6UNW
 
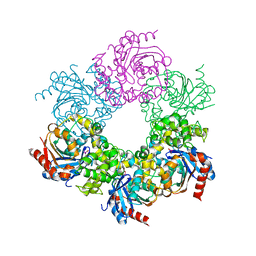 | | Epoxide hydrolase from an endophytic Streptomyces | | Descriptor: | CACODYLATE ION, CHLORIDE ION, Soluble epoxide hydrolase | | Authors: | Wilson, C, dos Santos, J.C, Dias, M.V.B. | | Deposit date: | 2019-10-13 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | An epoxide hydrolase from endophytic Streptomyces shows unique structural features and wide biocatalytic activity.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
5URO
 
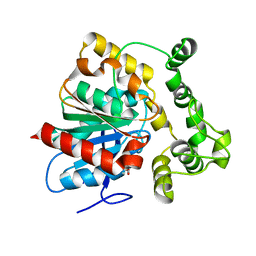 | | Structure of a soluble epoxide hydrolase identified in Trichoderma reesei | | Descriptor: | 2,2'-oxydi(ethyn-1-ol), Predicted protein | | Authors: | Wilson, C, de Oliveira, G, Chambergo, F, Dias, M.V.B. | | Deposit date: | 2017-02-11 | | Release date: | 2017-05-31 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.701 Å) | | Cite: | Structure of a soluble epoxide hydrolase identified in Trichoderma reesei.
Biochim. Biophys. Acta, 1865, 2017
|
|
1XNF
 
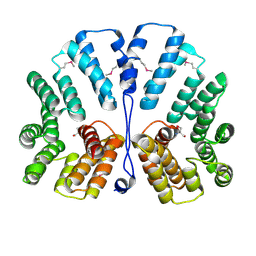 | | Crystal structure of E.coli TPR-protein NlpI | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Lipoprotein nlpI | | Authors: | Wilson, C.G, Kajander, T, Regan, L. | | Deposit date: | 2004-10-04 | | Release date: | 2004-11-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | The crystal structure of NlpI. A prokaryotic tetratricopeptide repeat protein with a globular fold.
FEBS J., 272, 2005
|
|
1LE2
 
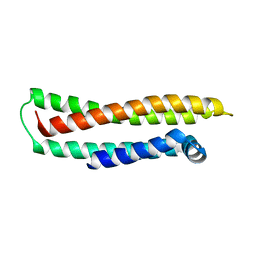 | |
1LE4
 
 | |
1LPE
 
 | |
1A6S
 
 | | M-DOMAIN FROM GAG POLYPROTEIN OF ROUS SARCOMA VIRUS, NMR, 20 STRUCTURES | | Descriptor: | GAG POLYPROTEIN | | Authors: | Mcdonnell, J.M, Fushman, D, Cahill, S.M, Zhou, W, Wolven, A, Wilson, C.B, Nelle, T.D, Resh, M.D, Wills, J, Cowburn, D. | | Deposit date: | 1998-03-02 | | Release date: | 1998-10-14 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the bioactive retroviral M domain from Rous sarcoma virus
J.Mol.Biol., 279, 1998
|
|
4UEU
 
 | | Tyrosine kinase AS - a common ancestor of Src and Abl | | Descriptor: | PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER, TYROSINE KINASE AS - A COMMON ANCESTOR OF SRC AND ABL | | Authors: | Kutter, S, Wilson, C, Agafonov, R.V, Hoemberger, M.S, Zorba, A, Halpin, J.C, Theobald, D.L, Kern, D. | | Deposit date: | 2014-12-18 | | Release date: | 2015-02-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Kinase Dynamics. Using Ancient Protein Kinases to Unravel a Modern Cancer Drug'S Mechanism.
Science, 347, 2015
|
|
1DDY
 
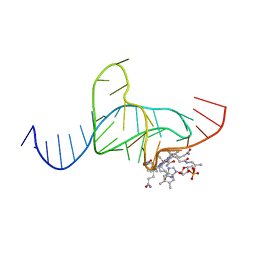 | |
7KBT
 
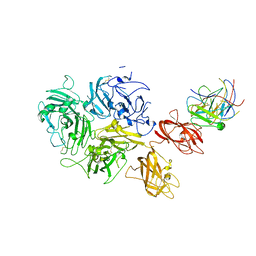 | | Factor VIII in complex with the anti-C2 domain antibody, G99 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Ronayne, E.K, Gish, J, Wilson, C, Peters, S, Spencer, H.T, Spiegel, P.C, Childers, K.C. | | Deposit date: | 2020-10-02 | | Release date: | 2020-11-18 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (4.15 Å) | | Cite: | Structure of Blood Coagulation Factor VIII in Complex With an Anti-C2 Domain Non-Classical, Pathogenic Antibody Inhibitor
Front Immunol, 12, 2021
|
|
1ET4
 
