3Q1X
 
 | |
5ZEP
 
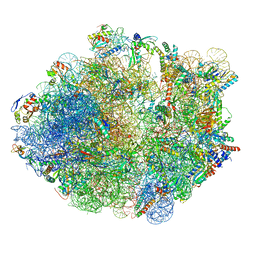 | | M. smegmatis hibernating state 70S ribosome structure | | Descriptor: | 16S rRNA, 23S rRNA, 30S ribosomal protein S10, ... | | Authors: | Mishra, S, Ahmed, T, Tyagi, A, Shi, J, Bhushan, S. | | Deposit date: | 2018-02-27 | | Release date: | 2018-09-26 | | Last modified: | 2025-09-17 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structures of Mycobacterium smegmatis 70S ribosomes in complex with HPF, tmRNA, and P-tRNA.
Sci Rep, 8, 2018
|
|
7F2D
 
 | |
7F2I
 
 | |
1PWA
 
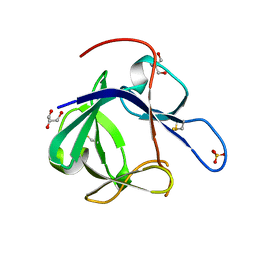 | | Crystal structure of Fibroblast Growth Factor 19 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Fibroblast growth factor-19, GLYCEROL, ... | | Authors: | Harmer, N.J, Pellegrini, L, Chirgadze, D, Fernandez-Recio, J, Blundell, T.L. | | Deposit date: | 2003-07-01 | | Release date: | 2004-01-27 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The crystal structure of fibroblast growth factor (FGF) 19 reveals novel features of the FGF family and offers a structural basis for its unusual receptor affinity.
Biochemistry, 43, 2004
|
|
1WCW
 | | Crystal Structure of Uroporphyrinogen III Synthase from an Extremely Thermophilic Bacterium Thermus thermophilus HB8 (Wild type, Native, Form-1 crystal) | | Descriptor: | GLYCEROL, Uroporphyrinogen III synthase | | Authors: | Mizohata, E, Matsuura, T, Sakai, H, Murayama, K, Terada, T, Shirouzu, M, Kuramitsu, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-06 | | Release date: | 2005-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structure of Uroporphyrinogen III Synthase from Thermus thermophilus HB8
To be Published
|
|
1CBQ
 
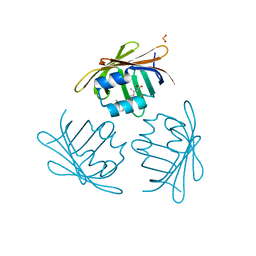 | | CRYSTAL STRUCTURE OF CELLULAR RETINOIC-ACID-BINDING PROTEINS I AND II IN COMPLEX WITH ALL-TRANS-RETINOIC ACID AND A SYNTHETIC RETINOID | | Descriptor: | 6-(2,3,4,5,6,7-HEXAHYDRO-2,4,4-TRIMETHYL-1-METYLENEINDEN-2-YL)-3-METHYLHEXA-2,4-DIENOIC ACID, CELLULAR RETINOIC ACID BINDING PROTEIN TYPE II, PHOSPHATE ION | | Authors: | Kleywegt, G.J, Bergfors, T, Jones, T.A. | | Deposit date: | 1994-09-28 | | Release date: | 1995-01-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of cellular retinoic acid binding proteins I and II in complex with all-trans-retinoic acid and a synthetic retinoid.
Structure, 2, 1994
|
|
5CAS
 
