1BVC
 
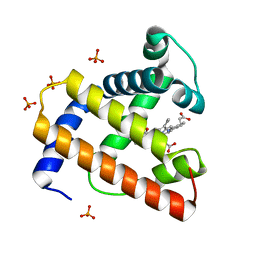 | | STRUCTURE OF A BILIVERDIN APOMYOGLOBIN COMPLEX (FORM D) AT 118 K | | Descriptor: | APOMYOGLOBIN, BILIVERDINE IX ALPHA, PHOSPHATE ION | | Authors: | Wagner, U.G, Mueller, N, Schmitzberger, W, Falk, H, Kratky, C. | | Deposit date: | 1994-12-16 | | Release date: | 1995-07-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure determination of the biliverdin apomyoglobin complex: crystal structure analysis of two crystal forms at 1.4 and 1.5 A resolution.
J.Mol.Biol., 247, 1995
|
|
5I9J
 
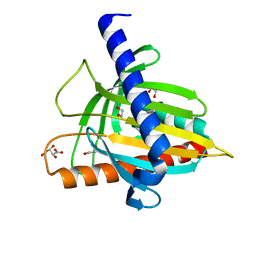 | | Structure of the cholesterol and lutein-binding domain of human STARD3 at 1.74A | | Descriptor: | 1,2-ETHANEDIOL, L(+)-TARTARIC ACID, SULFATE ION, ... | | Authors: | Horvath, M.P, Bernstein, P.S, Li, B, George, E.W, Tran, Q.T. | | Deposit date: | 2016-02-20 | | Release date: | 2016-03-16 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Structure of the lutein-binding domain of human StARD3 at 1.74 angstrom resolution and model of a complex with lutein.
Acta Crystallogr.,Sect.F, 72, 2016
|
|
3IKD
 
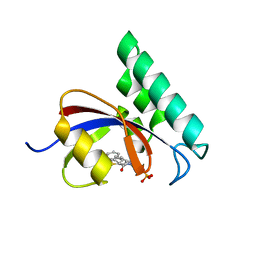 | | Structure-Based Design of Novel PIN1 Inhibitors (I) | | Descriptor: | (2R)-2-[(1-benzothiophen-2-ylcarbonyl)amino]-3-phenylpropyl phosphate, Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 | | Authors: | Matthews, D, Greasley, S, Ferre, R, Parge, H. | | Deposit date: | 2009-08-05 | | Release date: | 2009-09-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure-based design of novel human Pin1 inhibitors (I).
Bioorg.Med.Chem.Lett., 19, 2009
|
|
6KM0
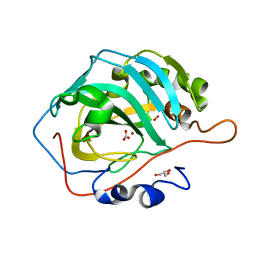 
 | | Human Carbonic Anhydrase II V143I variant 07 atm CO2 | | Descriptor: | BICARBONATE ION, CARBON DIOXIDE, Carbonic anhydrase 2, ... | | Authors: | Kim, C.U, Kim, J.K. | | Deposit date: | 2019-07-30 | | Release date: | 2020-08-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (0.93 Å) | | Cite: | Structural insights into the effect of active-site mutation on the catalytic mechanism of carbonic anhydrase.
Iucrj, 7, 2020
|
|
3IND
 
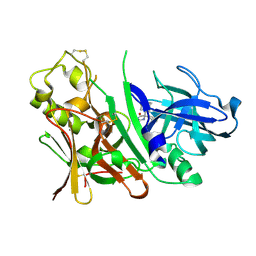 | | Bace1 with the aminohydantoin Compound 29 | | Descriptor: | (5S)-2-amino-3-methyl-5-phenyl-5-[(3S,5S,7S)-tricyclo[3.3.1.1~3,7~]dec-1-yl]-3,5-dihydro-4H-imidazol-4-one, Beta-secretase 1 | | Authors: | Olland, A.M, Chopra, R. | | Deposit date: | 2009-08-12 | | Release date: | 2010-03-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.246 Å) | | Cite: | Design and synthesis of 5,5'-disubstituted aminohydantoins as potent and selective human beta-secretase (BACE1) inhibitors.
J.Med.Chem., 53, 2010
|
|
7KIZ
 
