4ZBX
 
 | |
4ZBY
 
 | |
4ZBZ
 
 | |
5AYR
 
 | | The crystal structure of SAUGI/human UDG complex | | 分子名称: | MAGNESIUM ION, Uncharacterized protein, Uracil-DNA glycosylase | | 著者 | Wang, H.C, Ko, T.P, Huang, M.F, Wang, A.H.J. | | 登録日 | 2015-09-02 | | 公開日 | 2016-06-08 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Using structural-based protein engineering to modulate the differential inhibition effects of SAUGI on human and HSV uracil DNA glycosylase.
Nucleic Acids Res., 44, 2016
|
|
5AYS
 
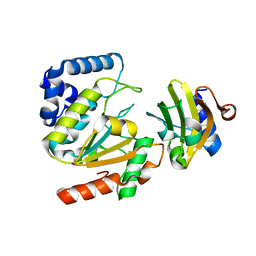 | | Crystal structure of SAUGI/HSV UDG complex | | 分子名称: | Uncharacterized protein, Uracil-DNA glycosylase | | 著者 | Wang, H.C, Ko, T.P, Huang, M.F, Wang, A.H.J. | | 登録日 | 2015-09-02 | | 公開日 | 2016-06-08 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.09 Å) | | 主引用文献 | Using structural-based protein engineering to modulate the differential inhibition effects of SAUGI on human and HSV uracil DNA glycosylase.
Nucleic Acids Res., 44, 2016
|
|
5CYS
 
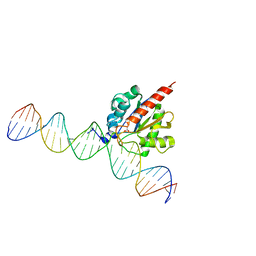 | |
5EUG
 
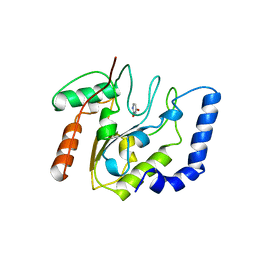 | | CRYSTALLOGRAPHIC AND ENZYMATIC STUDIES OF AN ACTIVE SITE VARIANT H187Q OF ESCHERICHIA COLI URACIL DNA GLYCOSYLASE: CRYSTAL STRUCTURES OF MUTANT H187Q AND ITS URACIL COMPLEX | | 分子名称: | PROTEIN (GLYCOSYLASE), URACIL | | 著者 | Xiao, G, Tordova, M, Drohat, A.C, Jagadeesh, J, Stivers, J.T, Gilliland, G.L. | | 登録日 | 1998-12-27 | | 公開日 | 1999-07-23 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Crystal structure of Escherichia coli uracil DNA glycosylase and its complexes with uracil and glycerol: structure and glycosylase mechanism revisited.
Proteins, 35, 1999
|
|
5FF8
 
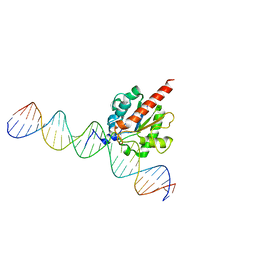 | | TDG enzyme-product complex | | 分子名称: | DNA, G/T mismatch-specific thymine DNA glycosylase | | 著者 | Pozharski, E, Malik, S.S, Drohat, A.C. | | 登録日 | 2015-12-18 | | 公開日 | 2016-09-28 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Structural basis of damage recognition by thymine DNA glycosylase: Key roles for N-terminal residues.
Nucleic Acids Res., 44, 2016
|
|
5H0J
 
