1D4T
 
 | | CRYSTAL STRUCTURE OF THE XLP PROTEIN SAP IN COMPLEX WITH A SLAM PEPTIDE | | Descriptor: | SIGNALING LYMPHOCYTIC ACTIVATION MOLECULE, T CELL SIGNAL TRANSDUCTION MOLECULE SAP | | Authors: | Poy, F, Yaffe, M.B, Sayos, J, Saxena, K, Eck, M.J. | | Deposit date: | 1999-10-06 | | Release date: | 1999-10-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal structures of the XLP protein SAP reveal a class of SH2 domains with extended, phosphotyrosine-independent sequence recognition.
Mol.Cell, 4, 1999
|
|
1CKT
 
 | | CRYSTAL STRUCTURE OF HMG1 DOMAIN A BOUND TO A CISPLATIN-MODIFIED DNA DUPLEX | | Descriptor: | Cisplatin, DNA (5'-D(*CP*CP*(5IU)P*CP*TP*CP*TP*GP*GP*AP*CP*CP*TP*TP*CP*C)-3'), DNA (5'-D(*GP*GP*AP*AP*GP*GP*TP*CP*CP*AP*GP*AP*GP*AP*GP*G)-3'), ... | | Authors: | Ohndorf, U.-M, Rould, M.A, Pabo, C.O, Lippard, S.J. | | Deposit date: | 1999-04-23 | | Release date: | 1999-06-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Basis for recognition of cisplatin-modified DNA by high-mobility-group proteins.
Nature, 399, 1999
|
|
1VFQ
 
 | |
1CG6
 
 | |
1D4W
 
 | | CRYSTAL STRUCTURE OF THE XLP PROTEIN SAP IN COMPLEX WITH SLAM PHOSPHOPEPTIDE | | Descriptor: | SIGNALING LYMPHOCYTIC ACTIVATION MOLECULE, T CELL SIGNAL TRANSDUCTION MOLECULE SAP | | Authors: | Poy, F, Yaffe, M.B, Sayos, J, Saxena, K, Eck, M.J. | | Deposit date: | 1999-10-06 | | Release date: | 1999-10-14 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structures of the XLP protein SAP reveal a class of SH2 domains with extended, phosphotyrosine-independent sequence recognition.
Mol.Cell, 4, 1999
|
|
1DZ1
 
 | | Mouse HP1 (M31) C terminal (shadow chromo) domain | | Descriptor: | MODIFIER 1 PROTEIN | | Authors: | Brasher, S.V, Smith, B.O, Fogh, R.H, Nietlispach, D, Thiru, A, Nielsen, P.R, Broadhurst, R.W, Ball, L.J, Murzina, N, Laue, E.D. | | Deposit date: | 2000-02-11 | | Release date: | 2000-04-09 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Structure of Mouse Hp1 Suggests a Unique Mode of Single Peptide Recognition by the Shadow Chromo Domain Dimer
Embo J., 19, 2000
|
|
1E33
 
 | | Crystal structure of an Arylsulfatase A mutant P426L | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Arylsulfatase A, MAGNESIUM ION | | Authors: | von Buelow, R, Schmidt, B, Dierks, T, von Figura, K, Uson, I. | | Deposit date: | 2000-06-06 | | Release date: | 2001-05-25 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Defective oligomerization of arylsulfatase a as a cause of its instability in lysosomes and metachromatic leukodystrophy.
J. Biol. Chem., 277, 2002
|
|
1T2K
 
 | | Structure Of The DNA Binding Domains Of IRF3, ATF-2 and Jun Bound To DNA | | Descriptor: | 31-MER, Cyclic-AMP-dependent transcription factor ATF-2, Interferon regulatory factor 3, ... | | Authors: | Panne, D, Maniatis, T, Harrison, S.C. | | Deposit date: | 2004-04-21 | | Release date: | 2004-11-16 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of ATF-2/c-Jun and IRF-3 bound to the interferon-beta enhancer.
Embo J., 23, 2004
|
|
1DWV
 
