6M7Z
 
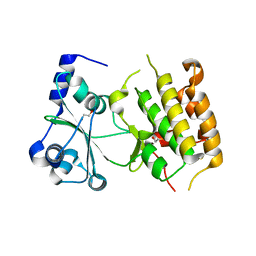 | | A divergent kinase lacking the glycine-rich loop regulates membrane ultrastructure of the Toxoplasma parasitophorous vacuole | | Descriptor: | 1,2-ETHANEDIOL, Bradyzoite pseudokinase 1, CHLORIDE ION | | Authors: | Beraki, T, Borek, D.M, Reese, M.L. | | Deposit date: | 2018-08-21 | | Release date: | 2018-10-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Divergent kinase regulates membrane ultrastructure of theToxoplasmaparasitophorous vacuole.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
6LRA
 
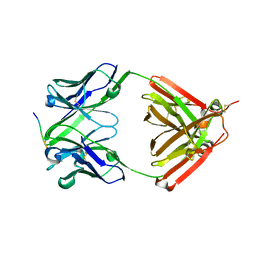 | | The complex structure of PHF core domain peptide of tau and antibody's Fab domain. | | Descriptor: | Fab Heavy Chain, Fab Light Chain, VQIINK | | Authors: | Tomohiro, T, Kouki, S, Tomohiro, S, Takahiro, T, Katsushiro, M, Yasuko, I, Katsuhiko, M, Taizo, T, Toshimitsu, I, Koji, T. | | Deposit date: | 2020-01-15 | | Release date: | 2020-08-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the human tau PHF core domain VQIINK complexed with the Fab domain of monoclonal antibody Tau2r3.
Febs Lett., 2020
|
|
6MEW
 
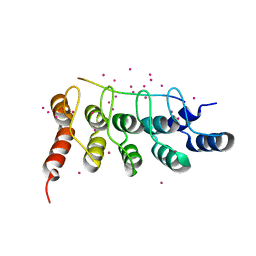 | | RFXANK ankyrin repeats in complex with a RFX7 peptide | | Descriptor: | DNA-binding protein RFXANK, RFX7 peptide, UNKNOWN ATOM OR ION | | Authors: | Tempel, W, Xu, C, Dong, A, Li, Y, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Min, J, Structural Genomics Consortium (SGC) | | Deposit date: | 2018-09-07 | | Release date: | 2018-10-03 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | RFXANK ankyrin repeats in complex with a RFX7 peptide
to be published
|
|
6MF9
 
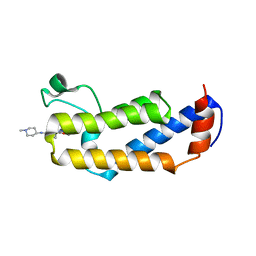 | | Crystal structure of CGD4-650 with compound BI2536 | | Descriptor: | 4-{[(7R)-8-cyclopentyl-7-ethyl-5-methyl-6-oxo-5,6,7,8-tetrahydropteridin-2-yl]amino}-3-methoxy-N-(1-methylpiperidin-4-yl)benzamide, ZnKn (C2HC)+Athook+bromo domain protein, Taf250, ... | | Authors: | Dong, A, Lin, Y.L, Hou, D, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Vedadi, M, Hui, R, Structural Genomics Consortium (SGC) | | Deposit date: | 2018-09-10 | | Release date: | 2018-10-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.037 Å) | | Cite: | Crystal structure of CGD4-650 with compound BI2536
to be published
|
|
6LXS
 
 | | CWA protein-SdrC | | Descriptor: | CALCIUM ION, GLYCEROL, Ser-Asp rich fibrinogen-binding, ... | | Authors: | Pi, Y, Ji, Q. | | Deposit date: | 2020-02-11 | | Release date: | 2020-04-22 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Structural Basis ofStaphylococcus aureusSurface Protein SdrC.
Biochemistry, 59, 2020
|
|
6M2E
 
 | | Sirohydrochlorin nickelochelatase CfbA in complex with sirohydrochlorin | | Descriptor: | 3,3',3'',3'''-[(7S,8S,12S,13S)-3,8,13,17-tetrakis(carboxymethyl)-8,13-dimethyl-7,8,12,13-tetrahydroporphyrin-2,7,12,18-tetrayl]tetrapropanoic acid, Sirohydrochlorin cobaltochelatase | | Authors: | Fujishiro, T. | | Deposit date: | 2020-02-27 | | Release date: | 2021-01-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The nickel-sirohydrochlorin formation mechanism of the ancestral class II chelatase CfbA in coenzyme F430 biosynthesis.
Chem Sci, 12, 2021
|
|
3RI0
 
