6CW9
 
 | | Structure of alpha-GC[8,16P] bound by CD1d and in complex with the Va14Vb8.2 TCR | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Wang, J, Zajonc, D. | | Deposit date: | 2018-03-30 | | Release date: | 2019-04-03 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A molecular switch in mouse CD1d modulates natural killer T cell activation by alpha-galactosylsphingamides.
J.Biol.Chem., 294, 2019
|
|
1IQH
 
 | | Human coagulation factor Xa in complex with M55143 | | Descriptor: | 4-[(6-CHLORO-2-NAPHTHALENYL)SULFONYL]-1-[[4-HYDROXYMETHYL-1-(4-PYRIDINYL)-4-PIPERIDINYL]METHYL]PIPERAZINONE, CALCIUM ION, coagulation Factor Xa | | Authors: | Shiromizu, I, Matsusue, T. | | Deposit date: | 2001-07-23 | | Release date: | 2003-09-23 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Factor Xa Specific Inhibitor that Induces the Novel Binding Model in Complex with Human Fxa
To be Published
|
|
3WXR
 
 | | Yeast 20S proteasome with a mutation of alpha7 subunit | | Descriptor: | Probable proteasome subunit alpha type-7, Proteasome subunit alpha type-1, Proteasome subunit alpha type-2, ... | | Authors: | Yashiroda, H, Toda, Y, Otsu, S, Takagi, K, Mizushima, T, Murata, S. | | Deposit date: | 2014-08-06 | | Release date: | 2014-10-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | N-terminal alpha 7 deletion of the proteasome 20S core particle substitutes for yeast PI31 function
Mol.Cell.Biol., 35, 2015
|
|
7GYN
 | | Crystal Structure of HSP72 in complex with ligand 11 at 7.15 MGy X-ray dose | | Descriptor: | 1,2-ETHANEDIOL, 3-PYRIDINIUM-1-YLPROPANE-1-SULFONATE, 8-chloroadenosine, ... | | Authors: | Cabry, M, Rodrigues, M.J, Le Bihan, Y.V, van Montfort, R.L.M. | | Deposit date: | 2024-01-12 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Specific radiation damage to halogenated inhibitors and ligands in protein-ligand crystal structures.
J.Appl.Crystallogr., 57, 2024
|
|
6JGR
 
 | | Crystal structure of barley exohydrolaseI W434Y in complex with 4'-nitrophenyl thiolaminaribioside | | Descriptor: | 4'-NITROPHENYL-S-(BETA-D-GLUCOPYRANOSYL)-(1-3)-(3-THIO-BETA-D-GLUCOPYRANOSYL)-(1-3)-BETA-D-GLUCOPYRANOSIDE, ACETATE ION, BETA-D-GLUCAN GLUCOHYDROLASE ISOENZYME EXO1, ... | | Authors: | Luang, S, Streltsov, V.A, Hrmova, M. | | Deposit date: | 2019-02-14 | | Release date: | 2020-08-19 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The evolutionary advantage of an aromatic clamp in plant family 3 glycoside exo-hydrolases.
Nat Commun, 13, 2022
|
|
1QRN
 
 | | CRYSTAL STRUCTURE OF HUMAN A6 TCR COMPLEXED WITH HLA-A2 BOUND TO ALTERED HTLV-1 TAX PEPTIDE P6A | | Descriptor: | BETA-2 MICROGLOBULIN, HLA CLASS I HISTOCOMPATIBILITY ANTIGEN, A-2 ALPHA CHAIN, ... | | Authors: | Ding, Y.H, Baker, B.M, Garboczi, D.N, Biddison, W.E, Wiley, D.C. | | Deposit date: | 1999-06-14 | | Release date: | 1999-10-14 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Four A6-TCR/peptide/HLA-A2 structures that generate very different T cell signals are nearly identical.
Immunity, 11, 1999
|
|
7GYP
 