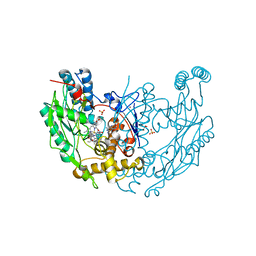 | | MURINE INDUCIBLE NITRIC OXIDE SYNTHASE OXYGENASE DIMER N-hydroxyarginine and 4-amino tetrahydrobiopterin | | Descriptor: | 2,4-DIAMINO-6-[2,3-DIHYDROXY-PROP-3-YL]-5,6,7,8-TETRAHYDROPTERIDINE, N-OMEGA-HYDROXY-L-ARGININE, NITRIC OXIDE SYNTHASE, ... | | Authors: | Crane, B.R, Arvai, A.S, Getzoff, E.D, Stuehr, D.J, Tainer, J.A. | | Deposit date: | 1999-12-14 | | Release date: | 2000-02-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structures of the N(Omega)-Hydroxy-L-Arginine Complex of Inducible Nitric Oxide Synthase Oxygenase Dimer with Active Andinactive Pterins
Biochemistry, 39, 2000
|
|
1TA3
 
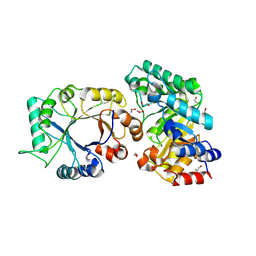 | | Crystal Structure of xylanase (GH10) in complex with inhibitor (XIP) | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Endo-1,4-beta-xylanase, ... | | Authors: | Payan, F, Leone, P, Furniss, C, Tahir, T, Durand, A, Porciero, S, Manzanares, P, Williamson, G, Gilbert, H.J, Juge, N, Roussel, A. | | Deposit date: | 2004-05-19 | | Release date: | 2004-07-20 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Dual Nature of the Wheat Xylanase Protein Inhibitor XIP-I: STRUCTURAL BASIS FOR THE INHIBITION OF FAMILY 10 AND FAMILY 11 XYLANASES.
J.Biol.Chem., 279, 2004
|
|
1TFC
 
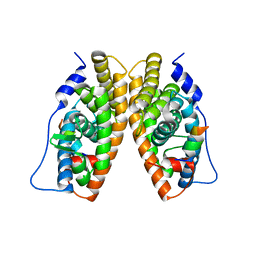 | | CRYSTAL STRUCTURE OF THE LIGAND-BINDING DOMAIN OF THE ESTROGEN-RELATED RECEPTOR GAMMA IN COMPLEX WITH A STEROID RECEPTOR COACTIVATOR-1 PEPTIDE | | Descriptor: | Estrogen-related receptor gamma, steroid receptor coactivator-1 | | Authors: | Greschik, H, Flaig, R, Renaud, J.P, Moras, D. | | Deposit date: | 2004-05-27 | | Release date: | 2004-07-27 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Basis for the Deactivation of the Estrogen-related Receptor {gamma} by Diethylstilbestrol or 4-Hydroxytamoxifen and Determinants of Selectivity.
J.Biol.Chem., 279, 2004
|
|
1W0H
 
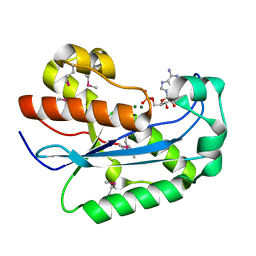 | | Crystallographic structure of the nuclease domain of 3'hExo, a DEDDh family member, bound to rAMP | | Descriptor: | 3'-5' EXONUCLEASE ERI1, ADENOSINE MONOPHOSPHATE, MAGNESIUM ION | | Authors: | Cheng, Y, Patel, D. | | Deposit date: | 2004-06-04 | | Release date: | 2004-09-30 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Crystallographic Structure of the Nuclease Domain of 3'Hexo, a Deddh Family Member, Bound to Ramp
J.Mol.Biol., 343, 2004
|
|
1W0E
 
 | | Crystal structure of human cytochrome P450 3A4 | | Descriptor: | CYTOCHROME P450 3A4, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Williams, P.A, Cosme, J, Vinkovic, D.M, Ward, A, Angove, H.C, Day, P.J, Vonrhein, C, Tickle, I.J, Jhoti, H. | | Deposit date: | 2004-06-03 | | Release date: | 2004-07-22 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structures of Human Cytochrome P450 3A4 Bound to Metyrapone and Progesterone
Science, 305, 2004
|
|
1W6S
 
