7S5X
 
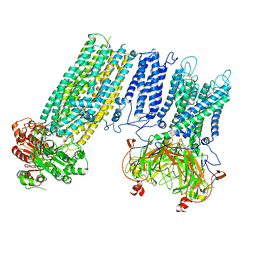 | | Human KATP channel in open conformation, focused on Kir and one SUR, position 1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ATP-binding cassette sub-family C member 8, ... | | Authors: | Zhao, C, MacKinnon, R. | | Deposit date: | 2021-09-12 | | Release date: | 2021-12-01 | | Last modified: | 2021-12-08 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Molecular structure of an open human K ATP channel.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7S8M
 
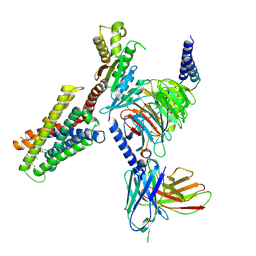 | | CryoEM structure of Gi-coupled MRGPRX2 with peptide agonist Cortistatin-14 | | Descriptor: | Cortistatin 14, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Cao, C, Fay, J.F, Gumpper, R.H, Roth, B.L. | | Deposit date: | 2021-09-18 | | Release date: | 2021-11-17 | | Last modified: | 2021-12-15 | | Method: | ELECTRON MICROSCOPY (2.54 Å) | | Cite: | Structure, function and pharmacology of human itch GPCRs.
Nature, 600, 2021
|
|
7S5Y
 
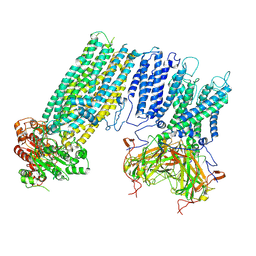 | | Human KATP channel in open conformation, focused on Kir and one SUR, position 2 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ATP-binding cassette sub-family C member 8, ... | | Authors: | Zhao, C, MacKinnon, R. | | Deposit date: | 2021-09-12 | | Release date: | 2021-12-01 | | Last modified: | 2021-12-08 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Molecular structure of an open human K ATP channel.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7S5T
 
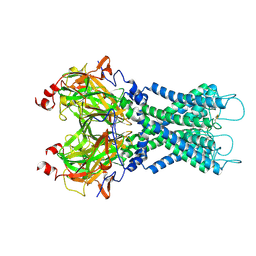 | |
7S61
 
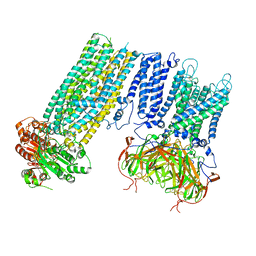 | | Human KATP channel in open conformation, focused on Kir and one SUR, position 5 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ATP-binding cassette sub-family C member 8, ... | | Authors: | Zhao, C, MacKinnon, R. | | Deposit date: | 2021-09-12 | | Release date: | 2021-12-01 | | Last modified: | 2021-12-08 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Molecular structure of an open human K ATP channel.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7S5Z
 
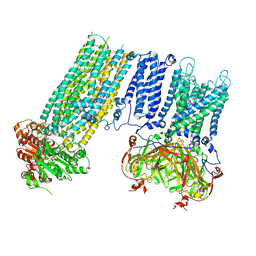 | | Human KATP channel in open conformation, focused on Kir and one SUR, position 3 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ATP-binding cassette sub-family C member 8, ... | | Authors: | Zhao, C, MacKinnon, R. | | Deposit date: | 2021-09-12 | | Release date: | 2021-12-01 | | Last modified: | 2021-12-08 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Molecular structure of an open human K ATP channel.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7S60
 
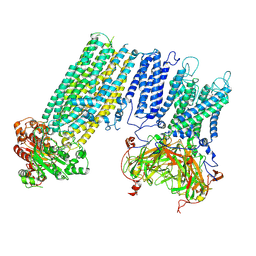 | | Human KATP channel in open conformation, focused on Kir and one SUR, position 4 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ATP-binding cassette sub-family C member 8, ... | | Authors: | Zhao, C, MacKinnon, R. | | Deposit date: | 2021-09-12 | | Release date: | 2021-12-01 | | Last modified: | 2021-12-08 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Molecular structure of an open human K ATP channel.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7S1M
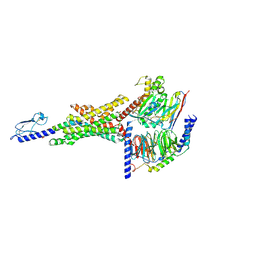 
 | |
7S3I
 
