9G35
 
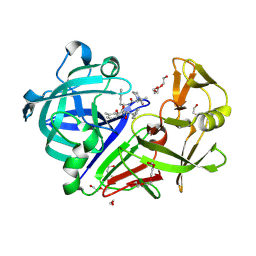 | | The HIV protease inhibitor lopinavir binding to the active site of Cryphonectria parasitica endothiapepsin | | Descriptor: | 1,2-ETHANEDIOL, Endothiapepsin, N-{1-BENZYL-4-[2-(2,6-DIMETHYL-PHENOXY)-ACETYLAMINO]-3-HYDROXY-5-PHENYL-PENTYL}-3-METHYL-2-(2-OXO-TETRAHYDRO-PYRIMIDIN-1-YL)-BUTYRAMIDE, ... | | Authors: | Falke, S, Senst, J.M, Guenther, S, Meents, A. | | Deposit date: | 2024-07-11 | | Release date: | 2024-07-24 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The HIV protease inhibitor lopinavir binding to the active site of Cryphonectria parasitica endothiapepsin
To Be Published
|
|
9G34
 
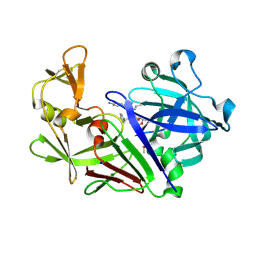 | | The HIV protease inhibitor darunavir binding to the active site of Cryphonectria parasitica endothiapepsin | | Descriptor: | (3R,3AS,6AR)-HEXAHYDROFURO[2,3-B]FURAN-3-YL(1S,2R)-3-[[(4-AMINOPHENYL)SULFONYL](ISOBUTYL)AMINO]-1-BENZYL-2-HYDROXYPROPYLCARBAMATE, 1,2-ETHANEDIOL, DIMETHYL SULFOXIDE, ... | | Authors: | Falke, S, Senst, J.M, Guenther, S, Meents, A. | | Deposit date: | 2024-07-11 | | Release date: | 2024-07-24 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The HIV protease inhibitor darunavir binding to the active site of Cryphonectria
parasitica endothiapepsin
To Be Published
|
|
9FVO
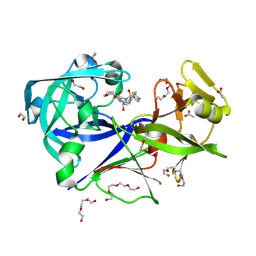 
 | | The HIV protease inhibitor amprenavir binding to the active site of Cryphonectria parasitica endothiapepsin | | Descriptor: | 1,2-ETHANEDIOL, ACETIC ACID, Endothiapepsin, ... | | Authors: | Falke, S, Senst, J.M, Guenther, S, Meents, A. | | Deposit date: | 2024-06-27 | | Release date: | 2024-07-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The HIV protease inhibitor amprenavir binding to the active site of Cryphonectria parasitica endothiapepsin
To Be Published
|
|
8TYH
 
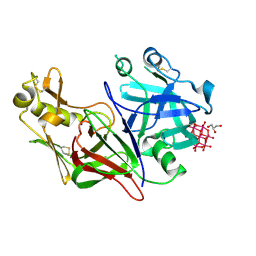 | | Plasmodium vivax PMX | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 6-tungstotellurate(VI), Aspartyl protease, ... | | Authors: | Hodder, A.N, Scally, S.W, Cowman, A.F. | | Deposit date: | 2023-08-25 | | Release date: | 2024-08-28 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Plasmodium vivax PMX
To Be Published
|
|
8TYG
 
