2HH4
 
 | | NMR structure of human insulin mutant GLY-B8-D-SER, HIS-B10-ASP PRO-B28-LYS, LYS-B29-PRO, 20 structures | | Descriptor: | insulin A chain, insulin B chain | | Authors: | Hua, Q.X, Nakagawa, S, Hu, S.Q, Jia, W, Weiss, M.A. | | Deposit date: | 2006-06-27 | | Release date: | 2006-07-18 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Toward the Active Conformation of Insulin: Stereospecific modulation of a structural switch in the B chain.
J.Biol.Chem., 281, 2006
|
|
2HHO
 
 | | NMR structure of human insulin mutant GLY-B8-SER, HIS-B10-ASP PRO-B28-LYS, LYS-B29-PRO, 20 structures | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Hua, Q.X, Nakagawa, S, Hu, S.Q, Jia, W, Weiss, M.A. | | Deposit date: | 2006-06-28 | | Release date: | 2006-07-18 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Toward the Active Conformation of Insulin: Stereospecific modulation of a structural switch in the B chain.
J.Biol.Chem., 281, 2006
|
|
2K3A
 
 | | NMR solution structure of Staphylococcus saprophyticus CHAP (cysteine, histidine-dependent amidohydrolases/peptidases) domain protein. Northeast Structural Genomics Consortium target SyR11 | | Descriptor: | CHAP domain protein | | Authors: | Rossi, P, Aramini, J.M, Chen, C.X, Nwosu, C, Cunningham, K.C, Owens, L.A, Xiao, R, Liu, J, Baran, M.C, Swapna, G, Acton, T.B, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-04-29 | | Release date: | 2008-05-13 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural elucidation of the Cys-His-Glu-Asn proteolytic relay in the secreted CHAP domain enzyme from the human pathogen Staphylococcus saprophyticus.
Proteins, 74, 2008
|
|
2JXD
 
 | |
1XOX
 
 | | SOLUTION STRUCTURE OF HUMAN SURVIVIN | | Descriptor: | Apoptosis inhibitor survivin, ZINC ION | | Authors: | Sun, C, Nettesheim, D, Liu, Z, Olejniczak, E.T. | | Deposit date: | 2004-10-07 | | Release date: | 2005-01-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of human survivin and its binding interface with smac/diablo
Biochemistry, 44, 2005
|
|
2KQQ
 
 | | NMR structure of human insulin mutant gly-b8-d-ala, his-b10-asp, pro-b28-lys, lys-b29-pro, 20 structures | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Hua, Q.X, Wan, Z.L, Zhao, M, Jia, W, Huang, K, Weiss, M.A. | | Deposit date: | 2009-11-13 | | Release date: | 2010-11-24 | | Last modified: | 2021-10-13 | | Method: | SOLUTION NMR | | Cite: | Structure and Dynamics of an Turn-Stabilize But Ina Insulin Analog.
To be Published
|
|
2KV7
 
 | | NMR solution structure of a soluble PrgI mutant from Salmonella Typhimurium | | Descriptor: | Protein prgI | | Authors: | Schmidt, H, Poyraz, O, Seidel, K, Delissen, F, Ader, C, Tenenboim, H, Goosmann, C, Laube, B, Thuenemann, A.F, Zychlinski, A, Baldus, M, Lange, A, Griesinger, C, Kolbe, M. | | Deposit date: | 2010-03-09 | | Release date: | 2010-06-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Protein refolding is required for assembly of the type three secretion needle.
Nat.Struct.Mol.Biol., 17, 2010
|
|
2IMU
 
 | | NMR structure of pep46 from the infectious bursal disease virus (IBDV) in dodecylphosphocholin (DPC). | | Descriptor: | Structural polyprotein (PP) p1 | | Authors: | Galloux, M, Libersou, S, Morellet, N, Bouaziz, S, Ouldali, M, Da Costa, B, Lepault, J, Delmas, B. | | Deposit date: | 2006-10-05 | | Release date: | 2007-06-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Cell entry of a dsRNA virus through pores induced by one of its structural peptides.
To be Published
|
|
2I2H
 
