6B0D
 
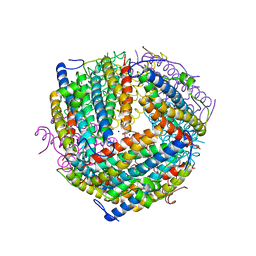 | | An E. coli DPS protein from ferritin superfamily | | 分子名称: | DNA protection during starvation protein, FORMIC ACID, SODIUM ION | | 著者 | Rui, W, Ruslan, S, Ronan, K, Adam, J.S. | | 登録日 | 2017-09-14 | | 公開日 | 2018-09-19 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | SIMBAD: a sequence-independent molecular-replacement pipeline.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
6AZH
 
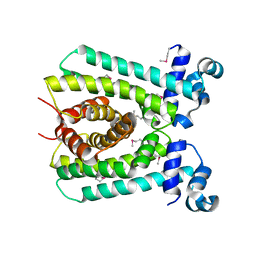 | |
4GIQ
 
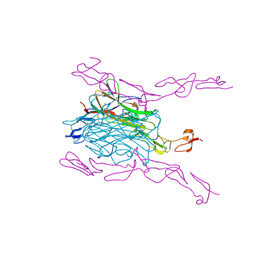 | | Crystal Structure of mouse RANK bound to RANKL | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, SODIUM ION, ... | | 著者 | Nelson, C.A, Wang, M.W.-H, Fremont, D.H. | | 登録日 | 2012-08-08 | | 公開日 | 2012-10-24 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | RANKL Employs Distinct Binding Modes to Engage RANK and the Osteoprotegerin Decoy Receptor.
Structure, 20, 2012
|
|
4A35
 
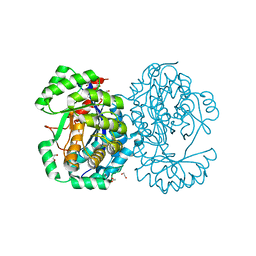 | | Crystal structure of human Mitochondrial enolase superfamily member 1 (ENOSF1) | | 分子名称: | 1,2-ETHANEDIOL, MAGNESIUM ION, MITOCHONDRIAL ENOLASE SUPERFAMILY MEMBER 1 | | 著者 | Muniz, J.R.C, Froese, D.S, Krojer, T, Vollmar, M, Canning, P, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Weigelt, J, Bountra, C, Oppermann, U, Yue, W.W. | | 登録日 | 2011-09-30 | | 公開日 | 2011-10-12 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (1.74 Å) | | 主引用文献 | Enzymatic and structural characterization of rTS gamma provides insights into the function of rTS beta.
Biochemistry, 53, 2014
|
|
4E4D
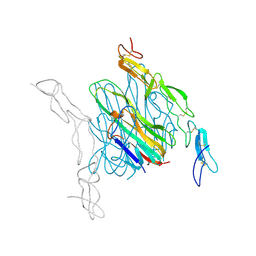 
 | | Crystal structure of mouse RANKL-OPG complex | | 分子名称: | CHLORIDE ION, Tumor necrosis factor ligand superfamily member 11, soluble form, ... | | 著者 | Nelson, C.A, Fremont, D.H. | | 登録日 | 2012-03-12 | | 公開日 | 2012-10-24 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | RANKL Employs Distinct Binding Modes to Engage RANK and the Osteoprotegerin Decoy Receptor.
Structure, 20, 2012
|
|
6D3N
 
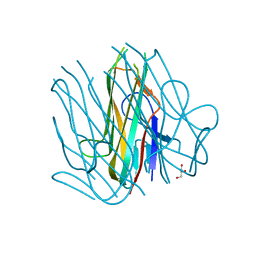 | | Crystal structure of h4-1BB ligand | | 分子名称: | GLYCEROL, Tumor necrosis factor ligand superfamily member 9 | | 著者 | Aruna, B, Zajonc, D.M, Doukov, T. | | 登録日 | 2018-04-16 | | 公開日 | 2018-05-09 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Crystal structures of the human 4-1BB receptor bound to its ligand 4-1BBL reveal covalent receptor dimerization as a potential signaling amplifier.
J. Biol. Chem., 293, 2018
|
|
6CHY
 