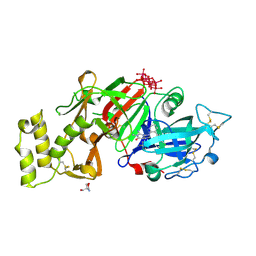 | | Plasmodium vivax PMV-WM08 inhibitor complex | | Descriptor: | (2E,4aR,7aS)-6-[3-(4-chlorophenyl)pyridin-2-yl]-7a-(2,5-difluorophenyl)-2-imino-3-methyloctahydro-4H-pyrrolo[3,4-d]pyrimidin-4-one, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Hodder, A.N, Scally, S.W, Cowman, A.F. | | Deposit date: | 2023-08-25 | | Release date: | 2024-08-28 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Plasmodium vivax PMV-WM08 inhibitor complex
To Be Published
|
|
8TYF
 
 | | Plasmodium vivax PMV-WM06 inhibitor complex | | Descriptor: | (2E,4aR,7aS)-6-[(3M)-3-(2-chlorophenyl)pyridin-2-yl]-7a-(2,5-difluorophenyl)-2-imino-3-methyloctahydro-4H-pyrrolo[3,4-d]pyrimidin-4-one, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Hodder, A.N, Scally, S.W, Czabotar, P.E, Cowman, A.F. | | Deposit date: | 2023-08-25 | | Release date: | 2024-08-28 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Plasmodium vivax PMX-XX inhibitor complex
To Be Published
|
|
8Q4B
 
 | | Endothiapepsin in complex with ligand (3R,5R)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)-5-(3-(2-nitrophenyl)-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol (CBWS-SE-163.1) | | Descriptor: | (3~{R},5~{R})-3-[2-[[ethynyl(methyl)amino]methyl]-1,3-thiazol-4-yl]-5-[3-(2-nitrophenyl)-1,2,4-oxadiazol-5-yl]pyrrolidin-3-ol, (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-08-05 | | Release date: | 2024-08-14 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q14
 
 | | Endothiapepsin in complex with ligand (3R,5R)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)-5-(3-(2-nitro-4-(trifluoromethyl)phenyl)-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol (CBWS-SE-171) | | Descriptor: | (3~{R},5~{R})-3-[2-[[methyl(prop-2-ynyl)amino]methyl]-1,3-thiazol-4-yl]-5-[3-[2-nitro-4-(trifluoromethyl)phenyl]-1,2,4-oxadiazol-5-yl]pyrrolidin-3-ol, (4S)-2-METHYL-2,4-PENTANEDIOL, DIMETHYL SULFOXIDE, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q13
 
 | | Endothiapepsin in complex with ligand (3R,5R)-5-(3-(2-amino-4-(trifluoromethyl)phenyl)-1,2,4-oxadiazol-5-yl)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)pyrrolidin-3-ol (CBWS-SE-168) | | Descriptor: | (3~{R},5~{R})-5-[3-[2-azanyl-4-(trifluoromethyl)phenyl]-1,2,4-oxadiazol-5-yl]-3-[2-[[methyl(prop-2-ynyl)amino]methyl]-1,3-thiazol-4-yl]pyrrolidin-3-ol, (4S)-2-METHYL-2,4-PENTANEDIOL, DIMETHYL SULFOXIDE, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.26 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q12
 
 | | Endothiapepsin in complex with ligand (3R,5R)-5-(3-(2-aminophenyl)-1,2,4-oxadiazol-5-yl)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)pyrrolidin-3-ol (CBWS-SE-146.2) | | Descriptor: | (3~{R},5~{R})-5-[3-(2-aminophenyl)-1,2,4-oxadiazol-5-yl]-3-[2-[[methyl(prop-2-ynyl)amino]methyl]-1,3-thiazol-4-yl]pyrrolidin-3-ol, (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.17 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q11
 
 | | Endothiapepsin in complex with ligand (3R,5R)-5-(3-(4-fluorophenyl)-1,2,4-oxadiazol-5-yl)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)pyrrolidin-3-ol (CBWS-SE-126) | | Descriptor: | (3~{R},5~{R})-5-[3-(4-fluorophenyl)-1,2,4-oxadiazol-5-yl]-3-[2-[[methyl(prop-2-ynyl)amino]methyl]-1,3-thiazol-4-yl]pyrrolidin-3-ol, ACETATE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q10
 
