1MAH
 
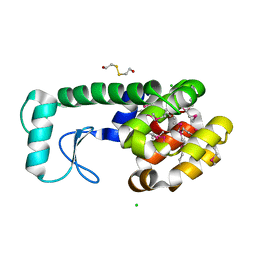
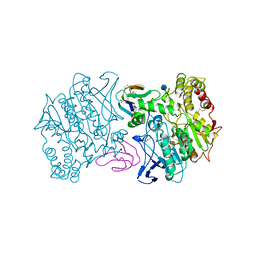 | | FASCICULIN2-MOUSE ACETYLCHOLINESTERASE COMPLEX | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETYLCHOLINESTERASE, FASCICULIN 2 | | Authors: | Bourne, Y, Taylor, P, Marchot, P. | | Deposit date: | 1995-11-21 | | Release date: | 1996-04-03 | | Last modified: | 2021-03-31 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Acetylcholinesterase inhibition by fasciculin: crystal structure of the complex.
Cell(Cambridge,Mass.), 83, 1995
|
|
4YIQ
 
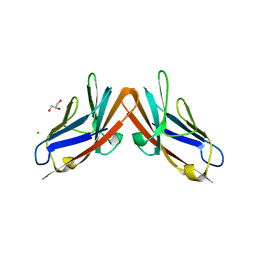 | | Structure of the CEACAM6-CEACAM8 heterodimer | | Descriptor: | CHLORIDE ION, Carcinoembryonic antigen-related cell adhesion molecule 6, Carcinoembryonic antigen-related cell adhesion molecule 8, ... | | Authors: | Bonsor, D.A, Sundberg, E.J. | | Deposit date: | 2015-03-02 | | Release date: | 2015-10-07 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Diverse oligomeric states of CEACAM IgV domains.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
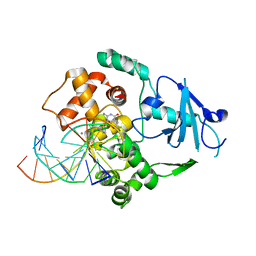
1LV2
 
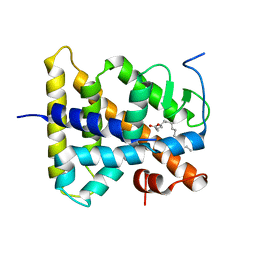 | | Hepatocyte Nuclear Factor 4 is a Transcription Factor that Constitutively Binds Fatty Acids | | Descriptor: | Hepatocyte nuclear factor 4-gamma, PALMITIC ACID | | Authors: | Wisely, B, Miller, A.B, Davis, R.G, Spitzer, T, Shearer, B, Moore, J.T, Johnson, R, Sefler, A, Willson, T.M, Williams, S.P. | | Deposit date: | 2002-05-24 | | Release date: | 2002-12-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Hepatocyte Nuclear Factor 4 Is a Transcription Factor
that Constitutively Binds Fatty Acids.
Structure, 10, 2002
|
|
4YIW
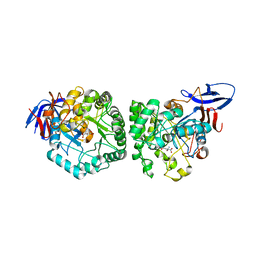 
 | | DIHYDROOROTASE FROM BACILLUS ANTHRACIS WITH SUBSTRATE BOUND | | Descriptor: | Dihydroorotase, N-CARBAMOYL-L-ASPARTATE, ZINC ION | | Authors: | Lei, H, Santarsiero, B.D, Rice, A.J, Lee, H, Johnson, M.E. | | Deposit date: | 2015-03-02 | | Release date: | 2016-08-10 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Ca-asp bound X-ray structure and inhibition of Bacillus anthracis dihydroorotase (DHOase).
Bioorg.Med.Chem., 24, 2016
|
|
1MBH
 
