3ZIX
 
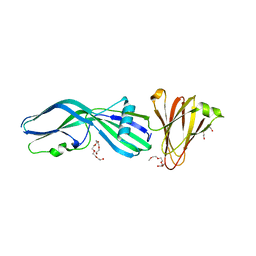 | | Clostridium perfringens Enterotoxin with the N-terminal 37 residues deleted | | Descriptor: | HEAT-LABILE ENTEROTOXIN B CHAIN, HEXAETHYLENE GLYCOL | | Authors: | Yelland, T, Naylor, C.E, Savva, C.G, Basak, A.K. | | Deposit date: | 2013-01-14 | | Release date: | 2014-01-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of a C. Perfringens Enterotoxin Mutant in Complex with a Modified Claudin-2 Extracellular Loop 2
J.Mol.Biol., 426, 2014
|
|
3ZIW
 
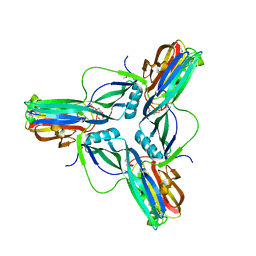 | | Clostridium perfringens enterotoxin, D48A mutation and N-terminal 37 residues deleted | | Descriptor: | HEAT-LABILE ENTEROTOXIN B CHAIN, HEXAETHYLENE GLYCOL | | Authors: | Yelland, T, Naylor, C.E, Savva, C.G, Basak, A.K. | | Deposit date: | 2013-01-14 | | Release date: | 2014-01-29 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of a C. Perfringens Enterotoxin Mutant in Complex with a Modified Claudin-2 Extracellular Loop 2
J.Mol.Biol., 426, 2014
|
|
7AJK
 
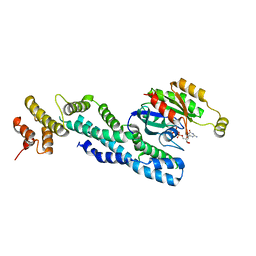 | | Crystal structure of CRYI-B Rac1 complex | | Descriptor: | CYFIP-related Rac1 interactor B, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Yelland, T, Anh, L, Insall, R, Machesky, L, Ismail, S. | | Deposit date: | 2020-09-29 | | Release date: | 2020-11-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural Basis of CYRI-B Direct Competition with Scar/WAVE Complex for Rac1.
Structure, 29, 2021
|
|
7AJL
 
 | | Cyrstal structure of CYRI-B/Fam49B | | Descriptor: | CYFIP-related Rac1 interactor B | | Authors: | Yelland, T, Anh, H, Insall, R, Machesky, L, Ismail, S. | | Deposit date: | 2020-09-29 | | Release date: | 2020-11-18 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Structural Basis of CYRI-B Direct Competition with Scar/WAVE Complex for Rac1.
Structure, 29, 2021
|
|
7OK7
 