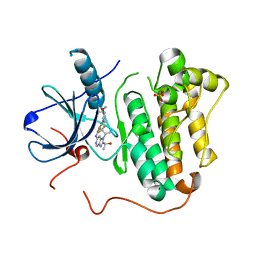 | | EGFR kinase domain mutant "TMLR" with compound 41a | | Descriptor: | (1R)-1-{6-({2-[(3R,4S)-3-fluoro-4-methoxypiperidin-1-yl]pyrimidin-4-yl}amino)-1-[(2S)-1,1,1-trifluoropropan-2-yl]-1H-imidazo[4,5-c]pyridin-2-yl}ethanol, Epidermal growth factor receptor, SULFATE ION | | Authors: | Eigenbrot, C, Yu, C. | | Deposit date: | 2015-06-29 | | Release date: | 2015-10-28 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Noncovalent Mutant Selective Epidermal Growth Factor Receptor Inhibitors: A Lead Optimization Case Study.
J.Med.Chem., 58, 2015
|
|
4RN2
 
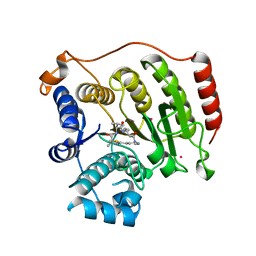 | | Crystal structure of S39D HDAC8 in complex with a largazole analogue. | | Descriptor: | (5R,8S,11S)-5-methyl-8-(propan-2-yl)-11-[(1E)-4-sulfanylbut-1-en-1-yl]-3-thia-7,10,14,17,21-pentaazatricyclo[14.3.1.1~2,5~]henicosa-1(20),2(21),16,18-tetraene-6,9,13-trione, Histone deacetylase 8, POTASSIUM ION, ... | | Authors: | Decroos, C, Christianson, D.W. | | Deposit date: | 2014-10-22 | | Release date: | 2015-04-08 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Variable Active Site Loop Conformations Accommodate the Binding of Macrocyclic Largazole Analogues to HDAC8.
Biochemistry, 54, 2015
|
|
2YIS
 
 | | triazolopyridine inhibitors of p38 kinase. | | Descriptor: | 1-[3-tert-butyl-1-(3-chloro-4-hydroxyphenyl)-1H-pyrazol-5-yl]-3-{2-[(3-{2-[(2-hydroxyethyl)sulfanyl]phenyl}[1,2,4]triazolo[4,3-a]pyridin-6-yl)sulfanyl]benzyl}urea, 2-fluoro-4-[4-(4-fluorophenyl)-1H-pyrazol-3-yl]pyridine, MITOGEN-ACTIVATED PROTEIN KINASE 14 | | Authors: | Millan, D.S, Anderson, M, Bazin, R, Bunnage, M.E, Burrows, J.L, Butcher, K.J, Dodd, P.G, Evans, T.J, Fairman, D.A, Hughes, S.J, Irving, S.L, Kilty, I.C, Lemaitre, A, Lewthwaite, R.A, Mahnke, A, Mathais, J.P, Philip, J, Phillips, C, Smith, R.T, Stefaniack, M.H, Yeadon, M. | | Deposit date: | 2011-05-16 | | Release date: | 2011-11-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design and Synthesis of Inhaled P38 Inhibitors for the Treatment of Chronic Obstructive Pulmonary Disease.
J.Med.Chem., 54, 2011
|
|
1BRY
 
 | | BRYODIN TYPE I RIP | | Descriptor: | BRYODIN I | | Authors: | Klei, H.E, Chang, C.Y. | | Deposit date: | 1997-02-14 | | Release date: | 1998-03-04 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Molecular, biological, and preliminary structural analysis of recombinant bryodin 1, a ribosome-inactivating protein from the plant Bryonia dioica.
Biochemistry, 36, 1997
|
|
4AIO
 
 | | Crystal structure of the starch debranching enzyme barley limit dextrinase | | Descriptor: | CALCIUM ION, GLYCEROL, IODIDE ION, ... | | Authors: | Moeller, M.S, Abou Hachem, M, Svensson, B, Henriksen, A. | | Deposit date: | 2012-02-11 | | Release date: | 2012-08-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the Starch-Debranching Enzyme Barley Limit Dextrinase Reveals Homology of the N-Terminal Domain to Cbm21.
Acta Crystallogr.,Sect.F, 68, 2012
|
|
1PW8
 