 | | NMR structure of TPC3 in TFE | | Descriptor: | signaling peptide TCP3 | | Authors: | Syvitski, R.T, Jakeman, D.L, Li, Y. | | Deposit date: | 2006-08-16 | | Release date: | 2006-10-17 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structure-Activity Analysis of Quorum-Sensing Signaling Peptides from Streptococcus mutans.
J.Bacteriol., 189, 2007
|
|
2KF0
 
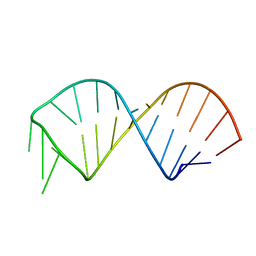 | | NMR structure of U6 ISL at pH 7.0 | | Descriptor: | RNA (5'-R(*GP*GP*UP*UP*CP*CP*CP*CP*UP*GP*CP*AP*UP*AP*AP*GP*GP*AP*UP*GP*AP*AP*CP*C)-3') | | Authors: | Venditti, V, Butcher, S.E. | | Deposit date: | 2009-02-08 | | Release date: | 2009-07-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Minimum-energy path for a u6 RNA conformational change involving protonation, base-pair rearrangement and base flipping.
J.Mol.Biol., 391, 2009
|
|
2I2J
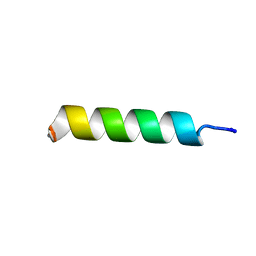 
 | | NMR structure of UA159sp in TFE | | Descriptor: | Competence stimulating peptide | | Authors: | Syvitski, R.T, Jakeman, D.L, Li, Y. | | Deposit date: | 2006-08-16 | | Release date: | 2006-10-17 | | Last modified: | 2020-03-04 | | Method: | SOLUTION NMR | | Cite: | Structure-Activity Analysis of Quorum-Sensing Signaling Peptides from Streptococcus mutans.
J.Bacteriol., 189, 2007
|
|
2GL1
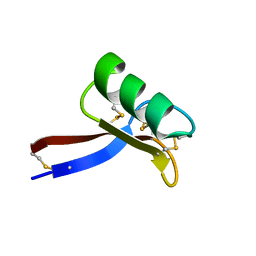 
 | | NMR solution structure of Vigna radiata Defensin 2 (VrD2) | | Descriptor: | PDF1 | | Authors: | Lin, K.F, Lee, T.R, Tsai, P.H, Hsu, M.P, Chen, C.S, Lyu, P.C. | | Deposit date: | 2006-04-04 | | Release date: | 2007-04-03 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structure-based protein engineering for alpha-amylase inhibitory activity of plant defensin.
Proteins, 68, 2007
|
|
2JPI
 
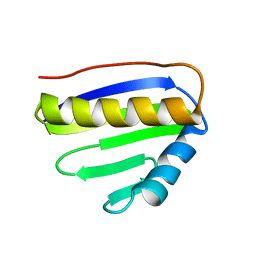 | | NMR structure of PA4090 from Pseudomonas aeruginosa | | Descriptor: | Hypothetical protein | | Authors: | Ai, X, Semesi, A, Yee, A, Arrowsmith, C.H, Li, S.S.C, Choy, W, Ontario Centre for Structural Proteomics (OCSP) | | Deposit date: | 2007-05-13 | | Release date: | 2007-10-16 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | chemical shift assignments of PA4090 from Pseudomonas aeruginosa
To be Published
|
|
2K65
 