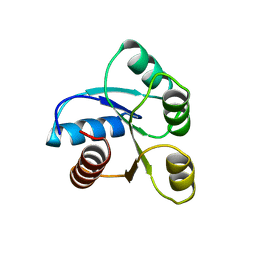 | | STRUCTURE OF CHEMOTAXIS PROTEIN CHEY | | 分子名称: | CHEY, SULFATE ION | | 著者 | Zhu, X, Rebello, J, Matsumura, P, Volz, K. | | 登録日 | 1996-08-29 | | 公開日 | 1996-12-07 | | 最終更新日 | 2024-05-22 | | 実験手法 | X-RAY DIFFRACTION (2.33 Å) | | 主引用文献 | Crystal structures of CheY mutants Y106W and T87I/Y106W. CheY activation correlates with movement of residue 106.
J.Biol.Chem., 272, 1997
|
|
4RSU
 
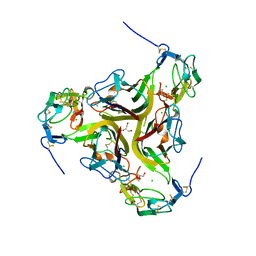 | | Crystal structure of the light and hvem complex | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, GLYCEROL, ... | | 著者 | Liu, W, Ramagoal, U.A, Himmel, D, Bonanno, J.B, Nathenson, S.G, Almo, S.C, Atoms-to-Animals: The Immune Function Network (IFN), New York Structural Genomics Research Consortium (NYSGRC) | | 登録日 | 2014-11-11 | | 公開日 | 2015-02-04 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | HVEM structures and mutants reveal distinct functions of binding to LIGHT and BTLA/CD160.
J.Exp.Med., 218, 2021
|
|
1JTZ
 
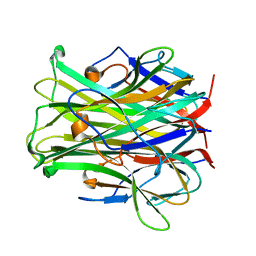 | |
5H6L
 
 | | DNA targeting ADP-ribosyltransferase Pierisin-1 in complex with beta-NAD+ | | 分子名称: | 1,2-ETHANEDIOL, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Pierisin-1 | | 著者 | Oda, T, Hirabayashi, H, Shikauchi, G, Takamura, R, Hiraga, K, Minami, H, Hashimoto, H, Yamamoto, M, Wakabayashi, K, Sugimura, T, Shimizu, T, Sato, M. | | 登録日 | 2016-11-14 | | 公開日 | 2017-08-09 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Structural basis of autoinhibition and activation of the DNA-targeting ADP-ribosyltransferase pierisin-1
J. Biol. Chem., 292, 2017
|
|
5H6M
 
 | | DNA targeting ADP-ribosyltransferase Pierisin-1 | | 分子名称: | 1,2-ETHANEDIOL, Pierisin-1 | | 著者 | Oda, T, Hirabayashi, H, Shikauchi, G, Takamura, R, Hiraga, K, Minami, H, Hashimoto, H, Yamamoto, M, Wakabayashi, K, Sugimura, T, Shimizu, T, Sato, M. | | 登録日 | 2016-11-14 | | 公開日 | 2017-08-09 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Structural basis of autoinhibition and activation of the DNA-targeting ADP-ribosyltransferase pierisin-1
J. Biol. Chem., 292, 2017
|
|
5DY6
 
 | |
6RTZ
 
 | |
1P94
 
 | | NMR Structure of ParG symmetric dimer | | 分子名称: | plasmid partition protein ParG | | 著者 | Golovanov, A.P, Barilla, D, Golovanova, M, Hayes, F, Lian, L.Y. | | 登録日 | 2003-05-09 | | 公開日 | 2004-01-13 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | ParG, a protein required for active partition of bacterial plasmids, has a dimeric ribbon-helix-helix structure.
Mol.Microbiol., 50, 2003
|
|
6RU0
 
 | |
8PNL
 
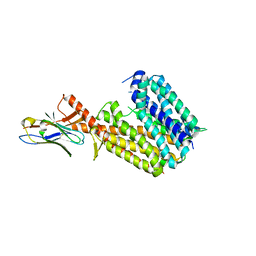 | | Outward-open conformation of a Major Facilitator Superfamily (MFS) transporter MHAS2168, a homologue of Rv1410 from M. tuberculosis, in complex with an alpaca nanobody | | 分子名称: | Nb_H2, Putative triacylglyceride transporter | | 著者 | Remm, S, Schoeppe, J, Hutter, C.A.J, Gonda, I, Seeger, M.A. | | 登録日 | 2023-06-30 | | 公開日 | 2023-10-18 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Structural basis for triacylglyceride extraction from mycobacterial inner membrane by MFS transporter Rv1410.
Nat Commun, 14, 2023
|
|
6XSX
 
