1WCH
 
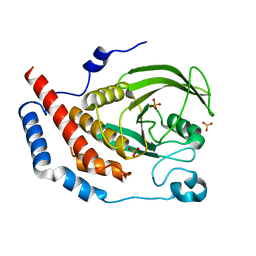 | | Crystal structure of PTPL1 human tyrosine phosphatase mutated in colorectal cancer - evidence for a second phosphotyrosine substrate recognition pocket | | Descriptor: | PHOSPHATE ION, PROTEIN TYROSINE PHOSPHATASE, NON-RECEPTOR TYPE 13 | | Authors: | Villa, F, Deak, M, Bloomberg, G.B, Alessi, D.R, Van Aalten, D.M.F. | | Deposit date: | 2004-11-16 | | Release date: | 2004-12-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure of Ptpl1/Fap-1 Human Tyrosine Phosphatase Mutated in Colorectal Cancer: Eveidence for a Second Phosphotyrosine Substrate Recognition Pocket
J.Biol.Chem., 280, 2005
|
|
2V3S
 
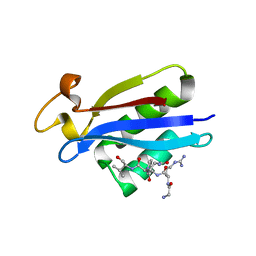 | | Structural insights into the recognition of substrates and activators by the OSR1 kinase | | Descriptor: | ACETATE ION, SERINE/THREONINE-PROTEIN KINASE OSR1, SERINE/THREONINE-PROTEIN KINASE WNK4 | | Authors: | Villa, F, Goebel, J, Rafiqi, F.H, Deak, M, Thastrup, J, Alessi, D.R, van Aalten, D.M.F. | | Deposit date: | 2007-06-21 | | Release date: | 2007-07-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Insights Into the Recognition of Substrates and Activators by the Osr1 Kinase
Embo Rep., 8, 2007
|
|
2VWI
 
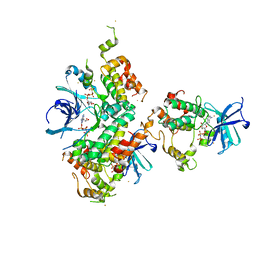 | | Structure of the OSR1 kinase, a hypertension drug target | | Descriptor: | GOLD ION, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Villa, F, Deak, M, Alessi, D.R, vanAalten, D.M.F. | | Deposit date: | 2008-06-25 | | Release date: | 2008-07-08 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structure of the Osr1 Kinase, a Hypertension Drug Target.
Proteins, 73, 2008
|
|
2WB8
 
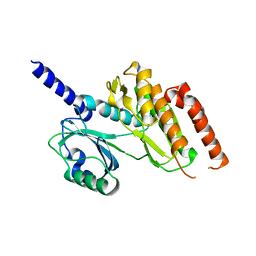 | | Crystal structure of Haspin kinase | | Descriptor: | SERINE/THREONINE-PROTEIN KINASE HASPIN | | Authors: | Villa, F, Tortorci, M, Forneris, F, Mattevi, A, Musacchio, A. | | Deposit date: | 2009-02-23 | | Release date: | 2009-11-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal Structure of the Catalytic Domain of Haspin, an Atypical Kinase Implicated in Chromatin Organization.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
4C2W
 
 | | Crystal structure of Aurora B in complex with AMP-PNP | | Descriptor: | AURORA KINASE B-A, INNER CENTROMERE PROTEIN A, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Sessa, F, Villa, F. | | Deposit date: | 2013-08-20 | | Release date: | 2014-02-26 | | Last modified: | 2014-03-19 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of Aurora B-Incenp in Complex with Barasertib Reveals a Potential Transinhibitory Mechanism
Acta Crystallogr.,Sect.F, 70, 2014
|
|
2CBJ
 
 | | Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase in complex with PUGNAc | | Descriptor: | CHLORIDE ION, HYALURONIDASE, O-(2-ACETAMIDO-2-DEOXY D-GLUCOPYRANOSYLIDENE) AMINO-N-PHENYLCARBAMATE | | Authors: | Rao, F.V, Dorfmueller, H.C, Villa, F, Allwood, M, Eggleston, I.M, van Aalten, D.M.F. | | Deposit date: | 2006-01-05 | | Release date: | 2006-02-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural insights into the mechanism and inhibition of eukaryotic O-GlcNAc hydrolysis.
EMBO J., 25, 2006
|
|
2CBI
 
 | | Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase | | Descriptor: | CHLORIDE ION, GAMMA-BUTYROLACTONE, GLYCEROL, ... | | Authors: | Rao, F.V, Dorfmueller, H.C, Villa, F, Allwood, M, Eggleston, I.M, van Aalten, D.M.F. | | Deposit date: | 2006-01-04 | | Release date: | 2006-02-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural insights into the mechanism and inhibition of eukaryotic O-GlcNAc hydrolysis.
EMBO J., 25, 2006
|
|
4C2V
 