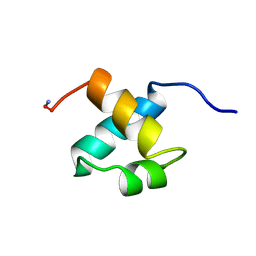 | | MOUSE C-MYB DNA-BINDING DOMAIN REPEAT 2 | | Descriptor: | C-MYB | | Authors: | Ogata, K, Morikawa, S, Nakamura, H, Hojo, H, Yoshimura, S, Zhang, R, Aimoto, S, Ametani, Y, Hirata, Z, Sarai, A, Ishii, S, Nishimura, Y. | | Deposit date: | 1995-05-19 | | Release date: | 1995-09-15 | | Last modified: | 2024-06-05 | | Method: | SOLUTION NMR | | Cite: | Comparison of the free and DNA-complexed forms of the DNA-binding domain from c-Myb.
Nat.Struct.Biol., 2, 1995
|
|
4YJ4
 
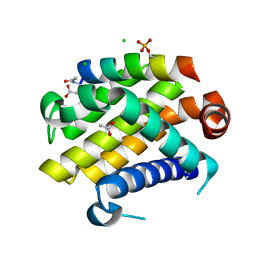 | | Crystal structure of Bcl-xL in complex with the BIM BH3 domain containing Ile155-to-Arg and Glu158-to-phosphoserine mutations | | Descriptor: | ACETATE ION, Bcl-2-like protein 1, BIM BH3 domain, ... | | Authors: | Ku, B, Ha, N.-C, Oh, B.-H. | | Deposit date: | 2015-03-03 | | Release date: | 2015-07-29 | | Last modified: | 2017-09-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Conversion of cell-survival activity of Akt into apoptotic death of cancer cells by two mutations on the BIM BH3 domain.
Cell Death Dis, 6, 2015
|
|
1LVF
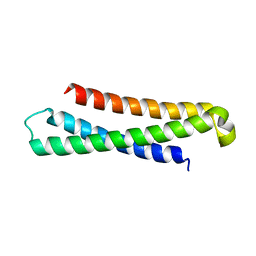 
 | | syntaxin 6 | | Descriptor: | syntaxin 6 | | Authors: | Misura, K.M.S, Bock, J.B, Gonzalez, L.C, Scheller, R.H, Weis, W.I. | | Deposit date: | 2002-05-28 | | Release date: | 2002-07-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Three-dimensional structure of the amino-terminal domain of syntaxin 6, a SNAP-25 C homolog.
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
4YJG
 
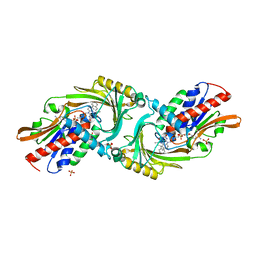 | | Crystal structure of DAAO(Y228L/R283G) variant (R-3-amino 1-phenylbutane binding form) | | Descriptor: | (2R)-4-phenylbutan-2-amine, D-amino-acid oxidase, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Nakano, S, Yasukawa, K, Kawahara, N, Ishitsubo, E, Tokiwa, H, Asano, Y. | | Deposit date: | 2015-03-03 | | Release date: | 2016-04-06 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of DAAO variant
To Be Published
|
|
1LV9
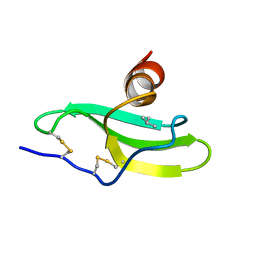 
 | | CXCR3 Binding Chemokine IP-10/CXCL10 | | Descriptor: | Small inducible cytokine B10 | | Authors: | Booth, V, Keizer, D.W, Kamphuis, M.B, Clark-Lewis, I, Sykes, B.D. | | Deposit date: | 2002-05-27 | | Release date: | 2002-09-18 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | The CXCR3 binding chemokine IP-10/CXCL10: structure and receptor interactions.
Biochemistry, 41, 2002
|
|
4YJH
 
