9C96
 | | Cryo-EM structure of TAP binding protein related (TAPBPR) in complex with HLA-A*02:01 bound to a suboptimal peptide. | | Descriptor: | Beta-2-microglobulin, LYS-ILE-LEU-GLY-PHE-VAL, MHC class I antigen, ... | | Authors: | Pumroy, R.P, Mallik, L, Sun, Y, Moiseenkova-Bell, Y.V, Sgourakis, N.G. | | Deposit date: | 2024-06-13 | | Release date: | 2025-01-22 | | Last modified: | 2025-01-29 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | CryoEM structure of an MHC-I/TAPBPR peptide-bound intermediate reveals the mechanism of antigen proofreading.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
9C42
 
 | | Structure of human MR1-ellagic acid in complex with human MAIT A-F7 TCR | | Descriptor: | 2,3,7,8-tetrahydroxychromeno[5,4,3-cde]chromene-5,10-dione, ACETIC ACID, Beta-2-microglobulin, ... | | Authors: | Wang, C.J.H, Le Nours, J, Rossjohn, J. | | Deposit date: | 2024-06-02 | | Release date: | 2025-06-11 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Ellagic acid metabolism as a source of dietary MR1 ligands
To Be Published
|
|
9HI7
 
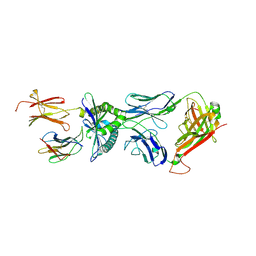 | | Structure of MC.7.G5 T cell receptor in complex with MR1 R9H | | Descriptor: | Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, T cell receptor alpha chain MC.7.G5, ... | | Authors: | Karuppiah, V. | | Deposit date: | 2024-11-24 | | Release date: | 2025-04-23 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Molecular basis underpinning MR1 allomorph recognition by an MR1-restricted T cell receptor.
Front Immunol, 16, 2025
|
|
9IIK
 
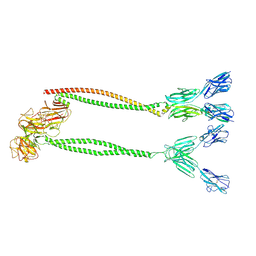 | | Cryo-EM structure of BTN2A1-BTN3A1-BTN3A2 | | Descriptor: | (2E)-4-hydroxy-3-methylbut-2-en-1-yl trihydrogen diphosphate, Butyrophilin subfamily 2 member A1, Butyrophilin subfamily 3 member A1, ... | | Authors: | Gao, W, Zheng, J, Zhu, Y, Huang, Z. | | Deposit date: | 2024-06-20 | | Release date: | 2025-06-25 | | Method: | ELECTRON MICROSCOPY (4.12 Å) | | Cite: | Cryo-EM structure of BTN2A1-BTN3A1-BTN3A2
To Be Published
|
|
9GH6
 
 | | CEACAM1 (35-234), focused refinement in the complex with CbpF | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Carcinoembryonic antigen-related cell adhesion molecule 1 | | Authors: | Marongiu, G.L, Fink, U, Roderer, D. | | Deposit date: | 2024-08-15 | | Release date: | 2025-04-02 | | Last modified: | 2025-04-23 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural basis for immune cell binding of Fusobacterium nucleatum via the trimeric autotransporter adhesin CbpF.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
9GH5
 | | Complex of Fusobacterium nucleatum CbpF with human CEACAM1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Carcinoembryonic antigen-related cell adhesion molecule 1, Cell surface protein | | Authors: | Marongiu, G.L, Fink, U, Roderer, D. | | Deposit date: | 2024-08-15 | | Release date: | 2025-04-02 | | Last modified: | 2025-04-23 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Structural basis for immune cell binding of Fusobacterium nucleatum via the trimeric autotransporter adhesin CbpF.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
9DFF
 
 | | G1 domain of human aggrecan bound to hyaluronan | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-deoxy-alpha-L-threo-hex-4-enopyranuronic acid-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-beta-D-glucopyranuronic acid-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-beta-D-glucopyranuronic acid-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-beta-D-glucopyranuronic acid-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-beta-D-glucopyranuronic acid-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose, Aggrecan core protein | | Authors: | Bouyain, S. | | Deposit date: | 2024-08-29 | | Release date: | 2025-04-30 | | Last modified: | 2025-06-11 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Aggrecan immobilizes to perineuronal nets through hyaluronan-dependent and hyaluronan-independent binding activities.
J.Biol.Chem., 301, 2025
|
|
9DFT
 
 | | G1 domain of human aggrecan | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Aggrecan core protein, SULFATE ION | | Authors: | Bouyains, S. | | Deposit date: | 2024-08-30 | | Release date: | 2025-04-30 | | Last modified: | 2025-06-11 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Aggrecan immobilizes to perineuronal nets through hyaluronan-dependent and hyaluronan-independent binding activities.
J.Biol.Chem., 301, 2025
|
|
9C3E
 