 | | Endothiapepsin in complex with ligand (3R,5R)-3-(2-(hydroxymethyl)thiazol-4-yl)-5-(3-(4-(trifluoromethyl)phenyl)-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol (CBWS-SE-125) | | Descriptor: | (3~{R},5~{R})-3-[2-(hydroxymethyl)-1,3-thiazol-4-yl]-5-[3-[4-(trifluoromethyl)phenyl]-1,2,4-oxadiazol-5-yl]pyrrolidin-3-ol, (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q0Z
 
 | | Endothiapepsin in complex with ligand (3R,5R)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)-5-(3-phenyl-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol (CBWS-SE-089) | | Descriptor: | (3~{R},5~{R})-3-[2-[[methyl(prop-2-ynyl)amino]methyl]-1,3-thiazol-4-yl]-5-(3-phenyl-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol, (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.21 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q0Y
 
 | | Endothiapepsin in complex with ligand (3R,5R)-5-(3-(4-bromophenyl)-1,2,4-oxadiazol-5-yl)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)pyrrolidin-3-ol (CBWS-SE-085.1) | | Descriptor: | (3~{R},5~{R})-5-[3-(4-bromophenyl)-1,2,4-oxadiazol-5-yl]-3-[2-[[methyl(prop-2-ynyl)amino]methyl]-1,3-thiazol-4-yl]pyrrolidin-3-ol, (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.23 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q0X
 
 | | Endothiapepsin in complex with ligand (3R,5R)-3-(2-((methyl(propyl)amino)methyl)thiazol-4-yl)-5-(3-(4-(trifluoromethyl)phenyl)-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol (CBWS-SE-087) | | Descriptor: | (3~{R},5~{R})-3-[2-[[methyl(propyl)amino]methyl]-1,3-thiazol-4-yl]-5-[3-[4-(trifluoromethyl)phenyl]-1,2,4-oxadiazol-5-yl]pyrrolidin-3-ol, ACETATE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q0W
 
 | | Endothiapepsin in complex with ligand (3R,5R)-3-(2-((allyl(methyl)amino)methyl)thiazol-4-yl)-5-(3-(4-(trifluoromethyl)phenyl)-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol (CBWS-SE-086) | | Descriptor: | (3~{R},5~{R})-3-[2-[[methyl(prop-2-enyl)amino]methyl]-1,3-thiazol-4-yl]-5-[3-[4-(trifluoromethyl)phenyl]-1,2,4-oxadiazol-5-yl]pyrrolidin-3-ol, ACETATE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8Q0V
 
 | | Endothiapepsin in complex with ligand (3R,5R)-3-(2-((methyl(prop-2-yn-1-yl)amino)methyl)thiazol-4-yl)-5-(3-(4-(trifluoromethyl)phenyl)-1,2,4-oxadiazol-5-yl)pyrrolidin-3-ol (CBWS-SE-073) | | Descriptor: | (3~{R},5~{R})-3-[2-[[methyl(prop-2-ynyl)amino]methyl]-1,3-thiazol-4-yl]-5-[3-[4-(trifluoromethyl)phenyl]-1,2,4-oxadiazol-5-yl]pyrrolidin-3-ol, ACETATE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-29 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8PXI
 
 | | Crystal structure of Endothiapepsin soaked with FRG283 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 4-(2-{[methyl(prop-2-yn-1-yl)amino]methyl}-1,3-thiazol-4-yl)piperidin-4-ol, DIMETHYL SULFOXIDE, ... | | Authors: | Mueller, J.M, Eckelt, S, Klebe, G, Glinca, S. | | Deposit date: | 2023-07-23 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal structures of Endothiapepsin with ligands derived from merged fragment hits
To Be Published
|
|
8FXQ
 
 | | The Crystal Sturucture of Rhizopuspepsin with a bound modified peptide inhibitor generated by de novo drug design. | | Descriptor: | ALA-CYS-VAL-LYS, CYCLOHEXANE, Rhizopuspepsin, ... | | Authors: | Satyshur, K.A, Rich, D.H, Ripka, A.S. | | Deposit date: | 2023-01-25 | | Release date: | 2023-02-08 | | Method: | X-RAY DIFFRACTION (1.21 Å) | | Cite: | Aspartic protease inhibitors designed from computer-generated templates bind as predicted.
Org Lett, 3, 2001
|
|
8DSR
 