 | | Crystal structure of the UNC119B ARL3 complex | | Descriptor: | 1,2-ETHANEDIOL, ADP-ribosylation factor-like protein 3, GLYCEROL, ... | | Authors: | Yelland, T, Ismail, S. | | Deposit date: | 2021-05-17 | | Release date: | 2021-06-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | The Structural and Biochemical Characterization of UNC119B Cargo Binding and Release Mechanisms.
Biochemistry, 60, 2021
|
|
6FMF
 | | Neuropilin-1 b1 domain in complex with EG01377; 2.8 Angstrom structure | | Descriptor: | (2~{S})-2-[[3-[[5-[4-(aminomethyl)phenyl]-1-benzofuran-7-yl]sulfonylamino]thiophen-2-yl]carbonylamino]-5-carbamimidamido-pentanoic acid, Neuropilin-1, trifluoroacetic acid | | Authors: | Yelland, T, Djordjevic, S, Selwood, D, Zachary, I, Frankel, P. | | Deposit date: | 2018-01-31 | | Release date: | 2018-10-17 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.811 Å) | | Cite: | Small Molecule Neuropilin-1 Antagonists Combine Antiangiogenic and Antitumor Activity with Immune Modulation through Reduction of Transforming Growth Factor Beta (TGF beta ) Production in Regulatory T-Cells.
J. Med. Chem., 61, 2018
|
|
6FMC
 | | Neuropilin1-b1 domain in complex with EG01377, 0.9 Angstrom structure | | Descriptor: | (2~{S})-2-[[3-[[5-[4-(aminomethyl)phenyl]-1-benzofuran-7-yl]sulfonylamino]thiophen-2-yl]carbonylamino]-5-carbamimidamido-pentanoic acid, Neuropilin-1 | | Authors: | Yelland, T, Djordjevic, S, Fotinou, K, Selwood, D, Zachary, I, Frankel, P. | | Deposit date: | 2018-01-30 | | Release date: | 2018-10-17 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (0.9 Å) | | Cite: | Small Molecule Neuropilin-1 Antagonists Combine Antiangiogenic and Antitumor Activity with Immune Modulation through Reduction of Transforming Growth Factor Beta (TGF beta ) Production in Regulatory T-Cells.
J. Med. Chem., 61, 2018
|
|
6H6A
 | | Crystal structure of UNC119 in complex with LCK peptide | | Descriptor: | GLY-CYS-GLY-CYS-SER-SER, GLYCEROL, MYRISTIC ACID, ... | | Authors: | Yelland, T, ElMaghloob, Y, McIlwraith, M, Stephen, L, Ismail, S. | | Deposit date: | 2018-07-26 | | Release date: | 2018-09-26 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Ciliary Machinery Is Repurposed for T Cell Immune Synapse Trafficking of LCK.
Dev.Cell, 47, 2018
|
|
7OK6
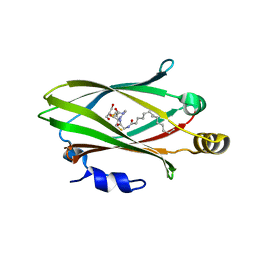 
 | |
7Q9S
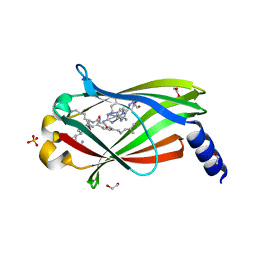 
 | | Crystal structure of PDE6D KRas peptide complex with Compound-1 | | Descriptor: | 1,2-ETHANEDIOL, 2-methyl-3-(1~{H}-pyrazol-5-yl)imidazo[1,2-a]pyridine, DIMETHYL SULFOXIDE, ... | | Authors: | Yelland, T, Ismail, S. | | Deposit date: | 2021-11-14 | | Release date: | 2022-01-19 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Stabilization of the RAS:PDE6D Complex Is a Novel Strategy to Inhibit RAS Signaling.
J.Med.Chem., 65, 2022
|
|
7Q9U
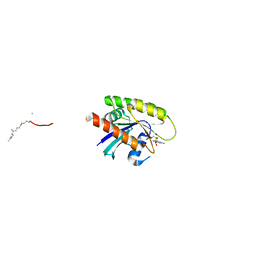 
 | |
5L73
 
 | | MAM domain of human neuropilin-1 | | Descriptor: | BICINE, CALCIUM ION, Neuropilin-1, ... | | Authors: | Yelland, T, Djordjevic, S. | | Deposit date: | 2016-06-01 | | Release date: | 2016-10-26 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Crystal Structure of the Neuropilin-1 MAM Domain: Completing the Neuropilin-1 Ectodomain Picture.
Structure, 24, 2016
|
|
7Q9R
 | | Cocrystal structure of PDE6D bound to NRAS peptide | | Descriptor: | 1,2-ETHANEDIOL, FARNESYL, Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta, ... | | Authors: | Yelland, T, Ismail, S. | | Deposit date: | 2021-11-14 | | Release date: | 2022-01-19 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Stabilization of the RAS:PDE6D Complex Is a Novel Strategy to Inhibit RAS Signaling.
J.Med.Chem., 65, 2022
|
|
7Q9Q
 
 | |
7QF9
 
 | | Crystal structure of PDE6D bound to HRas peptide | | Descriptor: | 1,2-ETHANEDIOL, FARNESYL, HRas peptide, ... | | Authors: | Yelland, T, Ismail, I. | | Deposit date: | 2021-12-05 | | Release date: | 2022-02-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Stabilization of the RAS:PDE6D Complex Is a Novel Strategy to Inhibit RAS Signaling.
J.Med.Chem., 65, 2022
|
|
7QJK
 
 | | Crystal structure of PDE6D in complex with Compound-2 | | Descriptor: | 1,2-ETHANEDIOL, N-(phenylmethyl)pyridin-2-amine, Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta, ... | | Authors: | Yelland, T, Ismail, S. | | Deposit date: | 2021-12-16 | | Release date: | 2022-01-26 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Stabilization of the RAS:PDE6D Complex Is a Novel Strategy to Inhibit RAS Signaling.
J.Med.Chem., 65, 2022
|
|
5IYY
 