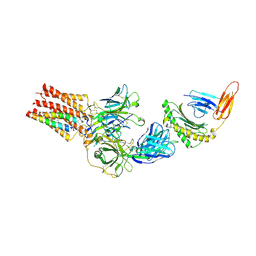 | | TCR - CD3 complex bound to HLA | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHOLESTEROL, Cancer/testis antigen 1,Beta-2-microglobulin,MHC class I antigen, ... | | Authors: | Notti, R.Q, Walz, T. | | Deposit date: | 2024-05-31 | | Release date: | 2024-10-02 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | The resting and ligand-bound states of the membrane-embedded human T-cell receptor-CD3 complex.
Biorxiv, 2024
|
|
9J5J
 
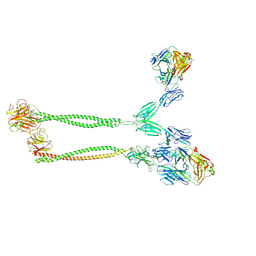 | | Cryo-EM structure of BTN2A1-BTN3A1-BTN3A2 in complex with gdTCR | | Descriptor: | (2E)-4-hydroxy-3-methylbut-2-en-1-yl trihydrogen diphosphate, Butyrophilin subfamily 2 member A1, Butyrophilin subfamily 3 member A1, ... | | Authors: | Zhu, Y, Gao, W, Huang, Z. | | Deposit date: | 2024-08-12 | | Release date: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (4.05 Å) | | Cite: | Phosphoantigen-induced inside-out stabilization of butyrophilin receptor complexes drives dimerization-dependent gamma delta TCR activation.
Immunity, 2025
|
|
9J5M
 
 | |
7RNO
 
 | | Model of the Ac-6-FP/hpMR1/bB2m/TAPBPR complex from integrated docking, NMR and restrained MD | | Descriptor: | Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, N-(6-formyl-4-oxo-3,4-dihydropteridin-2-yl)acetamide, ... | | Authors: | McShan, A.C, Sgourakis, N.G. | | Deposit date: | 2021-07-29 | | Release date: | 2022-05-11 | | Last modified: | 2022-08-10 | | Method: | SOLUTION NMR | | Cite: | TAPBPR employs a ligand-independent docking mechanism to chaperone MR1 molecules.
Nat.Chem.Biol., 18, 2022
|
|
7RYN
 
 | | CD1a-sulfatide-gdTCR complex | | Descriptor: | (15Z)-N-((1S,2R,3E)-2-HYDROXY-1-{[(3-O-SULFO-BETA-D-GALACTOPYRANOSYL)OXY]METHYL}HEPTADEC-3-ENYL)TETRACOS-15-ENAMIDE, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Wegrecki, M, Le Nours, J, Rossjohn, J. | | Deposit date: | 2021-08-25 | | Release date: | 2022-05-18 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Atypical sideways recognition of CD1a by autoreactive gamma delta T cell receptors.
Nat Commun, 13, 2022
|
|
5MTL
 
 | | Crystal structure of an amyloidogenic light chain | | Descriptor: | light chain dimer,IGL@ protein,IGL@ protein | | Authors: | Oberti, L, Rognoni, P, Russo, R, Bacarizo, J, Bolognesi, M, Ricagno, S. | | Deposit date: | 2017-01-10 | | Release date: | 2017-12-13 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Concurrent structural and biophysical traits link with immunoglobulin light chains amyloid propensity.
Sci Rep, 7, 2017
|
|
7RYO
 
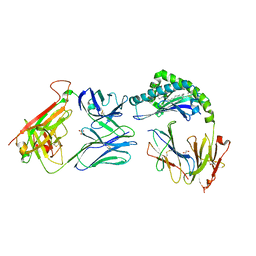 | | CD1a-dideoxymycobactin-gdTCR complex | | Descriptor: | 1,2-ETHANEDIOL, Beta-2-microglobulin, SULFATE ION, ... | | Authors: | Wegrecki, M, Le Nours, J, Rossjohn, J. | | Deposit date: | 2021-08-25 | | Release date: | 2022-05-18 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Atypical sideways recognition of CD1a by autoreactive gamma delta T cell receptors.
Nat Commun, 13, 2022
|
|
7RYL
 
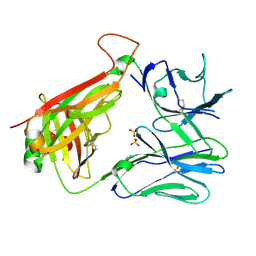 | | T cell receptor CO3 | | Descriptor: | 1,2-ETHANEDIOL, SULFATE ION, T cell receptor delta variable 1,T cell receptor alpha chain constant, ... | | Authors: | Wegrecki, M, Le Nours, J, Rossjohn, J. | | Deposit date: | 2021-08-25 | | Release date: | 2022-05-18 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Atypical sideways recognition of CD1a by autoreactive gamma delta T cell receptors.
Nat Commun, 13, 2022
|
|
7RYM
 
