5AV0
 
 | |
1DGV
 
 | |
3UIT
 
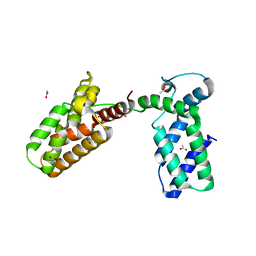 | | Overall structure of Patj/Pals1/Mals complex | | Descriptor: | ACETATE ION, InaD-like protein, MAGUK p55 subfamily member 5, ... | | Authors: | Zhang, J, Yang, X, Long, J, Shen, Y. | | Deposit date: | 2011-11-06 | | Release date: | 2012-02-22 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure of an L27 domain heterotrimer from cell polarity complex Patj/Pals1/Mals2 reveals mutually independent L27 domain assembly mode
J.Biol.Chem., 287, 2012
|
|
6TZ8
 
 | |
6TZ6
 
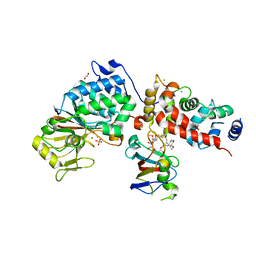 | | Crystal Structure of Candida Albicans Calcineurin A, Calcineurin B, FKBP12 and FK506 (Tacrolimus) | | Descriptor: | 1,2-ETHANEDIOL, 8-DEETHYL-8-[BUT-3-ENYL]-ASCOMYCIN, CALCIUM ION, ... | | Authors: | Fox III, D, Lukacs, C.M. | | Deposit date: | 2019-08-10 | | Release date: | 2019-09-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Harnessing calcineurin-FK506-FKBP12 crystal structures from invasive fungal pathogens to develop antifungal agents.
Nat Commun, 10, 2019
|
|
2RMO
 
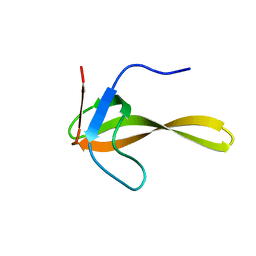 | | Solution structure of alpha-spectrin_SH3-bergerac from Chicken | | Descriptor: | Spectrin alpha chain, brain | | Authors: | Kutyshenko, V.P, Prokhorov, D.A, Timchenko, M.A, Kudrevatykh, Y.A, Fedyukina, D.V, Gushchina, L.V, Khristoforov, V.S, Filimonov, V.V. | | Deposit date: | 2007-11-07 | | Release date: | 2008-09-30 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Study of the structure and dynamics of a chimeric variant of the SH3 domain (SHA-Bergerac) by NMR spectroscopy
Russ.J.Bioorganic Chem., 34, 2008
|
|
2ROT
 
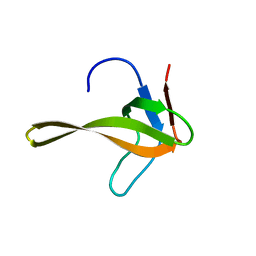 | | Structure of chimeric variant of SH3 domain- SHH | | Descriptor: | Spectrin alpha chain, brain | | Authors: | Kutyshenko, N.P, Prokhorov, D.A, Timchenko, M.A, Kudrevatykh, Y.A, Gushchina, L.V, Khristoforov, V.S, Filimonov, V.V. | | Deposit date: | 2008-04-10 | | Release date: | 2009-04-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the chimeric SH3 domains, SHH- and SHA-"Bergeracs".
Biochim.Biophys.Acta, 1794, 2009
|
|
2BHJ
 
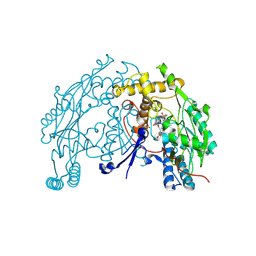 | | murine iNO synthase with coumarin inhibitor | | Descriptor: | 7,8-DIHYDROBIOPTERIN, NITRIC OXIDE SYNTHASE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Mathieu, M, Guilloteau, J.P. | | Deposit date: | 2005-01-12 | | Release date: | 2005-03-31 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Design, Synthesis and Characterization of a Novel Class of Coumarin-Based Inhibitors of Inducible Nitric Oxide Synthase
Bioorg.Med.Chem., 13, 2005
|
|
2RQ8
 
 | | Solution NMR structure of titin I27 domain mutant | | Descriptor: | Titin | | Authors: | Yagawa, K, Oguro, T, Momose, T, Kawano, S, Sato, T, Endo, T. | | Deposit date: | 2009-03-05 | | Release date: | 2010-02-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural basis for unfolding pathway-dependent stability of proteins: Vectorial unfolding vs. global unfolding
Protein Sci., 2010
|
|
1PSB
 