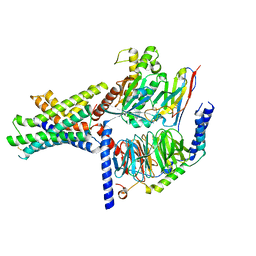 | | Ex4-D-Ala bound to the glucagon-like peptide-1 receptor/g protein complex (conformer 2) | | Descriptor: | Glucagon-like peptide 1 receptor, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Belousoff, M.J, Piper, S.J, Danev, R. | | Deposit date: | 2021-09-07 | | Release date: | 2022-01-05 | | Last modified: | 2022-03-16 | | Method: | ELECTRON MICROSCOPY (2.51 Å) | | Cite: | Structural and functional diversity among agonist-bound states of the GLP-1 receptor.
Nat.Chem.Biol., 18, 2022
|
|
6MF8
 
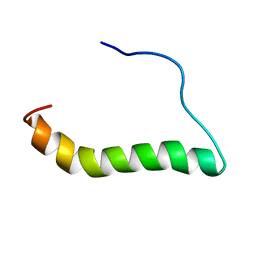 | | TCR alpha transmembrane domain | | Descriptor: | T-cell receptor alpha chain C region | | Authors: | Brazin, K.N, Reinherz, E.L. | | Deposit date: | 2018-09-10 | | Release date: | 2018-12-12 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The T Cell Antigen Receptor alpha Transmembrane Domain Coordinates Triggering through Regulation of Bilayer Immersion and CD3 Subunit Associations.
Immunity, 49, 2018
|
|
6M6E
 
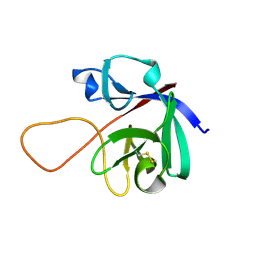 | |
6M6F
 
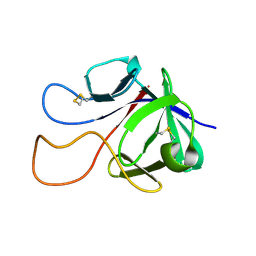 | |
8COO
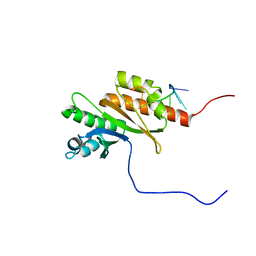 
 | | Solution structure of Zipcode binding protein 1 (ZBP1) KH3(DD)KH4 domains in complex with N6-Methyladenosine containing RNA | | Descriptor: | Insulin-like growth factor 2 mRNA-binding protein 1, RNA_(5'-R(*(UP*CP*GP*GP*(6MZ)P*CP*U)-3') | | Authors: | Nicastro, G, Abis, G, Taylor, I.A, Ramos, A. | | Deposit date: | 2023-02-28 | | Release date: | 2024-02-07 | | Method: | SOLUTION NMR | | Cite: | Direct m6A recognition by IMP1 underlays an alternative model of target selection for non-canonical methyl-readers.
Nucleic Acids Res., 51, 2023
|
|
6V9I
 