 | | Epitope backbone grafting by computational design for improved presentation of linear epitopes on scaffold proteins | | Descriptor: | BB_2cx5_001, GLYCEROL, SULFATE ION | | Authors: | Azoitei, M.L, Ban, Y.A, Julien, J.P, Bryson, S, Schroeter, A, Kalyuzhniy, O, Porter, J.R, Adachi, Y, Baker, D, Szabo, E, Pai, E.F, Schief, W.R. | | Deposit date: | 2011-04-12 | | Release date: | 2011-11-09 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Computational design of high-affinity epitope scaffolds by backbone grafting of a linear epitope.
J.Mol.Biol., 415, 2012
|
|
3RK9
 
 | | CDK2 in complex with inhibitor RC-2-74 | | Descriptor: | Cyclin-dependent kinase 2, [4-amino-2-(propan-2-ylamino)-1,3-thiazol-5-yl](pyridin-3-yl)methanone | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-04-17 | | Release date: | 2012-10-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Development of highly potent and selective diaminothiazole inhibitors of cyclin-dependent kinases.
J.Med.Chem., 56, 2013
|
|
3R8L
 
 | | CDK2 in complex with inhibitor L3-4 | | Descriptor: | (5R)-5-tert-butyl-4,5,6,7-tetrahydro-1H-indazole-3-carbohydrazide, Cyclin-dependent kinase 2 | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-03-24 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-guided optimization of novel CDK2 inhibitors discovered by high-throughput screening
To be Published
|
|
3RMY
 
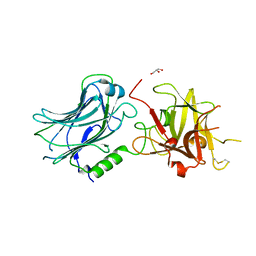 | | Crystal structure of HCR/D W1238A mutant | | Descriptor: | Botulinum neurotoxin type D, GLYCEROL | | Authors: | Fu, Z, Karalewitz, A, Kroken, A, Kim, J.-J.P, Barbieri, J.T. | | Deposit date: | 2011-04-21 | | Release date: | 2011-06-01 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Novel Ganglioside-mediated Entry of Botulinum Neurotoxin Serotype D into Neurons.
J.Biol.Chem., 286, 2011
|
|
6LYS
 
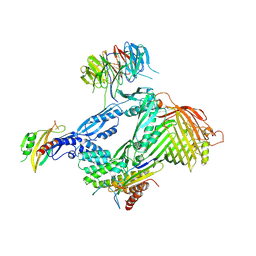 | | Structure of the BAM complex | | Descriptor: | Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, Outer membrane protein assembly factor BamC, ... | | Authors: | Xiao, L, Huang, Y. | | Deposit date: | 2020-02-15 | | Release date: | 2021-01-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Structures of the beta-barrel assembly machine recognizing outer membrane protein substrates.
Faseb J., 35, 2021
|
|
8PI5
 
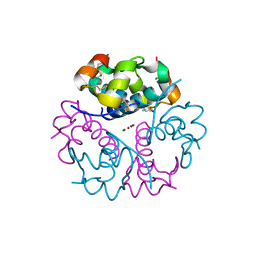 | |
3NS5
 | |
3RMF
 
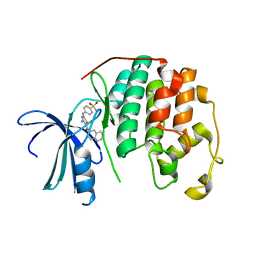 | | CDK2 in complex with inhibitor RC-2-33 | | Descriptor: | 4-{[4-amino-5-(naphthalen-2-ylcarbonyl)-1,3-thiazol-2-yl]amino}benzenesulfonamide, Cyclin-dependent kinase 2 | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-04-20 | | Release date: | 2012-10-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Development of highly potent and selective diaminothiazole inhibitors of cyclin-dependent kinases.
J.Med.Chem., 56, 2013
|
|
6M9G
 
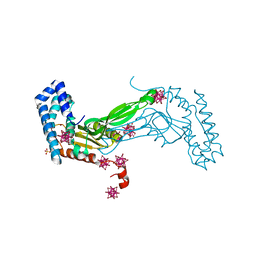 | | BbvCI B2 dimer with Ta6Br14 clusters | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, BROMIDE ION, ... | | Authors: | Shen, B.W, Stoddard, B.L. | | Deposit date: | 2018-08-23 | | Release date: | 2018-11-14 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structure, subunit organization and behavior of the asymmetric Type IIT restriction endonuclease BbvCI.
Nucleic Acids Res., 47, 2019
|
|
3ROY
 
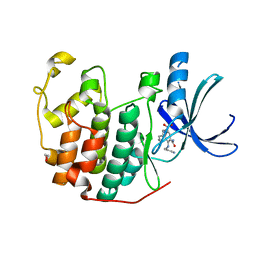 | | CDK2 in complex with inhibitor KVR-1-154 | | Descriptor: | 1,2-ETHANEDIOL, 4-(hexylamino)-5-nitro-2-[(pyridin-3-ylmethyl)amino]benzamide, Cyclin-dependent kinase 2 | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-04-26 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure-guided optimization of novel CDK2 inhibitors discovered by high-throughput screening
To be Published
|
|
3R8P
 
