4BL7
 
 | |
4BVY
 
 | |
7YRB
 
 | | UBR box of human UBR6 | | Descriptor: | F-box protein 11, isoform CRA_f, SULFATE ION, ... | | Authors: | Kim, B, Song, H.K. | | Deposit date: | 2022-08-09 | | Release date: | 2023-11-15 | | Last modified: | 2025-04-23 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | Revisiting the structure of UBR box from human UBR6.
Protein Sci., 34, 2025
|
|
6HR8
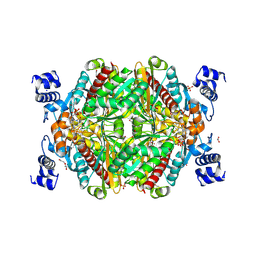 
 | | HMG-CoA reductase from Methanothermococcus thermolithotrophicus in complex with NADPH at 2.9 A resolution | | Descriptor: | DI(HYDROXYETHYL)ETHER, HMG-CoA reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Wagner, T, Voegeli, B, Erb, T.J, Shima, S. | | Deposit date: | 2018-09-26 | | Release date: | 2019-02-13 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of archaeal HMG-CoA reductase: insights into structural changes of the C-terminal helix of the class-I enzyme.
Febs Lett., 593, 2019
|
|
6HR7
 
 | | HMG-CoA reductase from Methanothermococcus thermolithotrophicus apo form at 2.4 A resolution | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, CHLORIDE ION, GLYCEROL, ... | | Authors: | Wagner, T, Voegeli, B, Erb, T.J, Shima, S. | | Deposit date: | 2018-09-26 | | Release date: | 2019-02-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of archaeal HMG-CoA reductase: insights into structural changes of the C-terminal helix of the class-I enzyme.
Febs Lett., 593, 2019
|
|
7F07
 
 | | Autonomous VH domain that interacts with eIF4E at the Capped mRNA Binding site. | | Descriptor: | Eukaryotic translation initiation factor 4E, VH domain (VH-DiFCAP-01) | | Authors: | Brown, C.J, Frosi, Y, Ng, S, Lin, Y.C. | | Deposit date: | 2021-06-03 | | Release date: | 2022-06-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Development of a novel peptide aptamer that interacts with the eIF4E capped-mRNA binding site using peptide epitope linker evolution (PELE).
Rsc Chem Biol, 3, 2022
|
|
2PAV
 
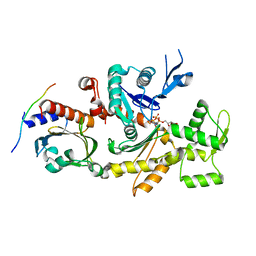 | | Ternary complex of Profilin-Actin with the Last Poly-Pro of Human VASP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin, alpha skeletal muscle, ... | | Authors: | Ferron, F, Rebowski, G, Dominguez, R. | | Deposit date: | 2007-03-27 | | Release date: | 2007-10-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for the recruitment of profilin-actin complexes during filament elongation by Ena/VASP.
Embo J., 26, 2007
|
|
5XLN
 
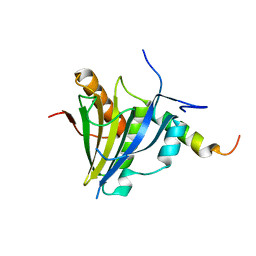 | |
7KCI
 
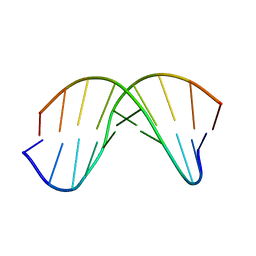 | | DETERMINANTS OF REPRESSOR/OPERATOR RECOGNITION FROM THE STRUCTURE OF THE TRP OPERATOR BINDING SITE | | Descriptor: | Self-complementary deoxyoligonucleotide decamer d(CCACTAGTGG) | | Authors: | Shakked, Z, Guzikevich-Guerstein, G, Frolow, F, Rabinovich, D, Joachimiak, A, Sigler, P.B. | | Deposit date: | 1994-09-12 | | Release date: | 2020-10-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Determinants of repressor/operator recognition from the structure of the trp operator binding site.
Nature, 368, 1994
|
|
7CJN
 