 | | Crystal Structure of HSP72 in complex with ligand 11 at 10.01 MGy X-ray dose | | Descriptor: | 1,2-ETHANEDIOL, 3-PYRIDINIUM-1-YLPROPANE-1-SULFONATE, 8-chloroadenosine, ... | | Authors: | Cabry, M, Rodrigues, M.J, Le Bihan, Y.V, van Montfort, R.L.M. | | Deposit date: | 2024-01-12 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Specific radiation damage to halogenated inhibitors and ligands in protein-ligand crystal structures.
J.Appl.Crystallogr., 57, 2024
|
|
7GYX
 
 | | Crystal Structure of HSP72 in complex with ligand 11 at 21.45 MGy X-ray dose | | Descriptor: | 1,2-ETHANEDIOL, 3-PYRIDINIUM-1-YLPROPANE-1-SULFONATE, 8-chloroadenosine, ... | | Authors: | Cabry, M, Rodrigues, M.J, Le Bihan, Y.V, van Montfort, R.L.M. | | Deposit date: | 2024-01-12 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Specific radiation damage to halogenated inhibitors and ligands in protein-ligand crystal structures.
J.Appl.Crystallogr., 57, 2024
|
|
7GYL
 | | Crystal Structure of HSP72 in complex with ligand 11 at 4.29 MGy X-ray dose | | Descriptor: | 1,2-ETHANEDIOL, 3-PYRIDINIUM-1-YLPROPANE-1-SULFONATE, 8-chloroadenosine, ... | | Authors: | Cabry, M, Rodrigues, M.J, Le Bihan, Y.V, van Montfort, R.L.M. | | Deposit date: | 2024-01-12 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Specific radiation damage to halogenated inhibitors and ligands in protein-ligand crystal structures.
J.Appl.Crystallogr., 57, 2024
|
|
4EGW
 
 | | The structure of the soluble domain of CorA from Methanocaldococcus jannaschii | | Descriptor: | 1,4-BUTANEDIOL, HEXANE-1,6-DIOL, MAGNESIUM ION, ... | | Authors: | Guskov, A, Nordin, N, Reynaud, A, Engman, H, Lundback, A.-K, Jong, A.J.O, Cornvik, T, Phua, T, Eshaghi, S. | | Deposit date: | 2012-04-02 | | Release date: | 2012-10-31 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into the mechanisms of Mg2+ uptake, transport, and gating by CorA
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
1QTI
 
 | | Acetylcholinesterase (E.C.3.1.1.7) | | Descriptor: | (-)-GALANTHAMINE, ACETYLCHOLINESTERASE | | Authors: | Bartolucci, C, Perola, E, Pilger, C, Fels, G, Lamba, D. | | Deposit date: | 1999-06-28 | | Release date: | 1999-12-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Three-dimensional structure of a complex of galanthamine (Nivalin) with acetylcholinesterase from Torpedo californica: implications for the design of new anti-Alzheimer drugs
Proteins, 42, 2001
|
|
5PB4
 
 | | human factor VIIa in complex with 1-[[3-[5-hydroxy-3-methyl-4-(1H-pyrrolo[3,2-c]pyridin-2-yl)pyrazol-1-yl]phenyl]methyl]-3-phenylurea at 2.43A | | Descriptor: | CALCIUM ION, CHLORIDE ION, Coagulation factor VII heavy chain, ... | | Authors: | Stihle, M, Mayweg, A, Roever, S, Rudolph, M.G. | | Deposit date: | 2016-11-16 | | Release date: | 2017-06-21 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Crystal Structure of a Factor VIIa complex
To be published
|
|
3LEV
 