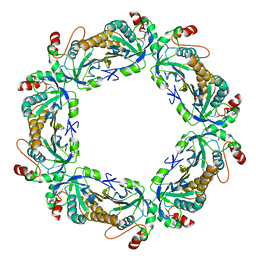 | | reduced human peroxiredoxin 2 | | Descriptor: | Peroxiredoxin-2 | | Authors: | Kean, K.M, Karplus, P.A. | | Deposit date: | 2020-10-25 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Modifying the resolving cysteine affects the structure and hydrogen peroxide reactivity of peroxiredoxin 2.
J.Biol.Chem., 296, 2021
|
|
7KJ0
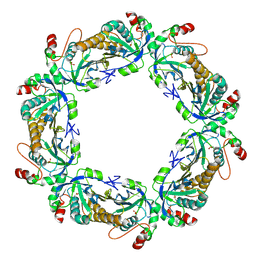 
 | | hyperoxidized human peroxiredoxin 2 | | Descriptor: | Peroxiredoxin-2 | | Authors: | Kean, K.M, Karplus, P.A. | | Deposit date: | 2020-10-25 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Modifying the resolving cysteine affects the structure and hydrogen peroxide reactivity of peroxiredoxin 2.
J.Biol.Chem., 296, 2021
|
|
3EI4
 
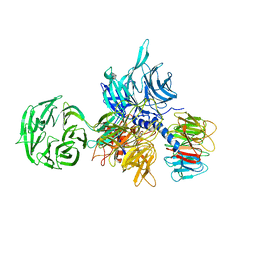 | | Structure of the hsDDB1-hsDDB2 complex | | Descriptor: | DNA damage-binding protein 1, DNA damage-binding protein 2 | | Authors: | Scrima, A, Pavletich, N.P, Thoma, N.H. | | Deposit date: | 2008-09-15 | | Release date: | 2009-01-20 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structural basis of UV DNA-damage recognition by the DDB1-DDB2 complex.
Cell(Cambridge,Mass.), 135, 2008
|
|
7KJ1
 
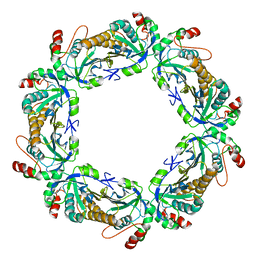 | | human peroxiredoxin 2 - C172S mutant | | Descriptor: | Peroxiredoxin-2 | | Authors: | Kean, K.M, Karplus, P.A. | | Deposit date: | 2020-10-25 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Modifying the resolving cysteine affects the structure and hydrogen peroxide reactivity of peroxiredoxin 2.
J.Biol.Chem., 296, 2021
|
|
3EJW
 
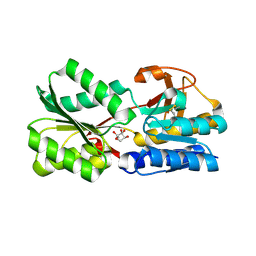 | | Crystal Structure of the Sinorhizobium meliloti AI-2 receptor, SmLsrB | | Descriptor: | (2R,4S)-2-methyl-2,3,3,4-tetrahydroxytetrahydrofuran, SmLsrB | | Authors: | Miller, S.T, McAuley, J.R, Pereira, C, Xavier, K.B, Taga, M.E. | | Deposit date: | 2008-09-18 | | Release date: | 2008-12-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Sinorhizobium meliloti, a bacterium lacking the autoinducer-2 (AI-2) synthase, responds to AI-2 supplied by other bacteria.
Mol.Microbiol., 70, 2008
|
|
1K02
 
 | | Crystal Structure of Old Yellow Enzyme Mutant Gln114Asn | | Descriptor: | FLAVIN MONONUCLEOTIDE, MAGNESIUM ION, NADPH DEHYDROGENASE 1 | | Authors: | Brown, B.J, Hyun, J, Duvvuri, S.D, Karplus, P.A, Massey, V. | | Deposit date: | 2001-09-18 | | Release date: | 2001-09-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The role of glutamine 114 in old yellow enzyme
J.Biol.Chem., 277, 2002
|
|
3INE
 