 | | The high resolution structure of methanol dehydrogenase from methylobacterium extorquens | | Descriptor: | CALCIUM ION, GLYCEROL, METHANOL DEHYDROGENASE SUBUNIT 1, ... | | Authors: | Williams, P.A, Coates, L, Mohammed, F, Gill, R, Erskine, P.T, Wood, S.P, Anthony, C, Cooper, J.B. | | Deposit date: | 2004-08-23 | | Release date: | 2004-12-21 | | Last modified: | 2019-05-22 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | The Atomic Resolution Structure of Methanol Dehydrogenase from Methylobacterium Extorquens
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1VFL
 
 | | Adenosine deaminase | | Descriptor: | Adenosine deaminase, ZINC ION | | Authors: | Kinoshita, T. | | Deposit date: | 2004-04-16 | | Release date: | 2005-08-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Basis of Compound Recognition by Adenosine Deaminase
Biochemistry, 44, 2005
|
|
1VGR
 
 | | Formyl-CoA transferase mutant Asp169 to Glu | | Descriptor: | COENZYME A, Formyl-coenzyme A transferase | | Authors: | Ricagno, S, Jonsson, S, Richards, N.G, Lindqvist, Y. | | Deposit date: | 2004-04-28 | | Release date: | 2004-08-03 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Kinetic and mechanistic characterization of the formyl-CoA transferase from Oxalobacter formigenes
J.Biol.Chem., 279, 2004
|
|
1VLR
 
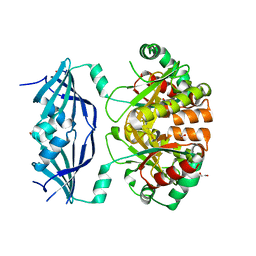 | |
1CZ8
 
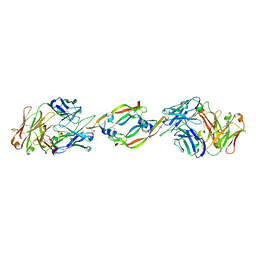 | | VASCULAR ENDOTHELIAL GROWTH FACTOR IN COMPLEX WITH AN AFFINITY MATURED ANTIBODY | | Descriptor: | HEAVY CHAIN OF NEUTRALIZING ANTIBODY, LIGHT CHAIN OF NEUTRALIZING ANTIBODY, SULFATE ION, ... | | Authors: | Chen, Y, Wiesmann, C, Fuh, G, Li, B, Christinger, H.W, McKay, P, de Vos, A.M, Lowman, H.B. | | Deposit date: | 1999-09-01 | | Release date: | 2000-03-20 | | Last modified: | 2017-04-05 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Selection and analysis of an optimized anti-VEGF antibody: crystal structure of an affinity-matured Fab in complex with antigen.
J.Mol.Biol., 293, 1999
|
|
1E3L
 
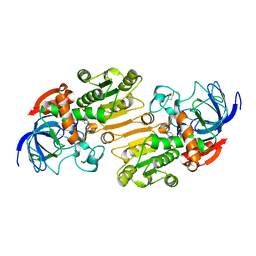 | | P47H mutant of mouse class II alcohol dehydrogenase complex with NADH | | Descriptor: | ALCOHOL DEHYDROGENASE, CLASS II, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Svensson, S, Hoog, J.O, Schneider, G, Sandalova, T. | | Deposit date: | 2000-06-19 | | Release date: | 2000-09-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of Mouse Class II Alcohol Dehydrogenase Reveal Determinants of Substrate Specificity and Catalytic Efficiency
J.Mol.Biol., 302, 2000
|
|
1E3E
 
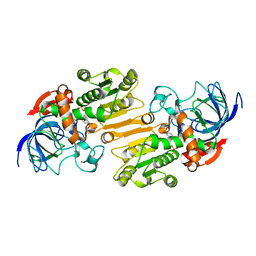 | | Mouse class II alcohol dehydrogenase complex with NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, ALCOHOL DEHYDROGENASE, CLASS II, ... | | Authors: | Svensson, S, Hoeoeg, J.O, Schneider, G, Sandalova, T. | | Deposit date: | 2000-06-14 | | Release date: | 2000-09-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Crystal Structure of Mouse Class II Alcohol Dehydrogenase Reveal Determinants of Substrate Specificity and Catalytic Efficiency
J.Mol.Biol., 302, 2000
|
|
1E3I
 
