2Y79
 
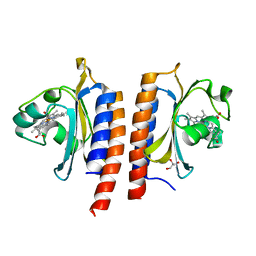 | | STRUCTURE OF THE FIRST GAF DOMAIN E87A MUTANT OF MYCOBACTERIUM TUBERCULOSIS DOSS | | Descriptor: | CALCIUM ION, GLYCEROL, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Cho, H.Y, Cho, H.J, Kang, B.S. | | Deposit date: | 2011-01-30 | | Release date: | 2011-02-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Blockage of the Channel to Heme by the E87 Side Chain in the Gaf Domain of Mycobacterium Tuberculosis Doss Confers the Unique Sensitivity of Doss to Oxygen.
FEBS Lett., 585, 2011
|
|
2XSS
 
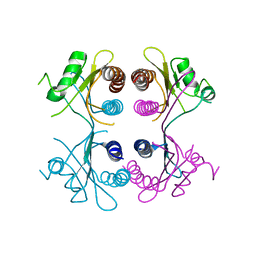 | |
3P01
 
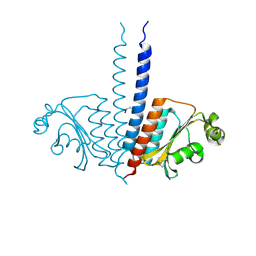 | |
3OOV
 
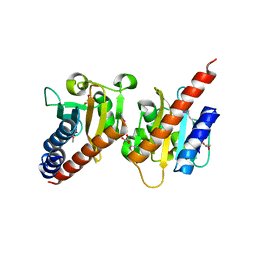 | | Crystal structure of a methyl-accepting chemotaxis protein, residues 122 to 287 | | Descriptor: | GLYCEROL, Methyl-accepting chemotaxis protein, putative | | Authors: | Joachimiak, A, Duke, N.E.C, Hatzos-Skintges, C, Mulligan, R, Clancy, S, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-08-31 | | Release date: | 2010-09-08 | | Last modified: | 2017-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of a methyl-accepting chemotaxis protein, residues 122 to 287
To be Published
|
|
3O5Y
 
 | |
3NOT
 
 | |
3NOP
 
 | |
3NOU
 
 | |
3NHQ
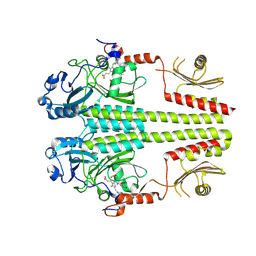 
 | | The dark Pfr structure of the photosensory core module of P. aeruginosa Bacteriophytochrome | | Descriptor: | BILIVERDINE IX ALPHA, Bacteriophytochrome | | Authors: | Yang, X, Ren, Z, Kuk, J, Moffat, K. | | Deposit date: | 2010-06-14 | | Release date: | 2011-11-30 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Temperature-scan cryocrystallography reveals reaction intermediates in bacteriophytochrome.
Nature, 479, 2011
|
|
3MF0
 
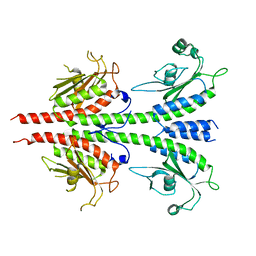 | | Crystal structure of PDE5A GAF domain (89-518) | | Descriptor: | cGMP-specific 3',5'-cyclic phosphodiesterase | | Authors: | Wang, H, Robinson, H, Ke, H. | | Deposit date: | 2010-04-01 | | Release date: | 2010-09-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Conformation changes, N-terminal involvement, and cGMP signal relay in the phosphodiesterase-5 GAF domain.
J.Biol.Chem., 285, 2010
|
|
3LFV
 
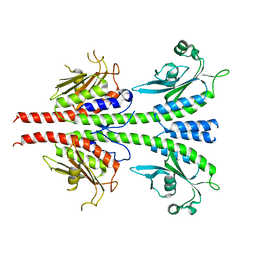 | | crystal structure of unliganded PDE5A GAF domain | | Descriptor: | cGMP-specific 3',5'-cyclic phosphodiesterase | | Authors: | Wang, H, Robinson, H, Ke, H. | | Deposit date: | 2010-01-18 | | Release date: | 2010-09-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Conformation changes, N-terminal involvement, and cGMP signal relay in the phosphodiesterase-5 GAF domain.
J.Biol.Chem., 285, 2010
|
|
3KO6
 
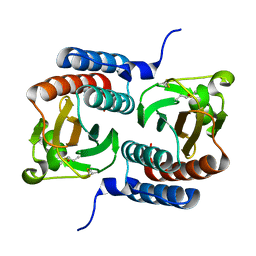 | |
3K2N
 
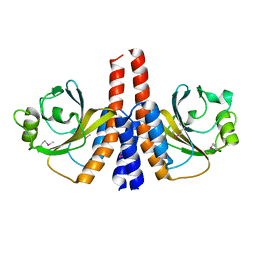 | |
2KOI
 
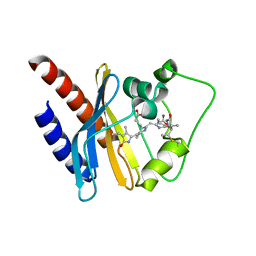 | | Refined solution structure of a cyanobacterial phytochrome GAF domain in the red light-absorbing ground state | | Descriptor: | PHYCOCYANOBILIN, Sensor protein | | Authors: | Cornilescu, C.C, Cornilescu, G, Ulijasz, A.T, Vierstra, R.D, Markley, J.L, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2009-09-22 | | Release date: | 2009-11-03 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the photoconversion of a phytochrome to the activated Pfr form
Nature, 463, 2010
|
|
3IBR
 
