksson, A.E, Hogg, M, Osterman, P, Johansson, E. | | Deposit date: | 2013-08-13 | | Release date: | 2013-11-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural basis for processive DNA synthesis by yeast DNA polymerase epsilon.
Nat.Struct.Mol.Biol., 21, 2014
|
|
6NKX
 
 | |
6NKR
 
 | | Ternary complex crystal structure of K289M variant of DNA polymerase Beta with dGTP | | Descriptor: | 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, CHLORIDE ION, DNA (5'-D(*CP*CP*GP*AP*CP*CP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3'), ... | | Authors: | Batra, V.K, Wilson, S.H. | | Deposit date: | 2019-01-07 | | Release date: | 2020-01-08 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Revealing an Internal Stabilization Deficiency in the DNA Polymerase beta K289M Cancer Variant through the Combined Use of Chemical Biology and X-ray Crystallography.
Biochemistry, 59, 2020
|
|
6NKU
 
 | |
6NL0
 
 | |
6NKS
 
 | |
6NKW
 
 | |
6CR6
 
 | | Ternary complex crystal structure of DNA polymerase Beta with a dideoxy terminated primer with CH-CH3, beta, gamma dATP analogue | | Descriptor: | 2'-deoxy-5'-O-[(R)-hydroxy({(R)-hydroxy[(1S)-1-phosphonoethyl]phosphoryl}oxy)phosphoryl]adenosine, DNA polymerase beta, Downstream Primer Strand, ... | | Authors: | Batra, V.K, Wilson, S.H. | | Deposit date: | 2018-03-16 | | Release date: | 2018-08-22 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.097 Å) | | Cite: | Mapping Functional Substrate-Enzyme Interactions in the pol beta Active Site through Chemical Biology: Structural Responses to Acidity Modification of Incoming dNTPs.
Biochemistry, 57, 2018
|
|
6CTN
 
 | |
6CR9
 
 | | Ternary complex crystal structure of DNA polymerase Beta with a dideoxy terminated primer with CFCL, beta, gamma dATP analogue | | Descriptor: | 9-{5-O-[(R)-{[(R)-[(S)-chloro(fluoro)phosphonomethyl](hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]-2-deoxy-alpha-D-threo-pentofuranosyl}-9H-purin-6-amine, CHLORIDE ION, DNA polymerase beta, ... | | Authors: | Batra, V.K, Wilson, S.H. | | Deposit date: | 2018-03-16 | | Release date: | 2018-07-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Mapping Functional Substrate-Enzyme Interactions in the pol beta Active Site through Chemical Biology: Structural Responses to Acidity Modification of Incoming dNTPs.
Biochemistry, 57, 2018
|
|