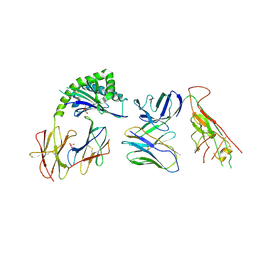 | | CD1a-endo-gdTCR complex | | Descriptor: | Beta-2-microglobulin, DI(HYDROXYETHYL)ETHER, SULFATE ION, ... | | Authors: | Wegrecki, M, Le Nours, J, Rossjohn, J. | | Deposit date: | 2021-08-25 | | Release date: | 2022-05-18 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Atypical sideways recognition of CD1a by autoreactive gamma delta T cell receptors.
Nat Commun, 13, 2022
|
|
8TW4
 
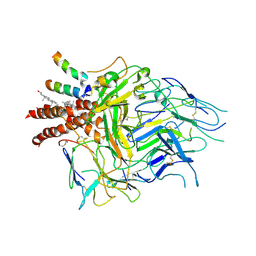 | | TCR in nanodisc ND-I | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHOLESTEROL, ... | | Authors: | Notti, R.Q, Walz, T. | | Deposit date: | 2023-08-20 | | Release date: | 2024-10-02 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | The resting and ligand-bound states of the membrane-embedded human T-cell receptor-CD3 complex.
Biorxiv, 2024
|
|
8TW6
 
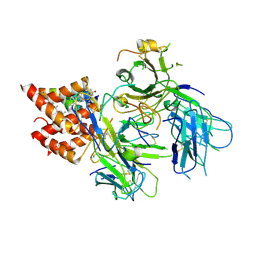 | | TCR in nanodisc ND-II | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHOLESTEROL, T cell receptor beta variable 6-5,T cell receptor beta chain MC.7.G5,MCHERRY, ... | | Authors: | Notti, R.Q, Walz, T. | | Deposit date: | 2023-08-20 | | Release date: | 2024-10-02 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | The resting and ligand-bound states of the membrane-embedded human T-cell receptor-CD3 complex.
Biorxiv, 2024
|
|
5O1R
 | |
5NHT
 
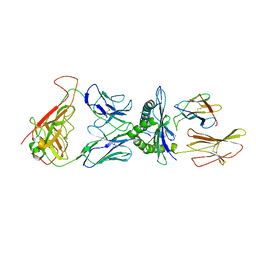 | | human 199.54-16 TCR in complex with Melan-A/MART-1 (26-35) peptide and HLA-A2 | | Descriptor: | Beta-2-microglobulin, CHLORIDE ION, HLA class I histocompatibility antigen, ... | | Authors: | Exertier, C, Reiser, J.-B, Lantez, V, Chouquet, A, Bonneville, M, Saulquin, X, Housset, D. | | Deposit date: | 2017-03-22 | | Release date: | 2018-05-16 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | human 199.54-16 TCR in complex with Melan-A/MART-1 (26-35) peptide and HLA-A2
To Be Published
|
|
5OR7
 
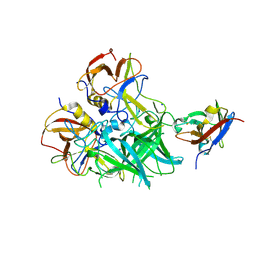 | |
4BFI
 
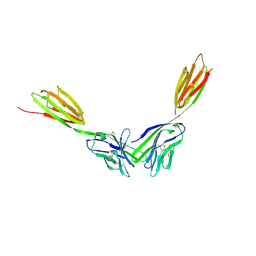 | | Structure of the complex of the extracellular portions of mouse CD200R and mouse CD200 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, CELL SURFACE GLYCOPROTEIN CD200 RECEPTOR 1, ... | | Authors: | Hatherley, D, Lea, S.M, Johnson, S, Barclay, A.N. | | Deposit date: | 2013-03-19 | | Release date: | 2013-05-01 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.22 Å) | | Cite: | Structures of Cd200/Cd200 Receptor Family and Implications for Topology, Regulation, and Evolution
Structure, 21, 2013
|
|
5OPI
 
 | | Crystal structure of the TAPBPR-MHC I peptide editing complex | | Descriptor: | Beta-2-microglobulin, H-2 class I histocompatibility antigen, D-B alpha chain, ... | | Authors: | Thomas, C, Tampe, R. | | Deposit date: | 2017-08-09 | | Release date: | 2017-10-18 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of the TAPBPR-MHC I complex defines the mechanism of peptide loading and editing.
Science, 358, 2017
|
|
7TY0
 
 | |