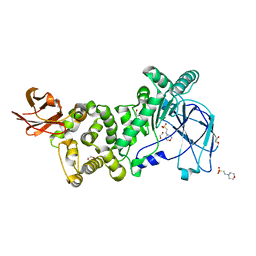 | |
6XT1
 
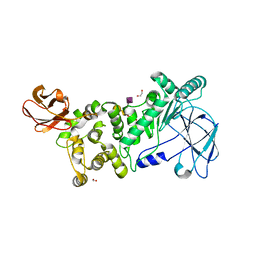 | |
6XNP
 
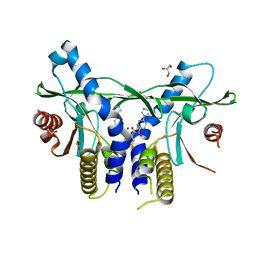 | | Crystal Structure of Human STING CTD complex with SR-717 | | 分子名称: | 1,2-ETHANEDIOL, 4,5-difluoro-2-{[6-(1H-imidazol-1-yl)pyridazine-3-carbonyl]amino}benzoic acid, GLYCEROL, ... | | 著者 | Chin, E.N, Yu, C, Wolan, D.W, Petrassi, H.M, Lairson, L.L. | | 登録日 | 2020-07-03 | | 公開日 | 2020-08-26 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.77 Å) | | 主引用文献 | Antitumor activity of a systemic STING-activating non-nucleotide cGAMP mimetic.
Science, 369, 2020
|
|
6SPZ
 
 | |
6XSZ
 
 | |
5H6J
 
 | | DNA targeting ADP-ribosyltransferase Pierisin-1 in complex with beta-NAD+ | | 分子名称: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Pierisin-1 | | 著者 | Oda, T, Hirabayashi, H, Shikauchi, G, Takamura, R, Hiraga, K, Minami, H, Hashimoto, H, Yamamoto, M, Wakabayashi, K, Sugimura, T, Shimizu, T, Sato, M. | | 登録日 | 2016-11-14 | | 公開日 | 2017-08-09 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Structural basis of autoinhibition and activation of the DNA-targeting ADP-ribosyltransferase pierisin-1
J. Biol. Chem., 292, 2017
|
|
6SPV
 
 | |
6XLP
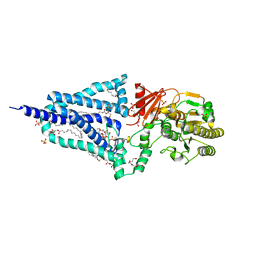 
 | | Structure of the essential inner membrane lipopolysaccharide-PbgA complex | | 分子名称: | (2S)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 1,2-Distearoyl-sn-glycerophosphoethanolamine, 2-deoxy-3-O-[(1R,3R)-1,3-dihydroxytetradecyl]-2-{[(3R)-3-hydroxytetradecanoyl]amino}-1-O-phosphono-alpha-D-glucopyranose-(6-1)-[3-deoxy-alpha-D-manno-oct-2-ulopyranosonic acid-(2-6)]1,5-anhydro-2-deoxy-2-{[(1S,3R)-1-hydroxy-3-(pentanoyloxy)undecyl]amino}-4-O-phosphono-D-glucitol, ... | | 著者 | Payandeh, J, Clairefeuille, T. | | 登録日 | 2020-06-29 | | 公開日 | 2020-08-26 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structure of the essential inner membrane lipopolysaccharide-PbgA complex.
Nature, 584, 2020
|
|
5H6K
 
 | | DNA targeting ADP-ribosyltransferase Pierisin-1 | | 分子名称: | 1,2-ETHANEDIOL, Pierisin-1 | | 著者 | Oda, T, Hirabayashi, H, Shikauchi, G, Takamura, R, Hiraga, K, Minami, H, Hashimoto, H, Yamamoto, M, Wakabayashi, K, Sugimura, T, Shimizu, T, Sato, M. | | 登録日 | 2016-11-14 | | 公開日 | 2017-08-09 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Structural basis of autoinhibition and activation of the DNA-targeting ADP-ribosyltransferase pierisin-1
J. Biol. Chem., 292, 2017
|
|