 | | Structure of Plasmepsin X (PM10, PMX) from Plasmodium falciparum 3D7 in complex with UCB7362 | | Descriptor: | (2E,6S)-6-{2-chloro-3-[(2-cyclopropylpyrimidin-5-yl)amino]phenyl}-2-imino-6-methyl-3-[(2S,4S)-2-methyloxan-4-yl]-1,3-diazinan-4-one, Plasmepsin X | | Authors: | Abendroth, J, Lorimer, D.D. | | Deposit date: | 2022-07-22 | | Release date: | 2022-10-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Discovery and Characterization of Potent, Efficacious, Orally Available Antimalarial Plasmepsin X Inhibitors and Preclinical Safety Assessment of UCB7362 .
J.Med.Chem., 65, 2022
|
|
8CIK
 
 | | Altai wapiti (Cervus elaphus sibiricus) chymosin at 2.2 A resolution | | Descriptor: | Chymosin, GLYCEROL | | Authors: | Diusenova, S.E, Shevtsov, M.B, Borshchevskiy, V.I, Belenkaya, S.V, Kolybalov, D.S, Arkhipov, S.G, Volosnikova, E.A, Elchaninov, V.V, Shcherbakov, D.N. | | Deposit date: | 2023-02-09 | | Release date: | 2023-03-08 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Altai wapiti (Cervus elaphus sibiricus) chymosin at 2.2 A resolution
To Be Published
|
|
8C74
 
 | | Pyrrolidine fragment 10d bound to endothiapepsin | | Descriptor: | (3~{R},4~{R})-4-[4-[(4-azanylphenoxy)methyl]-1,2,3-triazol-1-yl]pyrrolidin-3-ol, Endothiapepsin, GLYCEROL | | Authors: | Wiese, J.N, Buehrmann, M, Mueller, M.P, Rauh, D. | | Deposit date: | 2023-01-12 | | Release date: | 2023-05-24 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Fragtory: Pharmacophore-Focused Design, Synthesis, and Evaluation of an sp 3 -Enriched Fragment Library.
J.Med.Chem., 66, 2023
|
|
8C72
 
 | | Pyrrolidine fragment 10b bound to endothiapepsin | | Descriptor: | (3~{S},4~{S})-4-(4-pyridin-2-yl-1,2,3-triazol-1-yl)piperidin-3-ol, Endothiapepsin, GLYCEROL | | Authors: | Wiese, J.N, Buehrmann, M, Mueller, M.P, Rauh, D. | | Deposit date: | 2023-01-12 | | Release date: | 2023-05-24 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Fragtory: Pharmacophore-Focused Design, Synthesis, and Evaluation of an sp 3 -Enriched Fragment Library.
J.Med.Chem., 66, 2023
|
|
8C71
 
 | | Pyrrolidine fragment 5b bound to endothiapepsin | | Descriptor: | (3~{R},4~{R})-4-(3,4-dihydro-1~{H}-isoquinolin-2-yl)pyrrolidin-3-ol, Endothiapepsin, GLYCEROL | | Authors: | Wiese, J.N, Buehrmann, M, Mueller, M.P, Rauh, D. | | Deposit date: | 2023-01-12 | | Release date: | 2023-05-24 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Fragtory: Pharmacophore-Focused Design, Synthesis, and Evaluation of an sp 3 -Enriched Fragment Library.
J.Med.Chem., 66, 2023
|
|
8C70
 
 | | Pyrrolidine fragment 1 bound to endothiapepsin | | Descriptor: | (3~{R},4~{R})-4-[4-[(5-bromanylpyridin-3-yl)oxymethyl]-1,2,3-triazol-1-yl]pyrrolidin-3-ol, Endothiapepsin | | Authors: | Wiese, J.N, Buehrmann, M, Mueller, M.P, Rauh, D. | | Deposit date: | 2023-01-12 | | Release date: | 2023-05-24 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Fragtory: Pharmacophore-Focused Design, Synthesis, and Evaluation of an sp 3 -Enriched Fragment Library.
J.Med.Chem., 66, 2023
|
|