 | | Covalent Acyl Enzyme Complex Of The R61 DD-Peptidase with A Highly Specific Cephalosporin | | Descriptor: | (6R,7R)-3-[(ACETYLOXY)METHYL]-7-{[(6S)-6-(GLYCYLAMINO)-7-OXIDO-7-OXOHEPTANOYL]AMINO}-8-OXO-5-THIA-1-AZABICYCLO[4.2.0]OCTANE-2-CARBOXYLATE, D-alanyl-D-alanine carboxypeptidase, GLYCEROL | | Authors: | Silvaggi, N.R, Josephine, H.R, Pratt, R.F, Kelly, J.A. | | Deposit date: | 2003-07-01 | | Release date: | 2004-07-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal structures of complexes between the R61 DD-peptidase and peptidoglycan-mimetic beta-lactams: a non-covalent complex with a "perfect penicillin"
J.Mol.Biol., 345, 2005
|
|
2V3D
 
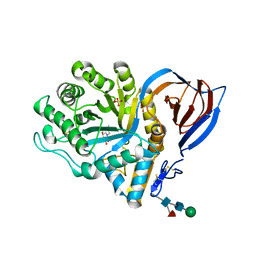 | | acid-beta-glucosidase with N-butyl-deoxynojirimycin | | Descriptor: | (2R,3R,4R,5S)-1-BUTYL-2-(HYDROXYMETHYL)PIPERIDINE-3,4,5-TRIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, GLUCOSYLCERAMIDASE, ... | | Authors: | Brumshtein, B, Greenblatt, H.M, Butters, T.D, Shaaltiel, Y, Aviezer, D, Silman, I, Futerman, A.H, Sussman, J.L. | | Deposit date: | 2007-06-17 | | Release date: | 2007-08-14 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Crystal Structures of Complexes of N-Butyl- and N-Nonyl-Deoxynojirimycin Bound to Acid Beta-Glucosidase: Insights Into the Mechanism of Chemical Chaperone Action in Gaucher Disease.
J.Biol.Chem., 282, 2007
|
|
1BWF
 
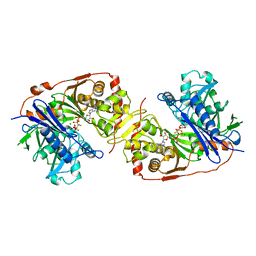 | | ESCHERICHIA COLI GLYCEROL KINASE MUTANT WITH BOUND ATP ANALOG SHOWING SUBSTANTIAL DOMAIN MOTION | | Descriptor: | GLYCEROL, GLYCEROL KINASE, MAGNESIUM ION, ... | | Authors: | Bystrom, C.E, Pettigrew, D.W, Branchaud, B.P, Remington, S.J. | | Deposit date: | 1998-09-23 | | Release date: | 1999-05-18 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structures of Escherichia coli glycerol kinase variant S58-->W in complex with nonhydrolyzable ATP analogues reveal a putative active conformation of the enzyme as a result of domain motion.
Biochemistry, 38, 1999
|
|
2LIG
 
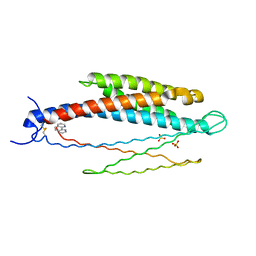 | | THREE-DIMENSIONAL STRUCTURES OF THE LIGAND-BINDING DOMAIN OF THE BACTERIAL ASPARTATE RECEPTOR WITH AND WITHOUT A LIGAND | | Descriptor: | 1,10-PHENANTHROLINE, ASPARTATE RECEPTOR, ASPARTIC ACID, ... | | Authors: | Kim, S.-H, Yeh, J.I, Prive, G.G, Milburn, M, Scott, W, Koshland Junior, D.E. | | Deposit date: | 1995-04-18 | | Release date: | 1995-09-15 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Three-dimensional structures of the ligand-binding domain of the bacterial aspartate receptor with and without a ligand.
Science, 254, 1991
|
|
6JGN
 