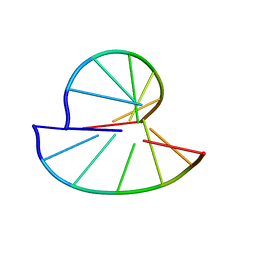 | |
2K7H
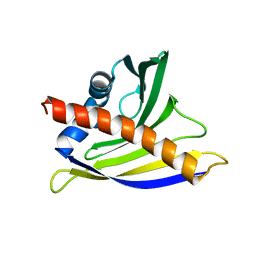 
 | | NMR solution structure of soybean allergen Gly m 4 | | Descriptor: | Stress-induced protein SAM22 | | Authors: | Berkner, H, Neudecker, P, Mittag, D, Ballmer-Weber, B.K, Schweimer, K, Vieths, S, Roesch, P. | | Deposit date: | 2008-08-11 | | Release date: | 2009-05-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Cross-reactivity of pollen and food allergens: soybean Gly m 4 is a member of the Bet v 1 superfamily and closely resembles yellow lupine proteins
Biosci.Rep., 29, 2009
|
|
2KGL
 
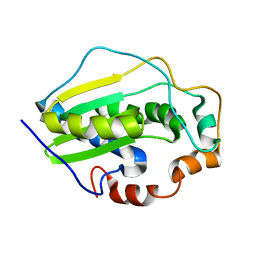 | | NMR solution structure of MESD | | Descriptor: | Mesoderm development candidate 2 | | Authors: | Chen, J, Wang, J. | | Deposit date: | 2009-03-12 | | Release date: | 2010-03-23 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Two Structural and Functional Domains of MESD Required for Proper Folding and Trafficking of LRP5/6.
Structure, 19, 2011
|
|
2KC1
 
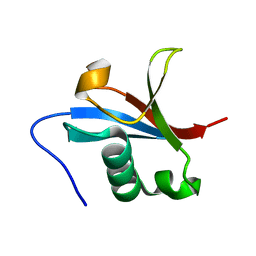 | | NMR structure of the F0 domain (residues 0-85) of the talin ferm domain | | Descriptor: | MKIAA1027 protein | | Authors: | Goult, B.T, Elliott, P.R, Roberts, G.C.K, Critchley, D.R, Barsukov, I.L. | | Deposit date: | 2008-12-13 | | Release date: | 2010-01-19 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure of a double ubiquitin-like domain in the talin head: a role in integrin activation.
Embo J., 29, 2010
|
|
2KMA
 
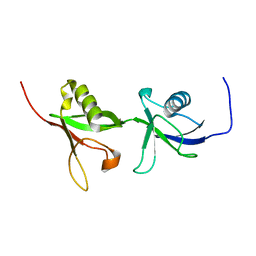 | | NMR structure of the F0F1 double domain (residues 1-202) of the talin ferm domain | | Descriptor: | Talin 1 | | Authors: | Goult, B.T, Elliott, P.R, Bate, N, Roberts, G.C, Critchley, D.R, Barsukov, I.L. | | Deposit date: | 2009-07-25 | | Release date: | 2010-03-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure of a double ubiquitin-like domain in the talin head: a role in integrin activation.
Embo J., 2010
|
|
2LA9
 
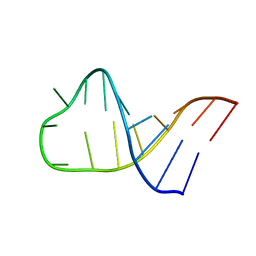 | | NMR structure of Pseudouridine_ASL_Tyr | | Descriptor: | RNA (5'-R(*GP*GP*GP*GP*AP*CP*UP*GP*UP*AP*AP*AP*(PSU)P*CP*CP*CP*C)-3') | | Authors: | Denmon, A.P, Wang, J, Nikonowicz, E.P. | | Deposit date: | 2011-03-09 | | Release date: | 2011-08-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Conformation Effects of Base Modification on the Anticodon Stem-Loop of Bacillus subtilis tRNA(Tyr).
J.Mol.Biol., 412, 2011
|
|
2JMN
 