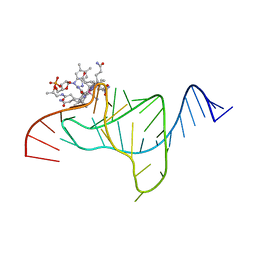 | |
1F1T
 
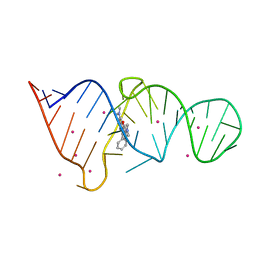 | | CRYSTAL STRUCTURE OF THE MALACHITE GREEN APTAMER COMPLEXED WITH TETRAMETHYL-ROSAMINE | | Descriptor: | MALACHITE GREEN APTAMER RNA, N,N'-TETRAMETHYL-ROSAMINE, STRONTIUM ION | | Authors: | Baugh, C, Grate, D, Wilson, C. | | Deposit date: | 2000-05-19 | | Release date: | 2000-09-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | 2.8 A crystal structure of the malachite green aptamer.
J.Mol.Biol., 301, 2000
|
|
4CSV
 
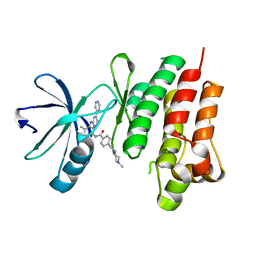 | | Tyrosine kinase AS - a common ancestor of Src and Abl bound to Gleevec | | Descriptor: | 4-(4-METHYL-PIPERAZIN-1-YLMETHYL)-N-[4-METHYL-3-(4-PYRIDIN-3-YL-PYRIMIDIN-2-YLAMINO)-PHENYL]-BENZAMIDE, SRC-ABL TYROSINE KINASE ANCESTOR | | Authors: | Kutter, S, Wilson, C, Cruz, L, Agafonov, R.V, Hoemberger, M.S, Zorba, A, Kern, D. | | Deposit date: | 2014-03-10 | | Release date: | 2015-03-04 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Kinase Dynamics. Using Ancient Protein Kinases to Unravel a Modern Cancer Drug'S Mechanism.
Science, 347, 2015
|
|
1F27
 
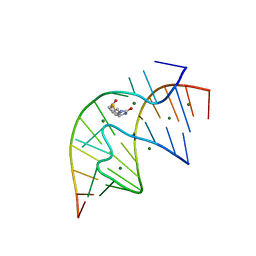 | | CRYSTAL STRUCTURE OF A BIOTIN-BINDING RNA PSEUDOKNOT | | Descriptor: | BIOTIN, MAGNESIUM ION, RNA (5'-R(*AP*AP*AP*AP*AP*GP*UP*CP*CP*UP*C)-3'), ... | | Authors: | Nix, J, Sussman, D, Wilson, C. | | Deposit date: | 2000-05-23 | | Release date: | 2000-06-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The 1.3 A crystal structure of a biotin-binding pseudoknot and the basis for RNA molecular recognition.
J.Mol.Biol., 296, 2000
|
|
2OO7
 
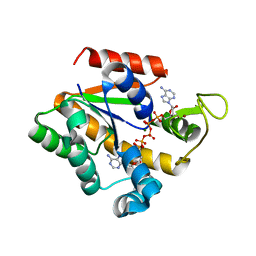 | | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (T179I/Q199R) | | Descriptor: | Adenylate kinase, BIS(ADENOSINE)-5'-PENTAPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Counago, R, Wilson, C.J, Wu, G, Myers, J.C, Wittung-Stafshede, P, Shamoo, Y. | | Deposit date: | 2007-01-25 | | Release date: | 2008-02-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (T179I/Q199R)
To be Published
|
|
2ORI
 
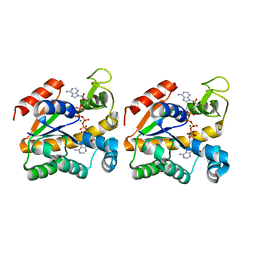 | | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (A193V/Q199R/) | | Descriptor: | Adenylate kinase, BIS(ADENOSINE)-5'-PENTAPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Counago, R, Shamoo, Y, Wilson, C.J, Wu, G, Myers, J. | | Deposit date: | 2007-02-02 | | Release date: | 2008-01-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (A193V/Q199R/)
To be Published
|
|
2OSB
 
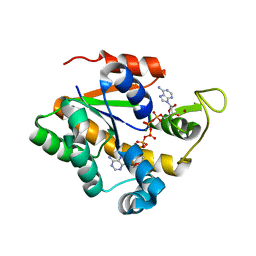 | | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (Q16L/Q199R/) | | Descriptor: | Adenylate kinase, BIS(ADENOSINE)-5'-PENTAPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Myers, J, Counago, R, Wilson, C.J, Wu, G, Shamoo, Y. | | Deposit date: | 2007-02-05 | | Release date: | 2008-01-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (Q16L/Q199R/)
To be Published
|
|
2P3S
 