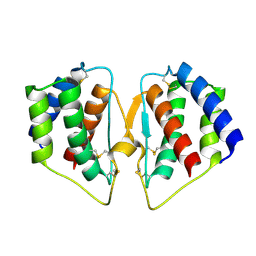 | | grass carp interleukin-2 | | Descriptor: | Interleukin | | Authors: | Wang, J. | | Deposit date: | 2020-07-12 | | Release date: | 2021-07-14 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.66 Å) | | Cite: | Structural insights into the co-evolution of IL-2 and its private receptor
To Be Published
|
|
8QJ7
 
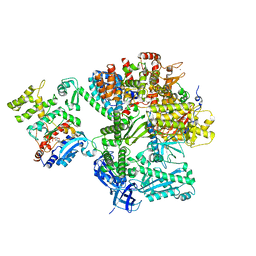 | |
3PMN
 
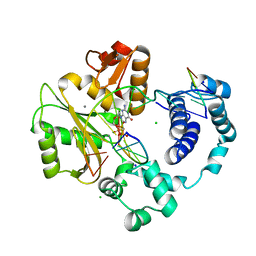 | | ternary crystal structure of polymerase lambda variant with a GT mispair at the primer terminus with Mn2+ in the active site | | Descriptor: | 2'-deoxy-5'-O-[(R)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]methyl}phosphoryl]guanosine, 5'-D(*CP*AP*GP*TP*AP*G)-3', 5'-D(*CP*GP*GP*CP*CP*TP*TP*AP*CP*TP*G)-3', ... | | Authors: | Bebenek, K, Pedersen, L.C, Kunkel, T.A. | | Deposit date: | 2010-11-17 | | Release date: | 2011-02-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Replication infidelity via a mismatch with Watson-Crick geometry.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
6YVH
 
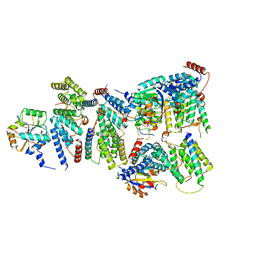 | | CWC22-CWC27-EIF4A3 Complex | | Descriptor: | Eukaryotic initiation factor 4A-III, Pre-mRNA-splicing factor CWC22 homolog, Spliceosome-associated protein CWC27 homolog | | Authors: | Basquin, J, Busetto, V, LeHir, H, Conti, E. | | Deposit date: | 2020-04-28 | | Release date: | 2020-05-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.19 Å) | | Cite: | Structural and functional insights into CWC27/CWC22 heterodimer linking the exon junction complex to spliceosomes.
Nucleic Acids Res., 48, 2020
|
|
3PNC
 
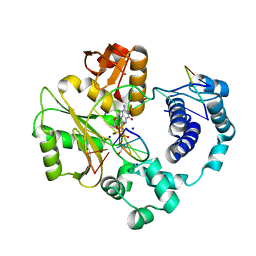 | | Ternary crystal structure of a polymerase lambda variant with a GT mispair at the primer terminus and sodium at catalytic metal site | | Descriptor: | 2'-deoxy-5'-O-[(R)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]methyl}phosphoryl]guanosine, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 5'-D(*CP*AP*GP*TP*AP*G)-3', ... | | Authors: | Bebenek, K, Pedersen, L.C, Kunkel, T.A. | | Deposit date: | 2010-11-18 | | Release date: | 2011-02-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Replication infidelity via a mismatch with Watson-Crick geometry.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
7FB6
 