 | | Bace1 with the aminohydantoin Compound S-34 | | Descriptor: | (5S)-2-amino-5-(4-methoxy-3-methylphenyl)-3-methyl-5-[(3S,5S,7S)-tricyclo[3.3.1.1~3,7~]dec-1-yl]-3,5-dihydro-4H-imidazol-4-one, Beta-secretase 1 | | Authors: | Olland, A.M, Chopra, R. | | Deposit date: | 2009-08-12 | | Release date: | 2010-03-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.996 Å) | | Cite: | Design and synthesis of 5,5'-disubstituted aminohydantoins as potent and selective human beta-secretase (BACE1) inhibitors.
J.Med.Chem., 53, 2010
|
|
4PC0
 
 | | Structure of the human RbAp48-MTA1(670-711) complex | | Descriptor: | CALCIUM ION, GLYCEROL, Histone-binding protein RBBP4, ... | | Authors: | Alqarni, S.S.M, Silva, A.P.G, Mackay, J.P, Laue, E.D. | | Deposit date: | 2014-04-14 | | Release date: | 2014-06-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Insight into the architecture of the NuRD complex: Structure of the RbAp48-MTA1 sub-complex.
J.Biol.Chem., 289, 2014
|
|
6GZE
 
 | | Tubulin-GDP.BeF complex | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, BERYLLIUM TRIFLUORIDE ION, CALCIUM ION, ... | | Authors: | Oliva, M.A, Diaz, J.F. | | Deposit date: | 2018-07-04 | | Release date: | 2020-01-22 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Structural model for differential cap maturation at growing microtubule ends.
Elife, 9, 2020
|
|
3EU0
 
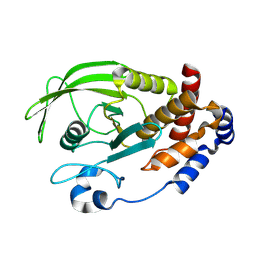 | | Crystal structure of the S-nitrosylated Cys215 of PTP1B | | Descriptor: | Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Chu, H.M, Wang, A.H.J, Chen, Y.Y, Pan, K.T, Wang, D.L, Khoo, K.H, Meng, T.C. | | Deposit date: | 2008-10-09 | | Release date: | 2008-11-11 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Cysteine S-Nitrosylation Protects Protein-tyrosine Phosphatase 1B against Oxidation-induced Permanent Inactivation
J.Biol.Chem., 283, 2008
|
|
3E78
 
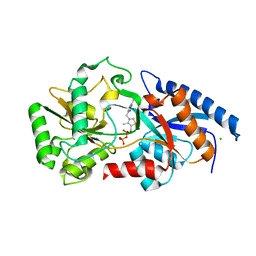 | | Structure determination of the cancer-associated Mycoplasma hyorhinis protein Mh-p37 | | Descriptor: | CALCIUM ION, CHLORIDE ION, High affinity transport system protein p37, ... | | Authors: | Sippel, K.H, Robbins, A.H, Reutzel, R, Domsic, J, McKenna, R. | | Deposit date: | 2008-08-18 | | Release date: | 2008-10-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure determination of the cancer-associated Mycoplasma hyorhinis protein Mh-p37.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
1CHL
 
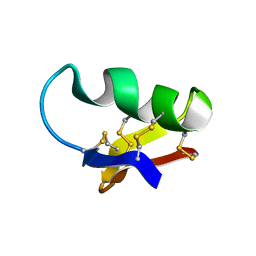 | | NMR SEQUENTIAL ASSIGNMENTS AND SOLUTION STRUCTURE OF CHLOROTOXIN, A SMALL SCORPION TOXIN THAT BLOCKS CHLORIDE CHANNELS | | Descriptor: | CHLOROTOXIN | | Authors: | Lippens, G, Najib, J, Wodak, S.J, Tartar, A. | | Deposit date: | 1994-11-09 | | Release date: | 1995-02-07 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | NMR sequential assignments and solution structure of chlorotoxin, a small scorpion toxin that blocks chloride channels.
Biochemistry, 34, 1995
|
|
3EMP
 