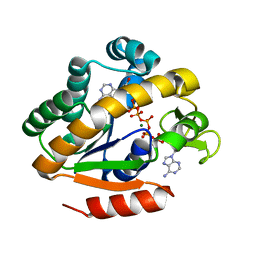 | | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (G214R/Q199R) | | Descriptor: | Adenylate kinase, BIS(ADENOSINE)-5'-PENTAPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Counago, R, Wilson, C.J, Myers, J, Wu, G, Shamoo, Y. | | Deposit date: | 2007-03-09 | | Release date: | 2008-01-22 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (G214R/Q199R)
To be Published
|
|
2QAJ
 
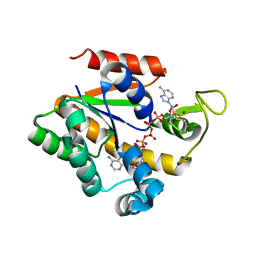 | | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (Q199R/G213E) | | Descriptor: | Adenylate kinase, BIS(ADENOSINE)-5'-PENTAPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Counago, R, Wilson, C.J, Myers, J, Wu, G, Shamoo, Y. | | Deposit date: | 2007-06-15 | | Release date: | 2008-04-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a thermostable mutant of Bacillus subtilis Adenylate Kinase (Q199R/G213E)
To be Published
|
|
8OS5
 
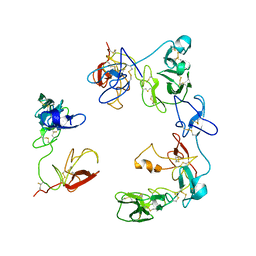 | | Crystal structure of the Factor XII heavy chain reveals an interlocking dimer with a FnII to kringle domain interaction | | Descriptor: | Coagulation factor XII-Mie | | Authors: | Li, C, Saleem, M, Kaira, B.G, Brown, A, Wilson, C, Philippou, H, Emsley, J. | | Deposit date: | 2023-04-18 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Factor XII and kininogen asymmetric assembly with gC1qR/C1QBP/P32 is governed by allostery.
Blood, 136, 2020
|
|
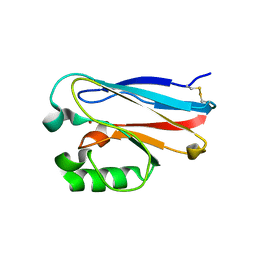
4KO7
 
 | |
4KOB
 
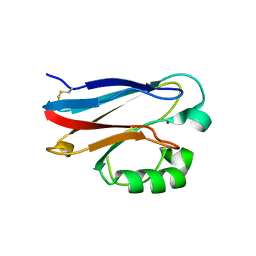 | |
4KO5
 
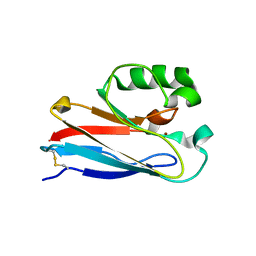 | | Investigating the functional significance of the interlocked pair structural determinants in Pseudomonas aeruginosa azurin (V31I/W48L/V95I/Y108F) | | Descriptor: | Azurin, COPPER (II) ION | | Authors: | Inampudi, K.K, Wang, Y, Meng, W, Tobin, P.H, Wilson, C.J. | | Deposit date: | 2013-05-11 | | Release date: | 2014-05-14 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Investigating the functional significance of the interlocked pair structural determinants in Pseudomonas aeruginosa azurin (V31I/W48L/V95I/Y108F)
To be Published
|
|
1Q8N
 
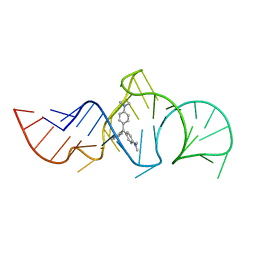 | | Solution Structure of the Malachite Green RNA Binding Aptamer | | Descriptor: | MALACHITE GREEN, RNA Aptamer | | Authors: | Flinders, J, DeFina, S.C, Brackett, D.M, Baugh, C, Wilson, C, Dieckmann, T. | | Deposit date: | 2003-08-21 | | Release date: | 2004-03-23 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Recognition of planar and nonplanar ligands in the malachite green-RNA aptamer complex.
Chembiochem, 5, 2004
|
|
4K4F
 
 | | Co-crystal structure of TNKS1 with compound 18 [4-[(4S)-5,5-dimethyl-2-oxo-4-phenyl-1,3-oxazolidin-3-yl]-N-(quinolin-8-yl)benzamide] | | Descriptor: | 4-[(4S)-5,5-dimethyl-2-oxo-4-phenyl-1,3-oxazolidin-3-yl]-N-(quinolin-8-yl)benzamide, Tankyrase-1, ZINC ION | | Authors: | Huang, X, Bregman, H, Wilson, C, DiMauro, E, Gunaydin, H. | | Deposit date: | 2013-04-12 | | Release date: | 2013-06-05 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Discovery of novel, induced-pocket binding oxazolidinones as potent, selective, and orally bioavailable tankyrase inhibitors.
J.Med.Chem., 56, 2013
|
|