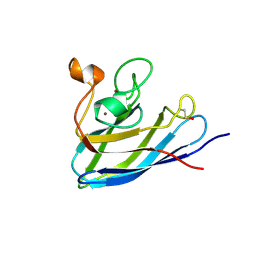 | | C57D/C146D mutant of Human Cu, Zn Superoxide Dismutase (SOD1) | | Descriptor: | CHLORIDE ION, COPPER (II) ION, Superoxide dismutase [Cu-Zn], ... | | Authors: | Baek, Y, Ha, N.-C. | | Deposit date: | 2021-07-08 | | Release date: | 2022-10-12 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural analysis of the overoxidized Cu/Zn-superoxide dismutase in ROS-induced ALS filament formation.
Commun Biol, 5, 2022
|
|
7FB9
 
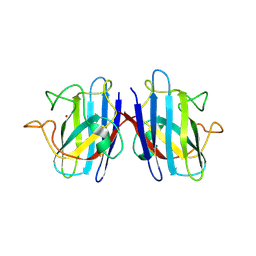 | | Crystal Structure of Human Cu, Zn Superoxide Dismutase (SOD1) | | Descriptor: | COPPER (II) ION, Superoxide dismutase [Cu-Zn], ZINC ION | | Authors: | Baek, Y, Ha, N.-C. | | Deposit date: | 2021-07-08 | | Release date: | 2022-10-12 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural analysis of the overoxidized Cu/Zn-superoxide dismutase in ROS-induced ALS filament formation.
Commun Biol, 5, 2022
|
|
6EO1
 | | The electron crystallography structure of the cAMP-bound potassium channel MloK1 (PCO-refined) | | Descriptor: | Cyclic nucleotide-gated potassium channel mll3241, POTASSIUM ION | | Authors: | Kowal, J, Biyani, N, Chami, M, Scherer, S, Rzepiela, A, Baumgartner, P, Upadhyay, V, Nimigean, C, Stahlberg, H. | | Deposit date: | 2017-10-08 | | Release date: | 2017-12-27 | | Last modified: | 2024-11-06 | | Method: | ELECTRON CRYSTALLOGRAPHY (4.5 Å) | | Cite: | High-Resolution Cryoelectron Microscopy Structure of the Cyclic Nucleotide-Modulated Potassium Channel MloK1 in a Lipid Bilayer.
Structure, 26, 2018
|
|
3FLO
 
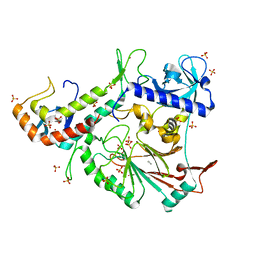 | |
5K2H
 | |
2W5Z
 
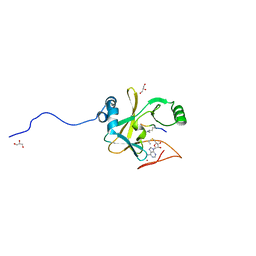 | | Ternary Complex of the Mixed Lineage Leukaemia (MLL1) SET Domain with the cofactor product S-Adenosylhomocysteine and histone peptide. | | Descriptor: | GLYCEROL, HISTONE PEPTIDE, HISTONE-LYSINE N-METHYLTRANSFERASE HRX, ... | | Authors: | Southall, S.M, Wong, P.S, Odho, Z, Roe, S.M, Wilson, J.R. | | Deposit date: | 2008-12-15 | | Release date: | 2009-02-10 | | Last modified: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Basis for the Requirement of Additional Factors for Mll1 Set Domain Activity and Recognition of Epigenetic Marks.
Mol.Cell, 33, 2009
|
|
6H6T
 | |
6H6U
 | |
7JK2
 
 | |
7JK4
 
 | | Structure of Drosophila ORC bound to AT-rich DNA and Cdc6 | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Cell division control protein, DNA (34-MER), ... | | Authors: | Schmidt, J.M, Bleichert, F. | | Deposit date: | 2020-07-27 | | Release date: | 2020-09-09 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6.
Nat Commun, 11, 2020
|
|
7JGS
 
 | |