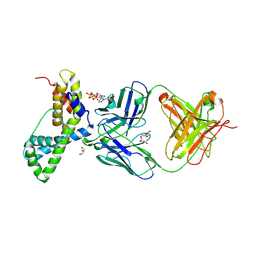 | | HIV-1 antibody 2F5 in complex with epitope scaffold ES2 | | Descriptor: | 2F5 ANTIBODY HEAVY CHAIN, 2F5 ANTIBODY LIGHT CHAIN, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Ofek, G, Guenaga, F.J, Schief, W.R, Skinner, J, Baker, D, Wyatt, R, Kwong, P.D. | | Deposit date: | 2010-01-15 | | Release date: | 2010-09-29 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Elicitation of structure-specific antibodies by epitope scaffolds.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
4I9J
 
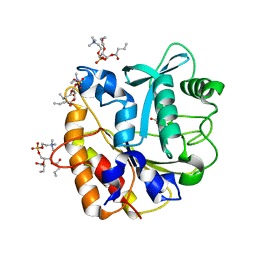 | | Structure of the N254Y/H258Y mutant of the phosphatidylinositol-specific phospholipase C from S. aureus bound to diC4PC | | Descriptor: | (4S,7R)-7-(heptanoyloxy)-4-hydroxy-N,N,N-trimethyl-10-oxo-3,5,9-trioxa-4-phosphahexadecan-1-aminium 4-oxide, 1-phosphatidylinositol phosphodiesterase, ACETATE ION | | Authors: | Goldstein, R.I, Cheng, J, Stec, B, Gershenson, A, Roberts, M.F. | | Deposit date: | 2012-12-05 | | Release date: | 2013-04-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | The cation-pi box is a specific phosphatidylcholine membrane targeting motif.
J.Biol.Chem., 288, 2013
|
|
7D5K
 
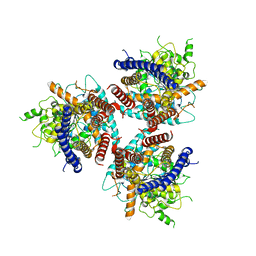 | | CryoEM structure of cotton cellulose synthase isoform 7 | | Descriptor: | Cellulose synthase, beta-D-glucopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Guan, Z.Y, Xue, Y, Yin, P, Zhang, X.L. | | Deposit date: | 2020-09-26 | | Release date: | 2021-07-28 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structural insights into homotrimeric assembly of cellulose synthase CesA7 from Gossypium hirsutum.
Plant Biotechnol J, 19, 2021
|
|
4IAA
 
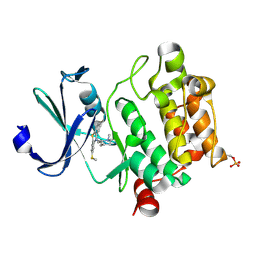 | | Crystal structure of Ser/Thr kinase Pim1 in complex with thioridazine | | Descriptor: | 10-{2-[(2R)-1-methylpiperidin-2-yl]ethyl}-2-(methylsulfanyl)-10H-phenothiazine, Serine/threonine-protein kinase pim-1 | | Authors: | Zhang, W, Wan, X, Li, W, Xie, Y, Huang, N. | | Deposit date: | 2012-12-06 | | Release date: | 2013-12-18 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Crystal structure of Ser/Thr kinase Pim1 in complex with thioridazine
To be Published
|
|
3WF4
 
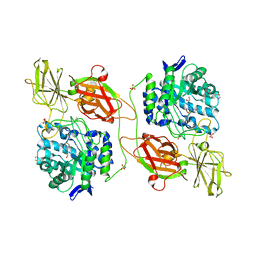 | | Crystal structure of human beta-galactosidase mutant I51T in complex with 6S-NBI-DGJ | | Descriptor: | (3Z,6S,7R,8S,8aS)-3-(butylimino)hexahydro[1,3]thiazolo[3,4-a]pyridine-6,7,8-triol, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Suzuki, H, Ohto, U, Shimizu, T. | | Deposit date: | 2013-07-16 | | Release date: | 2014-04-16 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis of pharmacological chaperoning for human beta-galactosidase
to be published
|
|
6IX4
 