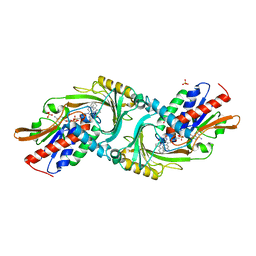 | | Crystal structure of DAAO(Y228L/R283G) variant (R-2-phenylpyrrolidine binding form) | | Descriptor: | (2R)-2-phenylpyrrolidine, D-amino-acid oxidase, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Nakano, S, Yasukawa, K, Kawahara, N, Ishitsubo, E, Tokiwa, H, Asano, Y. | | Deposit date: | 2015-03-03 | | Release date: | 2016-04-06 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of DAAO variant
To Be Published
|
|
1LVW
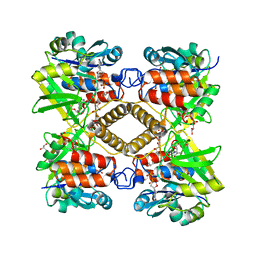 
 | | Crystal structure of glucose-1-phosphate thymidylyltransferase, RmlA, complex with dTDP | | Descriptor: | CHLORIDE ION, GLYCEROL, SULFATE ION, ... | | Authors: | Dong, A, Christendat, D, Pai, E.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2002-05-29 | | Release date: | 2003-07-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of glucose-1-phosphate thymidylyltransferase, RmlA, complex with dTDP
To be Published
|
|
4YJR
 
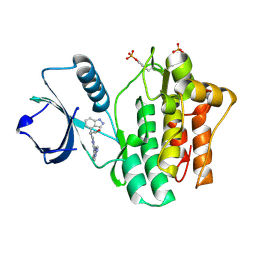 | | SYK kinase domain in complex with inhibitor GTC000225 | | Descriptor: | 3-(1H-indazol-4-yl{2-[(1-methyl-1H-indazol-5-yl)amino]pyrimidin-4-yl}amino)propan-1-ol, Tyrosine-protein kinase SYK | | Authors: | Somers, D.O, Neu, M, Stuckey, J. | | Deposit date: | 2015-03-03 | | Release date: | 2015-09-30 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Discovery of GSK143, a highly potent, selective and orally efficacious spleen tyrosine kinase inhibitor.
Bioorg. Med. Chem. Lett., 21, 2011
|
|
1LVO
 
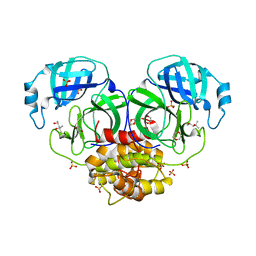 | | Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, 1,4-DIETHYLENE DIOXIDE, Replicase, ... | | Authors: | Anand, K, Palm, G.J, Mesters, J.R, Siddell, S.G, Ziebuhr, J, Hilgenfeld, R. | | Deposit date: | 2002-05-29 | | Release date: | 2002-07-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain.
EMBO J., 21, 2002
|
|
4YJQ
 
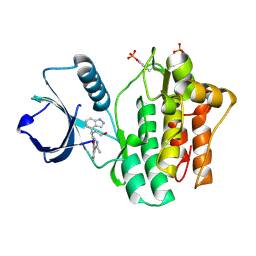 | | SYK kinase domain in complex with inhibitor GTC000224 | | Descriptor: | 3-[1H-indazol-4-yl(2-{[3-(4-methyl-1,3-oxazol-5-yl)phenyl]amino}pyrimidin-4-yl)amino]propan-1-ol, Tyrosine-protein kinase SYK | | Authors: | Somers, D.O, Neu, M, Stuckey, J. | | Deposit date: | 2015-03-03 | | Release date: | 2015-09-30 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Discovery of GSK143, a highly potent, selective and orally efficacious spleen tyrosine kinase inhibitor.
Bioorg. Med. Chem. Lett., 21, 2011
|
|
1LWL
 