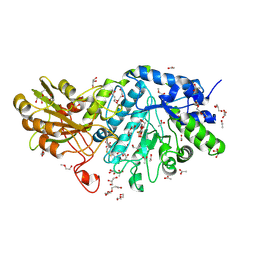 | | Crystal structure of barley exohydrolaseI W434H in complex with 4'-nitrophenyl thiolaminaribioside | | Descriptor: | 4'-NITROPHENYL-S-(BETA-D-GLUCOPYRANOSYL)-(1-3)-(3-THIO-BETA-D-GLUCOPYRANOSYL)-(1-3)-BETA-D-GLUCOPYRANOSIDE, ACETATE ION, BETA-D-GLUCAN GLUCOHYDROLASE ISOENZYME EXO1, ... | | Authors: | Luang, S, Streltsov, V.A, Hrmova, M. | | Deposit date: | 2019-02-14 | | Release date: | 2020-08-19 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | The evolutionary advantage of an aromatic clamp in plant family 3 glycoside exo-hydrolases.
Nat Commun, 13, 2022
|
|
4RUQ
 
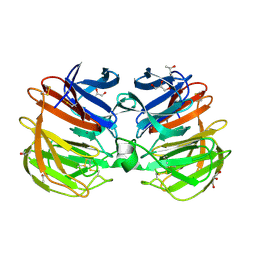 | | Carp Fishelectin, apo form | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Fish-egg lectin, ... | | Authors: | Capaldi, S, Faggion, B, Carrizo, M.E, Destefanis, L, Gonzalez, M.C, Perduca, M, Bovi, M, Galliano, M, Monaco, H.L. | | Deposit date: | 2014-11-21 | | Release date: | 2015-04-08 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Three-dimensional structure and ligand-binding site of carp fishelectin (FEL).
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4JVL
 
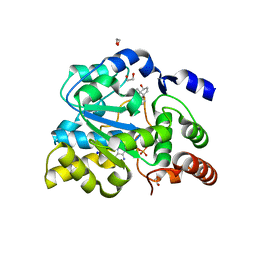 | | Crystal structure of human estrogen sulfotransferase (SULT1E1) in complex with inactive cofactor PAP and estradiol (E2) | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-3'-5'-DIPHOSPHATE, ESTRADIOL, ... | | Authors: | Gosavi, R.A, Knudsen, G.A, Birnbaum, L.S, Pedersen, L.C. | | Deposit date: | 2013-03-25 | | Release date: | 2013-09-04 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.943 Å) | | Cite: | Mimicking of Estradiol Binding by Flame Retardants and Their Metabolites: A Crystallographic Analysis.
Environ.Health Perspect., 121, 2013
|
|
4J8A
 
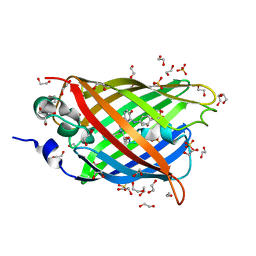 | | Irradiated-state structure of sfGFP containing the unnatural amino acid p-azido-phenylalanine at residue 145 | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Green fluorescent protein, ... | | Authors: | Reddington, S.C, Jones, D.D, Rizkallah, P.J, Tippmann, E.M. | | Deposit date: | 2013-02-14 | | Release date: | 2013-05-15 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.26 Å) | | Cite: | Different Photochemical Events of a Genetically Encoded Phenyl Azide Define and Modulate GFP Fluorescence.
Angew.Chem.Int.Ed.Engl., 52, 2013
|
|
3RLU
 