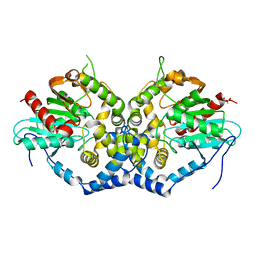 | | Structure of an epoxide hydrolase from Aspergillus usamii E001 (AuEH2) at 1.51 Angstroms resolution | | Descriptor: | ACETATE ION, CHLORIDE ION, GLYCEROL, ... | | Authors: | Hu, D, Hu, B.C, Hou, X.D, Wu, L, Rao, Y.J, Wu, M.C. | | Deposit date: | 2018-12-09 | | Release date: | 2019-12-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.511 Å) | | Cite: | Nearly perfect kinetic resolution of racemic o-nitrostyrene oxide by AuEH2, a microsomal epoxide hydrolase from Aspergillus usamii, with high enantio- and regio-selectivity.
Int.J.Biol.Macromol., 169, 2021
|
|
7GYM
 
 | | Crystal Structure of HSP72 in complex with ligand 11 at 5.72 MGy X-ray dose | | Descriptor: | 1,2-ETHANEDIOL, 3-PYRIDINIUM-1-YLPROPANE-1-SULFONATE, 8-chloroadenosine, ... | | Authors: | Cabry, M, Rodrigues, M.J, Le Bihan, Y.V, van Montfort, R.L.M. | | Deposit date: | 2024-01-12 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Specific radiation damage to halogenated inhibitors and ligands in protein-ligand crystal structures.
J.Appl.Crystallogr., 57, 2024
|
|
160L
 
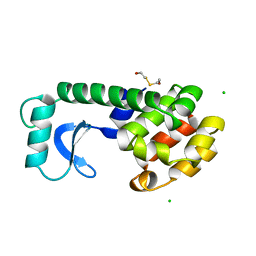 | | CONTROL OF ENZYME ACTIVITY BY AN ENGINEERED DISULFIDE BOND | | Descriptor: | BETA-MERCAPTOETHANOL, CHLORIDE ION, T4 LYSOZYME | | Authors: | Blaber, M, Matthews, B.W. | | Deposit date: | 1994-06-20 | | Release date: | 1994-08-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Alanine scanning mutagenesis of the alpha-helix 115-123 of phage T4 lysozyme: effects on structure, stability and the binding of solvent.
J.Mol.Biol., 246, 1995
|
|
173L
 
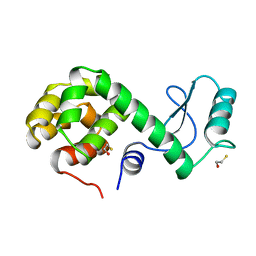 | | PROTEIN FLEXIBILITY AND ADAPTABILITY SEEN IN 25 CRYSTAL FORMS OF T4 LYSOZYME | | Descriptor: | BETA-MERCAPTOETHANOL, T4 LYSOZYME | | Authors: | Xiong, X.-P, Zhang, X.-J, Sun, D, Matthews, B.W. | | Deposit date: | 1995-03-24 | | Release date: | 1995-07-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Protein flexibility and adaptability seen in 25 crystal forms of T4 lysozyme.
J.Mol.Biol., 250, 1995
|
|
2Z6P
 
 | | Crystal Structure of the Ufc1, Ufm1 conjugating enzyme 1 | | Descriptor: | Ufm1-conjugating enzyme 1 | | Authors: | Mizushima , T, Tatsumi, K, Ozaki, Y, Kawakami, T, Suzuki, A, Ogasahara, K, Komatsu, M, Kominami, E, Tanaka, K, Yamane, T. | | Deposit date: | 2007-08-06 | | Release date: | 2007-09-25 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Ufc1, the Ufm1-conjugating enzyme
Biochem.Biophys.Res.Commun., 362, 2007
|
|
177L
 
 | |
2DG9
 
 | |
3H6C
 
 | |