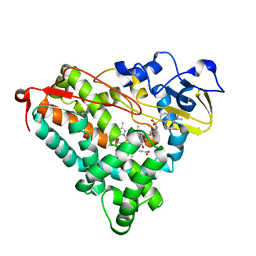 | | Crystal Structure of Cytochrome P450-cam with a Fluorescent Probe D-8-Ad (Adamantane-1-carboxylic acid-5-dimethylamino-naphthalene-1-sulfonylamino-octyl-amide) | | Descriptor: | ADAMANTANE-1-CARBOXYLIC ACID-5-DIMETHYLAMINO-NAPHTHALENE-1-SULFONYLAMINO-OCTYL-AMIDE, Cytochrome P450-cam, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Dunn, A.R, Hays, A.M, Goodin, D.B, Stout, C.D, Chiu, R, Winkler, J.R, Gray, H.B. | | Deposit date: | 2002-05-31 | | Release date: | 2002-12-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Fluorescent probes for cytochrome P450 structural characterization and inhibitor screening
J.AM.CHEM.SOC., 124, 2002
|
|
4YK3
 
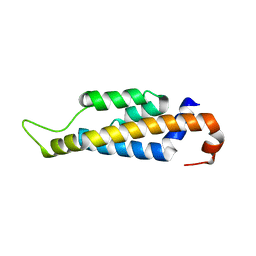 | |
1LWK
 
 | | Multiple Methionine Substitutions are Tolerated in T4 Lysozyme and have Coupled Effects on Folding and Stability | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, CHLORIDE ION, Lysozyme | | Authors: | Gassner, N.C, Baase, W.A, Mooers, B.H.M, Busam, R.D, Weaver, L.H, Lindstrom, J.D, Quillin, M.L, Matthews, B.M. | | Deposit date: | 2002-05-31 | | Release date: | 2003-05-20 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Multiple methionine substitutions are tolerated in T4 lysozyme and have coupled effects on folding and stability.
Biophys.Chem., 100, 2003
|
|
4YK1
 
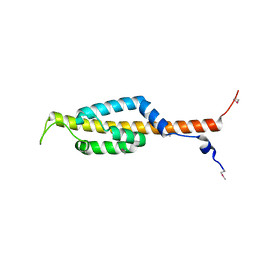 | |
1LWV
 
 | | Borohydride-trapped hOgg1 Intermediate Structure Co-Crystallized with 8-aminoguanine | | Descriptor: | 5'-D(*GP*CP*GP*TP*CP*CP*AP*(PED)P*GP*TP*CP*TP*AP*CP*C)-3', 5'-D(*GP*GP*TP*AP*GP*AP*CP*CP*TP*GP*GP*AP*CP*GP*C)-3', 8-AMINOGUANINE, ... | | Authors: | Fromme, J.C, Bruner, S.D, Yang, W, Karplus, M, Verdine, G.L. | | Deposit date: | 2002-06-03 | | Release date: | 2003-02-25 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Product-Assisted Catalysis in Base Excision DNA Repair
Nat.Struct.Biol., 10, 2003
|
|
4YKH
 
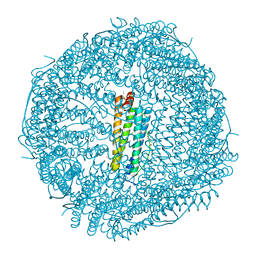 | | Thirty minutes iron loaded human H ferritin | | Descriptor: | BICINE, CHLORIDE ION, FE (II) ION, ... | | Authors: | Pozzi, C, Di Pisa, F, Mangani, S. | | Deposit date: | 2015-03-04 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Iron binding to human heavy-chain ferritin.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
1LWO
 