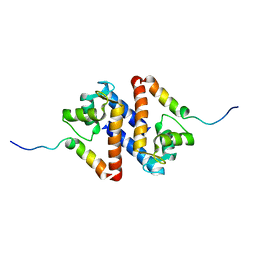 | | Solution structure of calcium loaded S100B complexed to a peptide from N-Terminal regulatory domain of NDR kinase. | | Descriptor: | Ndr Ser/Thr kinase-like protein, S-100 protein, beta chain | | Authors: | Bhattacharya, S, Large, E, Heizmann, C.W, Hemmings, B, Chazin, W.J. | | Deposit date: | 2003-06-21 | | Release date: | 2003-12-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the Ca(2+)/S100B/NDR Kinase Peptide Complex: Insights into S100 Target Specificity and Activation of the Kinase.
Biochemistry, 42, 2003
|
|
1RGW
 
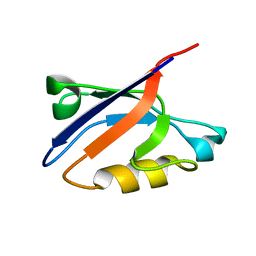 | | Solution Structure of ZASP's PDZ domain | | Descriptor: | ZASP protein | | Authors: | Au, Y, Atkinson, R.A, Pallavicini, A, Joseph, C, Martin, S.R, Muskett, F.W, Guerrini, R, Faulkner, G, Pastore, A. | | Deposit date: | 2003-11-13 | | Release date: | 2004-04-13 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of ZASP PDZ Domain; Implications for Sarcomere Ultrastructure and Enigma Family Redundancy.
Structure, 12, 2004
|
|
1ZL8
 
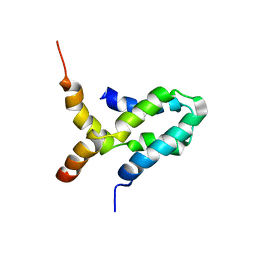 | | NMR structure of L27 heterodimer from C. elegans Lin-7 and H. sapiens Lin-2 scaffold proteins | | Descriptor: | LIN-7, Peripheral plasma membrane protein CASK | | Authors: | Petrosky, K.Y, Ou, H.D, Lohr, F, Dotsch, V, Lim, W.A. | | Deposit date: | 2005-05-05 | | Release date: | 2005-09-13 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A General Model for Preferential Hetero-oligomerization of LIN-2/7 Domains: Mechanism Underlying Directed Assembly of Supramolecular Signaling Complexes
J.Biol.Chem., 280, 2005
|
|
2V26
 
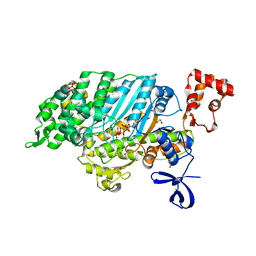 | | Myosin VI (MD) pre-powerstroke state (Mg.ADP.VO4) | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Menetrey, J, Llinas, P, Mukherjea, M, Sweeney, H.L, Houdusse, A. | | Deposit date: | 2007-06-03 | | Release date: | 2007-11-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Structural Basis for the Large Powerstroke of Myosin Vi.
Cell(Cambridge,Mass.), 131, 2007
|
|
1SRA
 
 | | STRUCTURE OF A NOVEL EXTRACELLULAR CA2+-BINDING MODULE IN BM-40(SLASH)SPARC(SLASH)OSTEONECTIN | | Descriptor: | CALCIUM ION, SPARC | | Authors: | Hohenester, E, Maurer, P, Hohenadl, C, Timpl, R, Jansonius, J.N, Engel, J. | | Deposit date: | 1995-08-21 | | Release date: | 1996-03-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of a novel extracellular Ca(2+)-binding module in BM-40.
Nat.Struct.Biol., 3, 1996
|
|
1SYM
 
 | |
3O96
 
 | | Crystal Structure of Human AKT1 with an Allosteric Inhibitor | | Descriptor: | 1-(1-(4-(7-phenyl-1H-imidazo[4,5-g]quinoxalin-6-yl)benzyl)piperidin-4-yl)-1H-benzo[d]imidazol-2(3H)-one, RAC-alpha serine/threonine-protein kinase | | Authors: | Voegtli, W.C, Wu, W.-I, Lord-Ondash, H.A, Dizon, F.P, Vigers, G.P.A, Brandhuber, B.J. | | Deposit date: | 2010-08-03 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of human AKT1 with an allosteric inhibitor reveals a new mode of kinase inhibition.
Plos One, 5, 2010
|
|
4IK5
 
 | | High resolution structure of Delta-REST-GCaMP3 | | Descriptor: | CALCIUM ION, RCaMP, Green fluorescent protein | | Authors: | Chen, Y, Song, X, Miao, L, Zhu, Y, Ji, G. | | Deposit date: | 2012-12-25 | | Release date: | 2014-01-29 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insight into enhanced calcium indicator GCaMP3 and GCaMPJ to promote further improvement.
Protein Cell, 4, 2013
|
|
4IK1
 