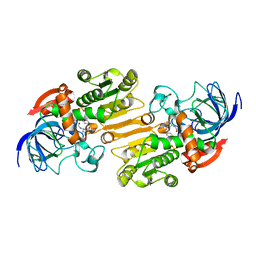 | | Mouse class II alcohol dehydrogenase complex with NADH and inhibitor | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, ALCOHOL DEHYDROGENASE, CLASS II, ... | | Authors: | Svensson, S, Hoog, J.O, Schneider, G, Sandalova, T. | | Deposit date: | 2000-06-16 | | Release date: | 2000-09-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Crystal Structure of Mouse Class II Alcohol Dehydrogenase Reveal Determinants of Substrate Specificity and Catalytic Efficiency
J.Mol.Biol., 302, 2000
|
|
1DLR
 
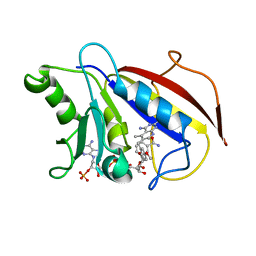 | | METHOTREXATE-RESISTANT VARIANTS OF HUMAN DIHYDROFOLATE REDUCTASE WITH SUBSTITUTION OF LEUCINE 22: KINETICS, CRYSTALLOGRAPHY AND POTENTIAL AS SELECTABLE MARKERS | | Descriptor: | 6-(2,5-DIMETHOXY-BENZYL)-5-METHYL-PYRIDO[2,3-D]PYRIMIDINE-2,4-DIAMINE, DIHYDROFOLATE REDUCTASE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Cody, V, Galitsky, N, Luft, J.R, Pangborn, W. | | Deposit date: | 1995-01-25 | | Release date: | 1995-04-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Methotrexate-resistant variants of human dihydrofolate reductase with substitutions of leucine 22. Kinetics, crystallography, and potential as selectable markers.
J.Biol.Chem., 270, 1995
|
|
1DEH
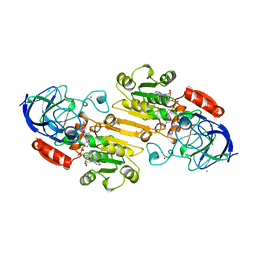 
 | | CRYSTALLIZATION OF HUMAN BETA1 ALCOHOL DEHYDROGENASE (15 MG/ML) IN 50 MM SODIUM PHOSPHATE (PH 7.5), 2.0 MM NAD+ AND 1 MM 4-IODOPYRAZOLE AT 25 OC, 13% (W/V) PEG 8000 | | Descriptor: | 4-IODOPYRAZOLE, CHLORIDE ION, HUMAN BETA1 ALCOHOL DEHYDROGENASE, ... | | Authors: | Hurley, T.D, Davis, G.J. | | Deposit date: | 1995-10-14 | | Release date: | 1996-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray structure of human beta3beta3 alcohol dehydrogenase. The contribution of ionic interactions to coenzyme binding.
J.Biol.Chem., 271, 1996
|
|
1DLS
 
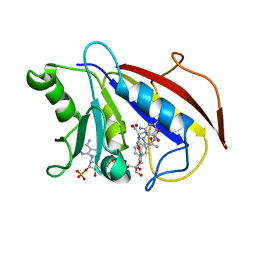 | | METHOTREXATE-RESISTANT VARIANTS OF HUMAN DIHYDROFOLATE REDUCTASE WITH SUBSTITUTION OF LEUCINE 22: KINETICS, CRYSTALLOGRAPHY AND POTENTIAL AS SELECTABLE MARKERS | | Descriptor: | DIHYDROFOLATE REDUCTASE, METHOTREXATE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Cody, V, Galitsky, N, Luft, J.R, Pangborn, W. | | Deposit date: | 1995-01-25 | | Release date: | 1995-04-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Methotrexate-resistant variants of human dihydrofolate reductase with substitutions of leucine 22. Kinetics, crystallography, and potential as selectable markers.
J.Biol.Chem., 270, 1995
|
|
1VEH
 
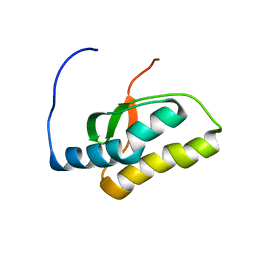 | | Solution structure of RSGI RUH-018, a NifU-like domain of hirip5 protein from mouse cDNA | | Descriptor: | NifU-like protein HIRIP5 | | Authors: | Ohashi, W, Hirota, H, Yamazaki, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-03-31 | | Release date: | 2004-09-30 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RSGI RUH-018, a NifU-like domain of hirip5 protein from mouse cDNA
To be published
|
|