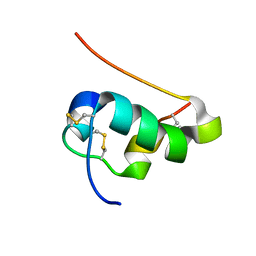 | | NMR structure of human insulin mutant His-B10-Asp, Pro-B28-Lys, Lys-B29-Pro, 20 structures | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Hua, Q.X, Hu, S.Q, Frank, B.H, Jia, W.H, Chu, Y.C, Wang, S.H, Burke, G.T, Katsoyannis, P.G, Weiss, M.A. | | Deposit date: | 2006-11-21 | | Release date: | 2006-12-05 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | Mapping the functional surface of insulin by design: structure and function of a novel A-chain analogue.
J.Mol.Biol., 264, 1996
|
|
2LTH
 
 | | NMR structure of major ampullate spidroin 1 N-terminal domain at pH 5.5 | | Descriptor: | Major ampullate spidroin 1 | | Authors: | Otikovs, M, Jaudzems, K, Nordling, K, Landreh, M, Rising, A, Askarieh, G, Knight, S, Johansson, J. | | Deposit date: | 2012-05-25 | | Release date: | 2013-11-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Sequential pH-driven dimerization and stabilization of the N-terminal domain enables rapid spider silk formation.
Nat Commun, 5, 2014
|
|
2LAC
 
 | | NMR structure of unmodified_ASL_Tyr | | Descriptor: | RNA (5'-R(*GP*GP*GP*GP*AP*CP*UP*GP*UP*AP*AP*AP*UP*CP*CP*CP*C)-3') | | Authors: | Denmon, A.P, Wang, J, Nikonowicz, E.P. | | Deposit date: | 2011-03-10 | | Release date: | 2011-08-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Conformation Effects of Base Modification on the Anticodon Stem-Loop of Bacillus subtilis tRNA(Tyr).
J.Mol.Biol., 412, 2011
|
|
2JYA
 
 | | NMR solution structure of protein ATU1810 from Agrobacterium tumefaciens. Northeast Structural Genomics Consortium target AtR23, Ontario Centre for Structural Proteomics Target ATC1776 | | Descriptor: | Uncharacterized protein Atu1810 | | Authors: | Lemak, A, Yee, A, Gutmanas, A, Fares, C, Karra, M, Semesi, A, Arrowsmith, C.H, Northeast Structural Genomics Consortium (NESG), Ontario Centre for Structural Proteomics (OCSP) | | Deposit date: | 2007-12-07 | | Release date: | 2007-12-25 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of protein ATU1810 form Agrobacterium tumefaciens.
To be Published
|
|
2JYQ
 
 | | NMR structure of the apo v-Src SH2 domain | | Descriptor: | Tyrosine-protein kinase transforming protein Src | | Authors: | Taylor, J.D, Ababou, A, Williams, M.A, Ladbury, J.E. | | Deposit date: | 2007-12-17 | | Release date: | 2008-06-24 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure, dynamics, and binding thermodynamics of the v-Src SH2 domain: Implications for drug design
Proteins, 73, 2008
|
|
2C0S
 
 | | NMR Solution Structure of a protein aspartic acid phosphate phosphatase from Bacillus Anthracis | | Descriptor: | CONSERVED DOMAIN PROTEIN | | Authors: | Grenha, R, Rzechorzek, N.J, Brannigan, J.A, Ab, E, Folkers, G.E, De Jong, R.N, Diercks, T, Wilkinson, A.J, Kaptein, R, Wilson, K.S. | | Deposit date: | 2005-09-07 | | Release date: | 2006-09-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of Spo0E-like protein-aspartic acid phosphatases that regulate sporulation in bacilli.
J. Biol. Chem., 281, 2006
|
|