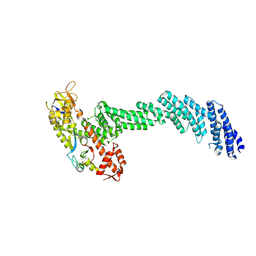 | | cryo-EM structure of Cullin5 bound to RING-box protein 2 (Cul5-Rbx2) | | Descriptor: | Immunoglobulin G-binding protein G,Cullin-5, RING-box protein 2, ZINC ION | | Authors: | Komives, E.A, Lumpkin, R.J, Baker, R.W, Leschziner, A.E. | | Deposit date: | 2019-12-13 | | Release date: | 2020-04-29 | | Last modified: | 2020-11-11 | | Method: | ELECTRON MICROSCOPY (5.2 Å) | | Cite: | Structure and dynamics of the ASB9 CUL-RING E3 Ligase.
Nat Commun, 11, 2020
|
|
6W6L
 
 | |
6MZA
 
 | |
6JNT
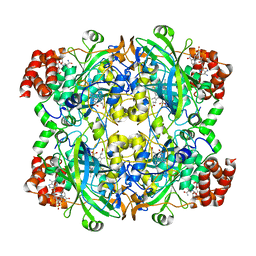 
 | | Catalase structure determined by eEFD (dataset 1) | | Descriptor: | Catalase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Yonekura, K, Maki-Yonekura, S. | | Deposit date: | 2019-03-18 | | Release date: | 2019-04-24 | | Last modified: | 2024-03-27 | | Method: | ELECTRON CRYSTALLOGRAPHY (3 Å) | | Cite: | A new cryo-EM system for electron 3D crystallography by eEFD.
J.Struct.Biol., 206, 2019
|
|
8CUG
 
 | | Synthetic epi-Novo29 (2R,3S), synchrotron structure | | Descriptor: | ACETATE ION, Synthetic epi-Novo29 (2R,3S) | | Authors: | Kreutzer, A.G, Li, X, Krumberger, M, Nowick, J.S. | | Deposit date: | 2022-05-17 | | Release date: | 2023-01-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.131 Å) | | Cite: | Synthesis and Stereochemical Determination of the Peptide Antibiotic Novo29.
J.Org.Chem., 88, 2023
|
|
8CUF
 
 | | Synthetic epi-Novo29 (2R,3S), X-ray diffractometer structure | | Descriptor: | ACETATE ION, IODIDE ION, Synthetic epi-Novo29 (2R,3S) | | Authors: | Kreutzer, A.G, Li, X, Krumberger, M, Nowick, J.S. | | Deposit date: | 2022-05-17 | | Release date: | 2023-01-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Synthesis and Stereochemical Determination of the Peptide Antibiotic Novo29.
J.Org.Chem., 88, 2023
|
|
6JNU
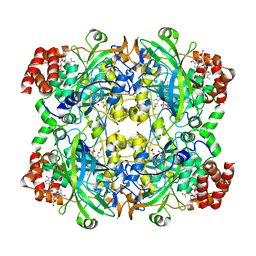 
 | | Catalase structure determined by eEFD (dataset 2) | | Descriptor: | Catalase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Yonekura, K, Maki-Yonekura, S. | | Deposit date: | 2019-03-18 | | Release date: | 2019-04-24 | | Last modified: | 2024-03-27 | | Method: | ELECTRON CRYSTALLOGRAPHY (3.001 Å) | | Cite: | A new cryo-EM system for electron 3D crystallography by eEFD.
J.Struct.Biol., 206, 2019
|
|
6NJF
 
 | | Solution NMR Structure of DANCER3-F34A, a rigid and natively folded single mutant of the dynamic protein DANCER-3 | | Descriptor: | Immunoglobulin G-binding protein G | | Authors: | Damry, A.M, Mayer, M.M, Goto, N.K, Chica, R.A. | | Deposit date: | 2019-01-03 | | Release date: | 2019-08-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Origin of conformational dynamics in a globular protein.
Commun Biol, 2, 2019
|
|
6NZ2
 
 | |
6OCV
 
 | |
6KCZ
 
 | |
6K3K
 
 | | Solution structure of APOBEC3G-CD2 with ssDNA, Product B | | Descriptor: | DNA dC->dU-editing enzyme APOBEC-3G, DNA/RNA (5'-D(*AP*TP*TP*CP*UP*(ICY)P*AP*AP*TP*T)-3'), ZINC ION | | Authors: | Cao, C, Yan, X, Lan, W, Wang, C. | | Deposit date: | 2019-05-19 | | Release date: | 2019-06-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural Investigations on the Interactions between Cytidine Deaminase Human APOBEC3G and DNA.
Chem Asian J, 14, 2019
|
|