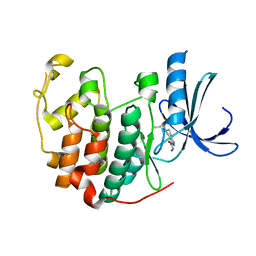 | | CDK2 in complex with inhibitor NSK-MC1-6 | | Descriptor: | (5R)-5-propyl-4,5,6,7-tetrahydro-1H-indazole-3-carbohydrazide, Cyclin-dependent kinase 2 | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-03-24 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure-guided optimization of novel CDK2 inhibitors discovered by high-throughput screening
To be Published
|
|
6M9J
 
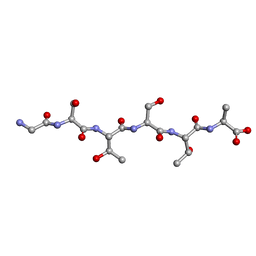 | | Racemic-GSTSTA from degenerate octameric repeats in InaZ, residues 707-712 | | Descriptor: | Ice nucleation protein | | Authors: | Zee, C, Glynn, C, Gallagher-Jones, M, Miao, J, Santiago, C.G, Cascio, D, Gonen, T, Sawaya, M.R, Rodriguez, J.A. | | Deposit date: | 2018-08-23 | | Release date: | 2019-03-27 | | Last modified: | 2024-03-13 | | Method: | ELECTRON CRYSTALLOGRAPHY (0.9 Å) | | Cite: | Homochiral and racemic MicroED structures of a peptide repeat from the ice-nucleation protein InaZ.
IUCrJ, 6, 2019
|
|
3R9N
 
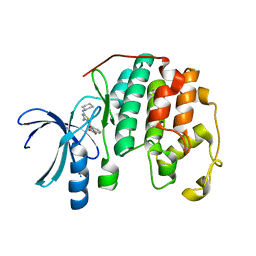 | | CDK2 in complex with inhibitor RC-2-21 | | Descriptor: | Cyclin-dependent kinase 2, [4-amino-2-(cyclohexylamino)-1,3-thiazol-5-yl](3-nitrophenyl)methanone | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-03-25 | | Release date: | 2012-10-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Development of highly potent and selective diaminothiazole inhibitors of cyclin-dependent kinases.
J.Med.Chem., 56, 2013
|
|
6M98
 | |
6MC0
 
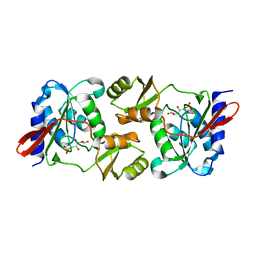 | |
6MFU
 
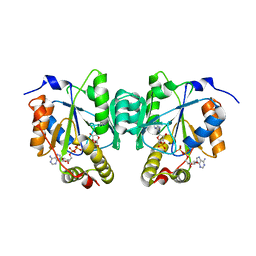 | |
6MAG
 
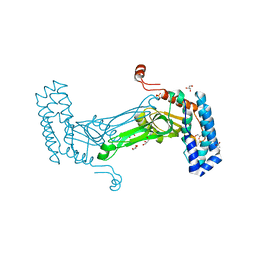 | | native BbvCI B2 dimer in space group C222 | | Descriptor: | ACETATE ION, BbvCI endonuclease subunit 2, GLYCEROL, ... | | Authors: | Shen, B.W, Stoddard, B.L. | | Deposit date: | 2018-08-27 | | Release date: | 2018-11-14 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Structure, subunit organization and behavior of the asymmetric Type IIT restriction endonuclease BbvCI.
Nucleic Acids Res., 47, 2019
|
|
3QXP
 
 | | CDK2 in complex with inhibitor RC-3-89 | | Descriptor: | 1,2-ETHANEDIOL, 4-{[4-amino-5-(2-nitrobenzoyl)-1,3-thiazol-2-yl]amino}benzenesulfonamide, Cyclin-dependent kinase 2 | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-03-02 | | Release date: | 2012-10-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Development of highly potent and selective diaminothiazole inhibitors of cyclin-dependent kinases.
J.Med.Chem., 56, 2013
|
|
3QZH
 
 | | CDK2 in complex with inhibitor KVR-1-124 | | Descriptor: | 1,2-ETHANEDIOL, 4-methoxy-5-nitro-2-[(pyridin-3-ylmethyl)amino]benzamide, Cyclin-dependent kinase 2 | | Authors: | Betzi, S, Alam, R, Han, H, Becker, A, Schonbrunn, E. | | Deposit date: | 2011-03-06 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure-guided optimization of novel CDK2 inhibitors discovered by high-throughput screening
To be Published
|
|