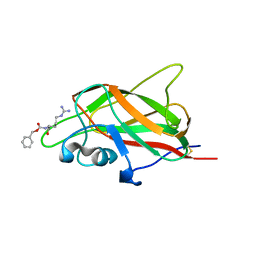 | | X-ray structure of neuropilin-1 b1 domain complexed with Arg-4 ligand. | | Descriptor: | Neuropilin-1, N~2~-[(benzyloxy)carbonyl]-L-arginine | | Authors: | Fotinou, C, Rana, R, Djordjevic, S, Yelland, T. | | Deposit date: | 2016-03-24 | | Release date: | 2017-04-05 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Architecture and hydration of the arginine-binding site of neuropilin-1.
FEBS J., 285, 2018
|
|
4P5H
 
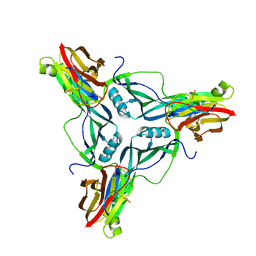 | |
5IJR
 
 | | X-ray structure of neuropilin-1 b1 domain complexed with Arg-1 ligand. | | Descriptor: | DIMETHYL SULFOXIDE, L-HOMOARGININE, Neuropilin-1 | | Authors: | Fotinou, C, Rana, R, Djordjevic, S, Yelland, T. | | Deposit date: | 2016-03-02 | | Release date: | 2017-03-29 | | Last modified: | 2018-07-11 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Architecture and hydration of the arginine-binding site of neuropilin-1.
FEBS J., 285, 2018
|
|
4RN5
 
 | | B1 domain of human Neuropilin-1 with acetate ion in a ligand-binding site | | Descriptor: | ACETATE ION, GLYCEROL, Neuropilin-1, ... | | Authors: | Allerston, C.K, Yelland, T.S, Jarvis, A, Jenkins, K, Winfield, N, Cheng, L, Jia, H, Zachary, I, Selwood, D.L, Djordjevic, S. | | Deposit date: | 2014-10-23 | | Release date: | 2015-10-28 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Conserved water molecules in a ligand-binding site of neuropilin-1
To be Published
|
|
7ZC8
 
 | |
6TDB
 | | Neuropilin2-b1 domain in a complex with the C-terminal VEGFB167 peptide | | Descriptor: | 1,2-ETHANEDIOL, ACETYL GROUP, C-terminal VEGFB167 peptide, ... | | Authors: | Eldrid, C, Yu, L, Yelland, T, Fotinou, C, Djordjevic, S. | | Deposit date: | 2019-11-08 | | Release date: | 2020-11-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | NRP2-b1 domain in a complex with the C-terminal VEGFB167 peptide
To Be Published
|
|
6TJT
 | | Neuropilin2-b1 domain in a complex with the C-terminal VEGFC peptide | | Descriptor: | 1,2-ETHANEDIOL, Neuropilin-2, SULFATE ION, ... | | Authors: | Eldrid, C, Yu, L, Yelland, T, Fotinou, C, Djordjevic, S. | | Deposit date: | 2019-11-26 | | Release date: | 2020-12-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.31 Å) | | Cite: | NRP2-b1 domain in a complex with the C-terminal VEGFC peptide
To Be Published
|
|
6TKK
 
 | | Neuropilin 1-b1 domain in a complex with the C-terminal VEGFB186 peptide | | Descriptor: | 1,2-ETHANEDIOL, ACE-ARG-PRO-GLN-PRO-ARG, Neuropilin-1 | | Authors: | Eldrid, C, Yu, L, Yelland, T, Fotinou, C, Djordjevic, S. | | Deposit date: | 2019-11-28 | | Release date: | 2020-12-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.06 Å) | | Cite: | Neuropilin 1-b1 domain in a complex with the C-terminal VEGFB186 peptide
To Be Published
|
|
5JHK
 | | X-ray structure of neuropilin-1 b1 domain complexed with Arg-6 ligand. | | Descriptor: | N-(benzenecarbonyl)glycyl-L-arginine, Neuropilin-1 | | Authors: | Fotinou, C, Rana, R, Djordjevic, S, Yelland, T. | | Deposit date: | 2016-04-21 | | Release date: | 2017-05-24 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Architecture and hydration of the arginine-binding site of neuropilin-1.
FEBS J., 285, 2018
|
|
