2MAI
 
 | | NMR structure of lassomycin | | Descriptor: | Lassomycin | | Authors: | Gavrish, E, Sit, C.S, Kandror, O, Spoering, A, Peoples, A, Ling, L, Fetterman, A, Hughes, D, Cao, S, Bissell, A, Torrey, H, Akopian, T, Mueller, A, Epstein, S, Goldberg, A, Clardy, J, Lewis, K. | | Deposit date: | 2013-07-09 | | Release date: | 2014-05-07 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Lassomycin, a Ribosomally Synthesized Cyclic Peptide, Kills Mycobacterium tuberculosis by Targeting the ATP-Dependent Protease ClpC1P1P2.
Chem.Biol., 21, 2014
|
|
2MAJ
 
 | |
2MAK
 
 | |
2MAL
 
 | |
2MAM
 
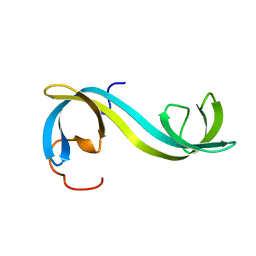 | |
2MAN
 
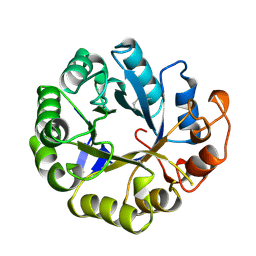 | | MANNOTRIOSE COMPLEX OF THERMOMONOSPORA FUSCA BETA-MANNANASE | | Descriptor: | PROTEIN (BETA-MANNANASE), beta-D-mannopyranose-(1-4)-alpha-D-mannopyranose | | Authors: | Hilge, M, Gloor, S.M, Piontek, K. | | Deposit date: | 1998-08-12 | | Release date: | 1999-08-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | High-resolution native and complex structures of thermostable beta-mannanase from Thermomonospora fusca - substrate specificity in glycosyl hydrolase family 5.
Structure, 6, 1998
|
|
2MAO
 
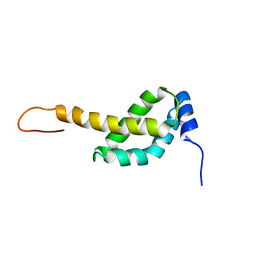 | |
2MAP
 
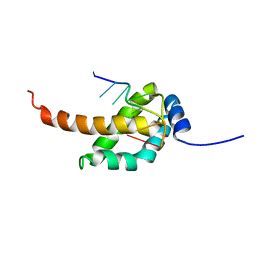 | |
2MAR
 
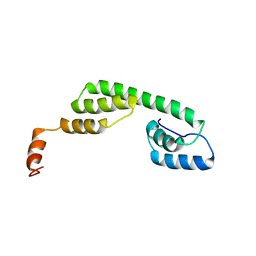 | | Solution structure of Ani s 5 Anisakis simplex allergen | | Descriptor: | SXP/RAL-2 family protein | | Authors: | Garcia-Mayoral, M.F, Trevino, M.A, Perez-Pinar, T, Caballero, M.L, Knaute, T, Umpierrez, A, Bruix, M, Rodriguez-Perez, R. | | Deposit date: | 2013-07-17 | | Release date: | 2014-07-02 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Relationships between IgE/IgG4 epitopes, structure and function in Anisakis simplex Ani s 5, a member of the SXP/RAL-2 protein family
Plos Negl Trop Dis, 8, 2014
|
|
2MAS
 
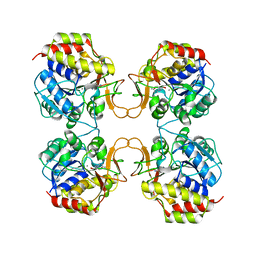 | | PURINE NUCLEOSIDE HYDROLASE WITH A TRANSITION STATE INHIBITOR | | Descriptor: | 2-(4-AMINO-PHENYL)-5-HYDROXYMETHYL-PYRROLIDINE-3,4-DIOL, CALCIUM ION, INOSINE-URIDINE NUCLEOSIDE N-RIBOHYDROLASE | | Authors: | Degano, M, Schramm, V.L, Sacchettini, J.C. | | Deposit date: | 1996-10-17 | | Release date: | 1997-08-12 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Trypanosomal nucleoside hydrolase. A novel mechanism from the structure with a transition-state inhibitor.
Biochemistry, 37, 1998
|
|
2MAT
 