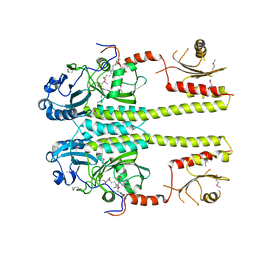 | |
3IBJ
 
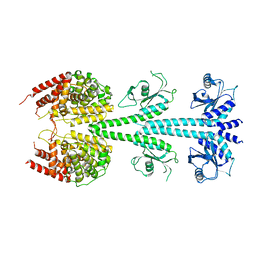 | | X-ray structure of PDE2A | | Descriptor: | MAGNESIUM ION, ZINC ION, cGMP-dependent 3',5'-cyclic phosphodiesterase | | Authors: | Pandit, J. | | Deposit date: | 2009-07-16 | | Release date: | 2009-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.02 Å) | | Cite: | Mechanism for the allosteric regulation of phosphodiesterase 2A deduced from the X-ray structure of a near full-length construct.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
2KLI
 
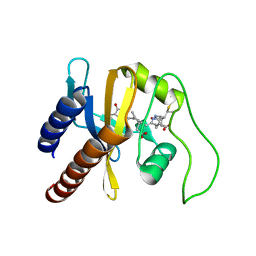 | | Structural Basis for the Photoconversion of A Phytochrome to the Activated FAR-RED LIGHT-ABSORBING Form | | Descriptor: | PHYCOCYANOBILIN, Sensor protein | | Authors: | Cornilescu, C.C, Cornilescu, G, Ulijasz, A.T, Zhang, J, Rivera, M, Vierstra, R.D, Markley, J.L, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2009-07-03 | | Release date: | 2009-11-03 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the photoconversion of a phytochrome to the activated Pfr form
Nature, 463, 2010
|
|
3G6O
 
 | |
2W3E
 
 | |
2W3D
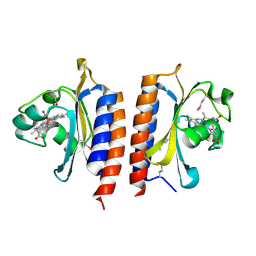 
 | |
2W3H
 
 | | Cyanide bound structure of the first GAF domain of Mycobacterium tuberculosis DosS | | Descriptor: | CALCIUM ION, CYANIDE ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Kang, B.S, Cho, H.Y, Cho, H.J. | | Deposit date: | 2008-11-12 | | Release date: | 2009-03-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Insight Into the Heme-Based Redox Sensing by Doss from Mycobacterium Tuberculosis.
J.Biol.Chem., 284, 2009
|
|
2W3F
 
 | |
2W3G
 
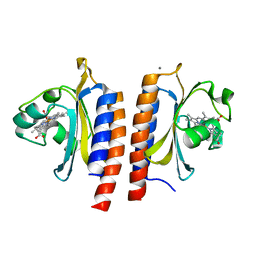 | |
2VZW
 
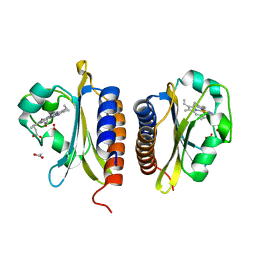 | | X-ray structure of the heme-bound GAF domain of sensory histidine kinase DosT of Mycobacterium tuberculosis | | Descriptor: | ACETATE ION, OXYGEN MOLECULE, PROBABLE HISTIDINE KINASE RESPONSE REGULATOR, ... | | Authors: | Podust, L.M, Ioanoviciu, A, Ortiz de Montellano, P.R. | | Deposit date: | 2008-08-06 | | Release date: | 2009-03-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | 2.3 A X-Ray Structure of the Heme-Bound Gaf Domain of Sensory Histidine Kinase Dost of Mycobacterium Tuberculosis.
Biochemistry, 47, 2008
|
|
3DBA
 
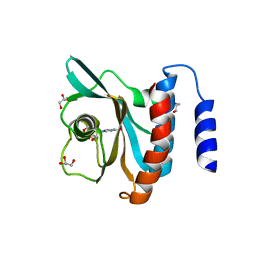 | | Crystal structure of the cGMP-bound GAF a domain from the photoreceptor phosphodiesterase 6C | | Descriptor: | Cone cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha', GLYCEROL, GUANOSINE-3',5'-MONOPHOSPHATE | | Authors: | Martinez, S.E, Heikaus, C.C, Klevit, R.E, Beavo, J.A. | | Deposit date: | 2008-05-30 | | Release date: | 2008-07-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | The Structure of the GAF A Domain from Phosphodiesterase 6C Reveals Determinants of cGMP Binding, a Conserved Binding Surface, and a Large cGMP-dependent Conformational Change.
J.Biol.Chem., 283, 2008
|
|