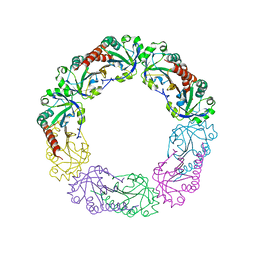 | |
3INF
 
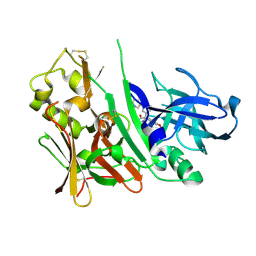 | | Bace1 with the aminohydantoin Compound 37 | | Descriptor: | (5S)-2-amino-5-(4-methoxy-3-methylphenyl)-3-methyl-5-(3-pyridin-3-ylphenyl)-3,5-dihydro-4H-imidazol-4-one, Beta-secretase 1 | | Authors: | Olland, A.M, Chopra, R. | | Deposit date: | 2009-08-12 | | Release date: | 2010-03-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.852 Å) | | Cite: | Design and synthesis of 5,5'-disubstituted aminohydantoins as potent and selective human beta-secretase (BACE1) inhibitors.
J.Med.Chem., 53, 2010
|
|
2J8N
 
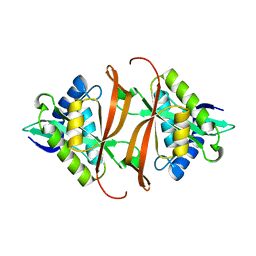 | | Structure of P. aeruginosa acetyltransferase PA4866 solved at room temperature | | Descriptor: | ACETYLTRANSFERASE PA4866 FROM P. AERUGINOSA | | Authors: | Davies, A.M, Tata, R, Beavil, R.L, Sutton, B.J, Brown, P.R. | | Deposit date: | 2006-10-26 | | Release date: | 2007-02-06 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | l-Methionine sulfoximine, but not phosphinothricin, is a substrate for an acetyltransferase (gene PA4866) from Pseudomonas aeruginosa: structural and functional studies.
Biochemistry, 46, 2007
|
|
3IK8
 
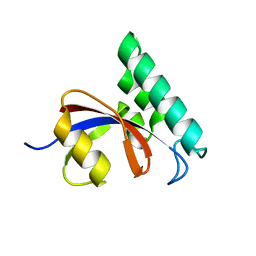 | |
7W7J
 
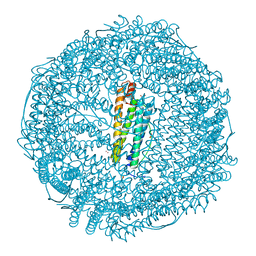 | | Crystal structure of IrCp* immobilized apo-R52H-rHLFr (25 equiv) | | Descriptor: | 1,2-ETHANEDIOL, CADMIUM ION, CHLORIDE ION, ... | | Authors: | Taher, M, Maity, B, Nakane, T, Abe, S, Mazumdar, S. | | Deposit date: | 2021-12-05 | | Release date: | 2022-03-02 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Controlled Uptake of an Iridium Complex inside Engineered apo-Ferritin Nanocages: Study of Structure and Catalysis.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
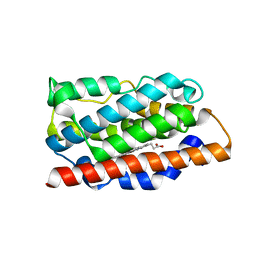
4MEC
 
 | |
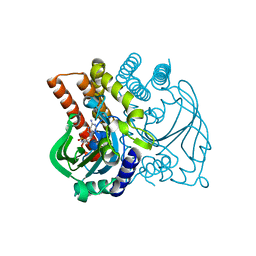
5F5W
 
 | |
3R7L
 
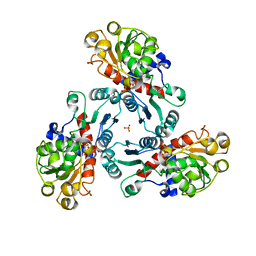 | | Crystal Structure of PALA-bound Aspartate Transcarbamoylase from Bacillus subtilis | | Descriptor: | Aspartate carbamoyltransferase, N-(PHOSPHONACETYL)-L-ASPARTIC ACID, PHOSPHATE ION | | Authors: | Harris, K.M, Cockrell, G.M, Puleo, D.E, Kantrowitz, E.R. | | Deposit date: | 2011-03-22 | | Release date: | 2011-06-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.585 Å) | | Cite: | Crystallographic Snapshots of the Complete Catalytic Cycle of the Unregulated Aspartate Transcarbamoylase from Bacillus subtilis.
J.Mol.Biol., 411, 2011
|
|