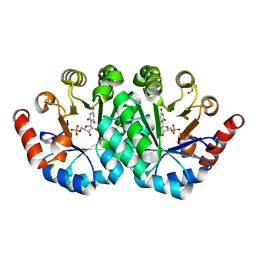 | | Crystal structure of the mutant K82A of orotidine 5'-monophosphate decarboxylase from Methanobacterium thermoautotrophicum complexed with the inhibitor BMP | | Descriptor: | 6-HYDROXYURIDINE-5'-PHOSPHATE, GLYCEROL, Orotidine 5'-phosphate decarboxylase | | Authors: | Fedorov, A.A, Fedorov, E.V, Desai, B, Gerlt, J.A, Almo, S.C. | | Deposit date: | 2011-04-20 | | Release date: | 2012-04-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Conformational changes in orotidine 5'-monophosphate decarboxylase: a structure-based explanation for how the 5'-phosphate group activates the enzyme.
Biochemistry, 51, 2012
|
|
1JZ2
 
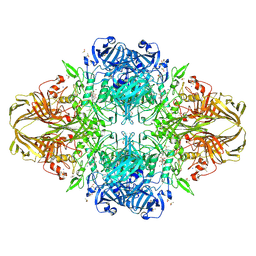 | | E. COLI (lacZ) BETA-GALACTOSIDASE-TRAPPED 2-F-GALACTOSYL-ENZYME INTERMEDIATE (ORTHORHOMBIC) | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 2-deoxy-2-fluoro-beta-D-galactopyranose, Beta-Galactosidase, ... | | Authors: | Juers, D.H, McCarter, J.D, Mackenzie, L, Withers, S.G, Matthews, B.W. | | Deposit date: | 2001-09-13 | | Release date: | 2001-12-07 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A Structural View of the Action of Escherichia Coli (Lacz) Beta-Galactosidase
Biochemistry, 40, 2001
|
|
4J88
 | | Dark-state structure of sfGFP containing the unnatural amino acid p-azido-phenylalanine at residue 66 | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Green fluorescent protein, ... | | Authors: | Reddington, S.C, Jones, D.D, Rizkallah, P.J, Tippmann, E.M. | | Deposit date: | 2013-02-14 | | Release date: | 2013-06-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Different Photochemical Events of a Genetically Encoded Phenyl Azide Define and Modulate GFP Fluorescence.
Angew.Chem.Int.Ed.Engl., 52, 2013
|
|
8D50
 
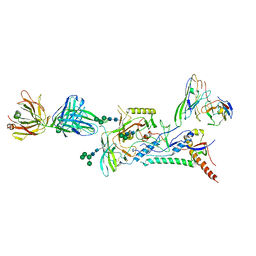 | |
4AW0
 
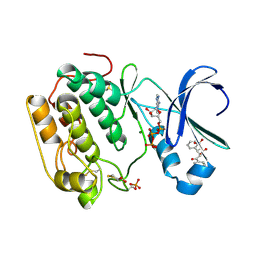 | | Human PDK1 Kinase Domain in Complex with Allosteric Compound PS182 Bound to the PIF-Pocket | | Descriptor: | 3-PHOSPHOINOSITIDE-DEPENDENT PROTEIN KINASE 1, ADENOSINE-5'-TRIPHOSPHATE, DIMETHYL SULFOXIDE, ... | | Authors: | Schulze, J.O, Busschots, K, Lopez-Garcia, L.A, Lammi, C, Stroba, A, Zeuzem, S, Piiper, A, Alzari, P.M, Neimanis, S, Arencibia, J.M, Engel, M, Biondi, R.M. | | Deposit date: | 2012-05-30 | | Release date: | 2012-10-03 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Substrate-Selective Inhibition of Protein Kinase Pdk1 by Small Compounds that Bind to the Pif-Pocket Allosteric Docking Site.
Chem.Biol., 19, 2012
|
|