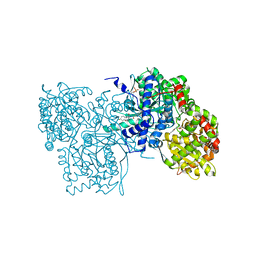 | | Crystal structure of rabbit muscle glycogen phosphorylase a in complex with a potential hypoglycaemic drug at 2.0 A resolution | | Descriptor: | 5-CHLORO-1H-INDOLE-2-CARBOXYLIC ACID [1-(4-FLUOROBENZYL)-2-(4-HYDROXYPIPERIDIN-1YL)-2-OXOETHYL]AMIDE, PYRIDOXAL-5'-PHOSPHATE, alpha-D-glucopyranose, ... | | Authors: | Oikonomakos, N.G, Chrysina, E.D, Kosmopoulou, M.N, Leonidas, D.D. | | Deposit date: | 2002-06-01 | | Release date: | 2002-06-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of rabbit muscle glycogen phosphorylase a in complex with a potential hypoglycaemic drug at 2.0 A resolution
BIOCHEM.BIOPHYS.ACTA PROTEINS & PROTEOMICS, 1647, 2003
|
|
4YKR
 
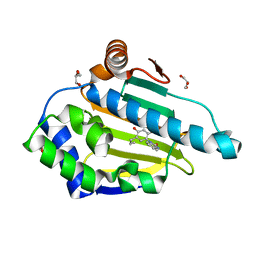 | | Heat Shock Protein 90 Bound to CS302 | | Descriptor: | 1-(2,4-dihydroxy-5-propylphenyl)-1,3-dihydro-2H-benzimidazol-2-one, GLYCEROL, Heat shock protein HSP 90-alpha | | Authors: | Kang, Y.N, Stuckey, J.A. | | Deposit date: | 2015-03-04 | | Release date: | 2016-03-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | Structure of Heat Shock Protein 90 Bound to CS302
To Be Published
|
|
1LXJ
 
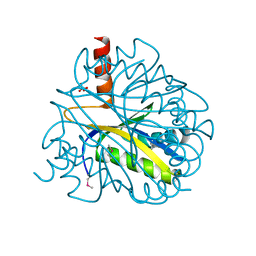 | | X-RAY STRUCTURE OF YBL001c NORTHEAST STRUCTURAL GENOMICS (NESG) CONSORTIUM TARGET YTYst72 | | Descriptor: | HYPOTHETICAL 11.5KDA PROTEIN IN HTB2-NTH2 INTERGENIC REGION, SULFATE ION | | Authors: | Tao, X, Khayat, R, Christendat, D, Savchenko, A, Xu, X, Edwards, A, Arrowsmith, C.H, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2002-06-05 | | Release date: | 2003-07-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | CRYSTAL STRUCTURES OF MTH1187 AND ITS YEAST ORTHOLOG YBL001C
Proteins, 52, 2003
|
|
1LXI
 
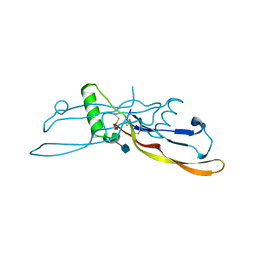 | | Refinement of BMP7 crystal structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, BONE MORPHOGENETIC PROTEIN 7 | | Authors: | Greenwald, J, Groppe, J, Kwiatkowski, W, Choe, S. | | Deposit date: | 2002-06-05 | | Release date: | 2003-04-01 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The BMP7/ActRII Extracellular Domain Complex Provides New Insights into
the Cooperative Nature of Receptor Assembly
Mol.Cell, 11, 2003
|
|
4YKA
 
 | | The structure of Agrobacterium tumefaciens ClpS2 in complex with L-tyrosinamide | | Descriptor: | ATP-dependent Clp protease adapter protein ClpS 2, L-TYROSINAMIDE, SULFATE ION | | Authors: | Stein, B, Grant, R.A, Sauer, R.T, Baker, T.A. | | Deposit date: | 2015-03-04 | | Release date: | 2016-01-27 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.801 Å) | | Cite: | Structural Basis of an N-Degron Adaptor with More Stringent Specificity.
Structure, 24, 2016
|
|