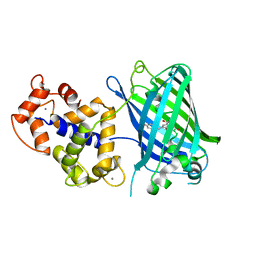 | | High resolution structure of GCaMPJ at pH 8.5 | | Descriptor: | CALCIUM ION, RCaMP, Green fluorescent protein | | Authors: | Chen, Y, Song, X, Miao, L, Zhu, Y, Ji, G. | | Deposit date: | 2012-12-25 | | Release date: | 2014-02-05 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insight into enhanced calcium indicator GCaMP3 and GCaMPJ to promote further improvement.
Protein Cell, 4, 2013
|
|
4IK4
 | | High resolution structure of GCaMP3 at pH 5.0 | | Descriptor: | CALCIUM ION, RCaMP, Green fluorescent protein | | Authors: | Chen, Y, Song, X, Miao, L, Zhu, Y, Ji, G. | | Deposit date: | 2012-12-25 | | Release date: | 2014-02-05 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Structural insight into enhanced calcium indicator GCaMP3 and GCaMPJ to promote further improvement.
Protein Cell, 4, 2013
|
|
6FSA
 
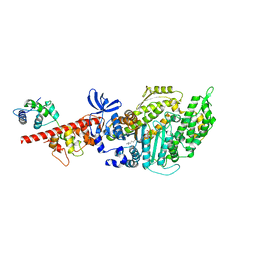 | | Beta-Cardiac myosin post-rigor | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, FORMIC ACID, ... | | Authors: | Robert-Paganin, J, Auguin, D, Houdusse, A. | | Deposit date: | 2018-02-19 | | Release date: | 2018-07-25 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Hypertrophic cardiomyopathy disease results from disparate impairments of cardiac myosin function and auto-inhibition.
Nat Commun, 9, 2018
|
|
4IK3
 
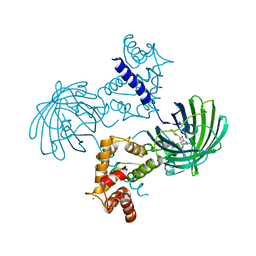 | | High resolution structure of GCaMP3 at pH 8.5 | | Descriptor: | CALCIUM ION, RCaMP, Green fluorescent protein | | Authors: | Chen, Y, Song, X, Miao, L, Zhu, Y, Ji, G. | | Deposit date: | 2012-12-25 | | Release date: | 2014-01-29 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (2.007 Å) | | Cite: | Structural insight into enhanced calcium indicator GCaMP3 and GCaMPJ to promote further improvement.
Protein Cell, 4, 2013
|
|
4IK9
 
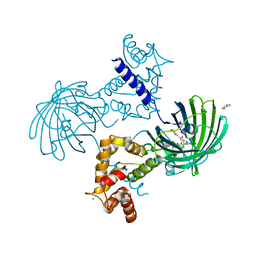 | | High resolution structure of GCaMP3 dimer form 2 at pH 7.5 | | Descriptor: | CALCIUM ION, DI(HYDROXYETHYL)ETHER, RCaMP, ... | | Authors: | Chen, Y, Song, X, Miao, L, Zhu, Y, Ji, G. | | Deposit date: | 2012-12-25 | | Release date: | 2014-01-29 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural insight into enhanced calcium indicator GCaMP3 and GCaMPJ to promote further improvement.
Protein Cell, 4, 2013
|
|
4IK8
 
 | | High resolution structure of GCaMP3 dimer form 1 at pH 7.5 | | Descriptor: | CALCIUM ION, RCaMP, Green fluorescent protein | | Authors: | Chen, Y, Song, X, Miao, L, Zhu, Y, Ji, G. | | Deposit date: | 2012-12-25 | | Release date: | 2014-02-05 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural insight into enhanced calcium indicator GCaMP3 and GCaMPJ to promote further improvement.
Protein Cell, 4, 2013
|
|
6HHF
 
 | | Crystal Structure of AKT1 in Complex with Covalent-Allosteric AKT Inhibitor Borussertib | | Descriptor: | Borussertib, RAC-alpha serine/threonine-protein kinase | | Authors: | Landel, I, Weisner, J, Mueller, M.P, Scheinpflug, R, Rauh, D. | | Deposit date: | 2018-08-28 | | Release date: | 2019-03-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Preclinical Efficacy of Covalent-Allosteric AKT Inhibitor Borussertib in Combination with Trametinib inKRAS-Mutant Pancreatic and Colorectal Cancer.
Cancer Res., 79, 2019
|
|
6HHI
 
 | | Crystal Structure of AKT1 in Complex with Covalent-Allosteric AKT Inhibitor 30b | | Descriptor: | RAC-alpha serine/threonine-protein kinase, ~{N}-[1-[[4-(5-oxidanylidene-3-phenyl-6~{H}-1,6-naphthyridin-2-yl)phenyl]methyl]piperidin-4-yl]-3-(propanoylamino)benzamide | | Authors: | Landel, I, Weisner, J, Mueller, M.P, Scheinpflug, R, Rauh, D. | | Deposit date: | 2018-08-28 | | Release date: | 2019-02-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural and chemical insights into the covalent-allosteric inhibition of the protein kinase Akt.
Chem Sci, 10, 2019
|
|