 | |
5H0K
 
 | |
5H93
 
 | |
5H98
 
 | |
5H99
 
 | |
5H9I
 
 | | Crystal structure of Geobacter metallireducens SMUG1 with xanthine | | 分子名称: | BETA-MERCAPTOETHANOL, GLYCEROL, Geobacter metallireducens SMUG1, ... | | 著者 | Xie, W, Cao, W, Zhang, Z, Shen, J. | | 登録日 | 2015-12-28 | | 公開日 | 2016-04-27 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.501 Å) | | 主引用文献 | Structural Basis of Substrate Specificity in Geobacter metallireducens SMUG1
Acs Chem.Biol., 11, 2016
|
|
5HF7
 
 | | TDG enzyme-substrate complex | | 分子名称: | DNA (28-MER), G/T mismatch-specific thymine DNA glycosylase | | 著者 | Pozharski, E, Malik, S.S, Drohat, A.C. | | 登録日 | 2016-01-06 | | 公開日 | 2016-09-28 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (1.54 Å) | | 主引用文献 | Structural basis of damage recognition by thymine DNA glycosylase: Key roles for N-terminal residues.
Nucleic Acids Res., 44, 2016
|
|
5JK7
 
 | | The X-ray structure of the DDB1-DCAF1-Vpr-UNG2 complex | | 分子名称: | DNA damage-binding protein 1, Protein VPRBP, Protein Vpr, ... | | 著者 | Calero, G, Ahn, J, Wu, Y. | | 登録日 | 2016-04-26 | | 公開日 | 2016-10-05 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (3.49 Å) | | 主引用文献 | The DDB1-DCAF1-Vpr-UNG2 crystal structure reveals how HIV-1 Vpr steers human UNG2 toward destruction.
Nat.Struct.Mol.Biol., 23, 2016
|
|
5JXY
 
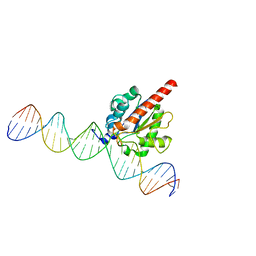 | | Enzyme-substrate complex of TDG catalytic domain bound to a G/U analog | | 分子名称: | DNA (28-MER), G/T mismatch-specific thymine DNA glycosylase | | 著者 | Pidugu, L.S, Pozharski, E, Malik, S.S, Drohat, A.C. | | 登録日 | 2016-05-13 | | 公開日 | 2016-09-28 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (1.71 Å) | | 主引用文献 | Structural basis of damage recognition by thymine DNA glycosylase: Key roles for N-terminal residues.
Nucleic Acids Res., 44, 2016
|
|
5NN7
 
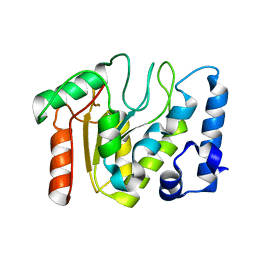 | | KSHV uracil-DNA glycosylase, apo form | | 分子名称: | Uracil-DNA glycosylase | | 著者 | Earl, C, Bagneris, C, Cole, A.R, Barrett, T, Savva, R. | | 登録日 | 2017-04-08 | | 公開日 | 2018-03-21 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | A structurally conserved motif in gamma-herpesvirus uracil-DNA glycosylases elicits duplex nucleotide-flipping.
Nucleic Acids Res., 46, 2018
|
|
5NNH
 
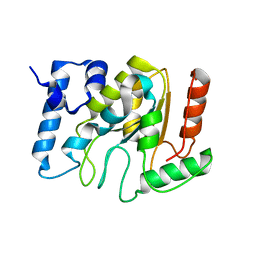 | | KSHV uracil-DNA glycosylase, apo form | | 分子名称: | SULFATE ION, Uracil-DNA glycosylase | | 著者 | Earl, C, Bagneris, C, Cole, A.R, Barrett, T, Savva, R. | | 登録日 | 2017-04-09 | | 公開日 | 2018-03-21 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | A structurally conserved motif in gamma-herpesvirus uracil-DNA glycosylases elicits duplex nucleotide-flipping.
Nucleic Acids Res., 46, 2018
|
|
5NNU
 