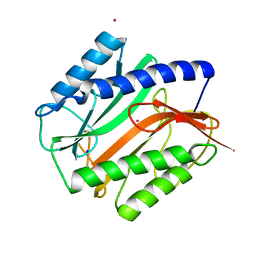 | | E.COLI METHIONINE AMINOPEPTIDASE AT 1.9 ANGSTROM RESOLUTION | | Descriptor: | COBALT (II) ION, PROTEIN (METHIONINE AMINOPEPTIDASE), SODIUM ION | | Authors: | Lowther, W.T, Orville, A.M, Madden, D.T, Lim, S, Rich, D.H, Matthews, B.W. | | Deposit date: | 1999-03-29 | | Release date: | 1999-06-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Escherichia coli methionine aminopeptidase: implications of crystallographic analyses of the native, mutant, and inhibited enzymes for the mechanism of catalysis.
Biochemistry, 38, 1999
|
|
2MAU
 
 | |
2MAV
 
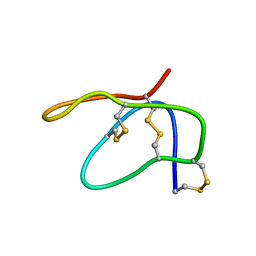
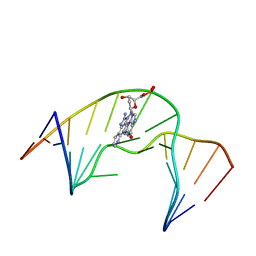 | | NMR Structure of N2-IQ-dG at the G3 position in the NarI recognition sequence | | Descriptor: | DNA_(5'-D(*CP*TP*CP*GP*GP*CP*GP*CP*CP*AP*TP*C)-3'), DNA_(5'-D(*GP*AP*TP*GP*GP*CP*GP*CP*CP*GP*AP*G)-3') | | Authors: | Stavros, K.M, Hawkins, E.K, Rizzo, C.J, Stone, M.P. | | Deposit date: | 2013-07-18 | | Release date: | 2014-05-14 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Base-displaced intercalation of the 2-amino-3-methylimidazo[4,5-f]quinolone N2-dG adduct in the NarI DNA recognition sequence.
Nucleic Acids Res., 42, 2014
|
|
2MAW
 
 | |
2MAX
 
 | |
2MAY
 
 | |
2MAZ
 
 | |
2MB0
 
 | |
2MB1
 | |
2MB2
 | | parallel-stranded G-quadruplex in DNA poly-G stretches | | Descriptor: | DNA_(5'-D(*TP*TP*GP*GP*GP*GP*GP*GP*GP*GP*GP*GP*GP*GP*GP*GP*GP*T)-3') | | Authors: | Sengar, A, Heddi, B, Phan, A.T. | | Deposit date: | 2013-07-24 | | Release date: | 2014-12-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Formation of g-quadruplexes in poly-g sequences: structure of a propeller-type parallel-stranded g-quadruplex formed by a g15 stretch.
Biochemistry, 53, 2014
|
|
2MB3
 
 | | Solution structure of an intramolecular (3+1) human telomeric G-quadruplex bound to a telomestatin derivative | | Descriptor: | (12S,27S)-12,27-bis(4-aminobutyl)-4,30-dimethyl-3,7,14,18,22,29-hexaoxa-11,26,31,32,33,34,35,36-octaazaheptacyclo[26.2. 1.1~2,5~.1~6,9~.1~13,16~.1~17,20~.1~21,24~]hexatriaconta-1(30),2(36),4,6(35),8,13(34),15,17(33),19,21(32),23,28(31)-dode caene-10,25-dione, DNA_(5'-D(*TP*TP*GP*GP*GP*TP*TP*AP*GP*GP*GP*TP*TP*AP*GP*GP*GP*TP*TP*AP*GP*GP*GP*A)-3') | | Authors: | Chung, W.J, Heddi, B, Tera, M, Iida, K, Nagasawa, K, Phan, A.T. | | Deposit date: | 2013-07-24 | | Release date: | 2013-08-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of an intramolecular (3 + 1) human telomeric g-quadruplex bound to a telomestatin derivative.
J.Am.Chem.Soc., 135, 2013
|
|
2MB4
 
 | | Solution structure of a stacked dimeric G-quadruplex formed by a segment of the human CEB1 minisatellite | | Descriptor: | DNA_(5'-D(*AP*GP*GP*GP*GP*GP*GP*AP*GP*GP*GP*AP*GP*GP*GP*TP*GP*G)-3') | | Authors: | Adrian, M, Ang, D.J, Lech, C, Heddi, B, Nicolas, A, Phan, A.T. | | Deposit date: | 2013-07-25 | | Release date: | 2014-05-28 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure and Conformational Dynamics of a Stacked Dimeric G-Quadruplex Formed by the Human CEB1 Minisatellite.
J.Am.Chem.Soc., 136, 2014
|
|
2MB5
 
 | |
2MB7
 | |
2MB9
 
 | | Human Bcl10 CARD | | Descriptor: | B-cell lymphoma/leukemia 10 | | Authors: | Zheng, C, Bracken, C, Wu, H. | | Deposit date: | 2013-07-26 | | Release date: | 2013-10-16 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural Architecture of the CARMA1/Bcl10/MALT1 Signalosome: Nucleation-Induced Filamentous Assembly.
Mol.Cell, 51, 2013
|
|