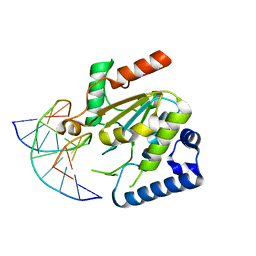 | | KSHV uracil-DNA glycosylase, product complex with dsDNA exhibiting duplex nucleotide flipping | | 分子名称: | DNA, DNA containing an abasic site, Uracil-DNA glycosylase | | 著者 | Earl, C, Bagneris, C, Barrett, T, Savva, R. | | 登録日 | 2017-04-10 | | 公開日 | 2018-03-21 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2.97 Å) | | 主引用文献 | A structurally conserved motif in gamma-herpesvirus uracil-DNA glycosylases elicits duplex nucleotide-flipping.
Nucleic Acids Res., 46, 2018
|
|
5T2W
 
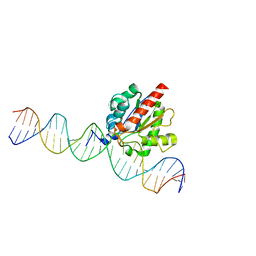 | |
5X55
 
 | |
6AIL
 
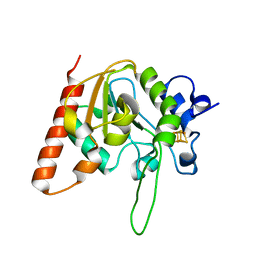 | | CRYSTAL STRUCTURE AT 1.3 ANGSTROMS RESOLUTION OF A NOVEL UDG, UdgX, FROM Mycobacterium smegmatis | | 分子名称: | IRON/SULFUR CLUSTER, Uracil DNA glycosylase X | | 著者 | Ahn, W.C, Aroli, S, Varshney, V, Woo, E.J. | | 登録日 | 2018-08-24 | | 公開日 | 2019-05-29 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (1.335 Å) | | 主引用文献 | Covalent binding of uracil DNA glycosylase UdgX to abasic DNA upon uracil excision.
Nat.Chem.Biol., 15, 2019
|
|
6AJO
 
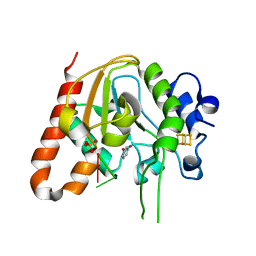 | | Complex form of Uracil DNA glycosylase X and uracil-DNA. | | 分子名称: | DNA (5'-D(P*(ORP)P*TP*T)-3'), IRON/SULFUR CLUSTER, PHOSPHATE ION, ... | | 著者 | Ahn, W.C, Aroli, S, Varshney, U, Woo, E.J. | | 登録日 | 2018-08-28 | | 公開日 | 2019-05-29 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.269 Å) | | 主引用文献 | Covalent binding of uracil DNA glycosylase UdgX to abasic DNA upon uracil excision.
Nat.Chem.Biol., 15, 2019
|
|
6AJP
 
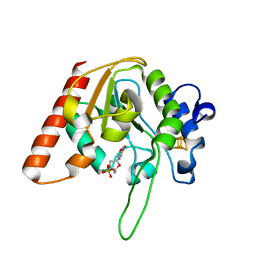 | | Complex form of Uracil DNA glycosylase X and deoxyuridine monophosphate. | | 分子名称: | 2'-DEOXYURIDINE-5'-MONOPHOSPHATE, IRON/SULFUR CLUSTER, Uracil DNA glycosylase superfamily protein | | 著者 | Ahn, W.C, Aroli, S, Varshney, U, Woo, E.J. | | 登録日 | 2018-08-28 | | 公開日 | 2019-05-29 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.334 Å) | | 主引用文献 | Covalent binding of uracil DNA glycosylase UdgX to abasic DNA upon uracil excision.
Nat.Chem.Biol., 15, 2019
|
|