 | | Aurora B kinase in complex with the specific inhibitor Barasertib | | Descriptor: | 2-[5-[[7-[3-[ethyl(2-hydroxyethyl)amino]propoxy]quinazolin-4-yl]amino]-1H-pyrazol-3-yl]-N-(3-fluorophenyl)ethanamide, AURORA KINASE B-A, INNER CENTROMERE PROTEIN A | | Authors: | Sessa, F, Villa, F. | | Deposit date: | 2013-08-20 | | Release date: | 2014-02-26 | | Last modified: | 2014-03-19 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Structure of Aurora B-Incenp in Complex with Barasertib Reveals a Potential Transinhibitory Mechanism
Acta Crystallogr.,Sect.F, 70, 2014
|
|
4B8M
 
 | | Aurora B kinase in complex with VX-680 | | Descriptor: | AURORA KINASE B-A, CYCLOPROPANECARBOXYLIC ACID {4-[4-(4-METHYL-PIPERAZIN-1-YL)-6-(5-METHYL-2H-PYRAZOL-3-YLAMINO)-PYRIMIDIN-2-YLSULFANYL]-PHENYL}-AMIDE, INNER CENTROMERE PROTEIN A | | Authors: | Sessa, F, Villa, F. | | Deposit date: | 2012-08-28 | | Release date: | 2013-07-10 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural and Biochemical Analysis of an Aurora B Kinase Mutant Reveals a Multistep Activation Mechanism
To be Published
|
|
2VRX
 
 | | Structure of Aurora B kinase in complex with ZM447439 | | Descriptor: | INNER CENTROMERE PROTEIN A, N-(4-{[6-methoxy-7-(3-morpholin-4-ylpropoxy)quinazolin-4-yl]amino}phenyl)benzamide, SERINE/THREONINE-PROTEIN KINASE 12-A | | Authors: | Girdler, F, Sessa, F, Patercoli, S, Villa, F, Ridgway, E, Musacchio, A, Taylor, S.S. | | Deposit date: | 2008-04-16 | | Release date: | 2008-07-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Molecular Basis of Drug Resistance in Aurora Kinases.
Chem.Biol., 15, 2008
|
|
2VGO
 
 | | Crystal structure of Aurora B kinase in complex with Reversine inhibitor | | Descriptor: | INNER CENTROMERE PROTEIN A, N~6~-cyclohexyl-N~2~-(4-morpholin-4-ylphenyl)-9H-purine-2,6-diamine, SERINE/THREONINE-PROTEIN KINASE 12-A | | Authors: | D'Alise, A.M, Amabile, G, Iovino, M, Di Giorgio, F.P, Bartiromo, M, Sessa, F, Villa, F, Musacchio, A, Cortese, R. | | Deposit date: | 2007-11-15 | | Release date: | 2008-10-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Reversine, a Novel Aurora Kinases Inhibitor, Inhibits Colony Formation of Human Acute Myeloid Leukemia Cells.
Mol.Cancer Ther., 7, 2008
|
|
4B8L
 
 | | Aurora B kinase P353G mutant | | Descriptor: | 9-{5-O-[(R)-hydroxy{[(R)-hydroxy(phosphonoamino)phosphoryl]oxy}phosphoryl]-alpha-L-arabinofuranosyl}-9H-purin-6-amine, AURORA KINASE B-A, INNER CENTROMERE PROTEIN A | | Authors: | Sessa, F, Villa, F. | | Deposit date: | 2012-08-28 | | Release date: | 2013-07-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and Biochemical Analysis of an Aurora B Kinase Mutant Reveals a Multistep Activation Mechanism
To be Published
|
|
2VGP
 
 | | Crystal structure of Aurora B kinase in complex with a aminothiazole inhibitor | | Descriptor: | 4-[(5-bromo-1,3-thiazol-2-yl)amino]-N-methylbenzamide, INNER CENTROMERE PROTEIN A, SERINE/THREONINE-PROTEIN KINASE 12-A | | Authors: | Andersen, C.B, Wan, Y, Chang, J.W, Lee, C, Liu, Y, Sessa, F, Villa, F, Nallan, L, Musacchio, A, Gray, N.S. | | Deposit date: | 2007-11-15 | | Release date: | 2008-02-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Discovery of Selective Aminothiazole Aurora Kinase Inhibitors
Acs Chem.Biol., 3, 2008
|
|
3ZTX
 
 | | Aurora kinase selective inhibitors identified using a Taxol-induced checkpoint sensitivity screen. | | Descriptor: | 2-((4-(4-HYDROXYPIPERIDIN-1-YL)PHENYL)AMINO)-5,11-DIMETHYL-5H-BENZO[E]PYRIMIDO [5,4-B][1,4]DIAZEPIN-6(11H)-ONE, INNER CENTROMERE PROTEIN A, SERINE/THREONINE-PROTEIN KINASE 12-A | | Authors: | Kwiatkowski, N, Villa, F, Musacchio, A, Gray, N. | | Deposit date: | 2011-07-12 | | Release date: | 2012-02-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Selective Aurora Kinase Inhibitors Identified Using a Taxol- Induced Checkpoint Sensitivity Screen.
Acs Chem.Biol., 7, 2012
|
|
8PFP
 
 | |
6SEN
 
 | | TEAD4 bound to a FAM181A peptide | | Descriptor: | Protein FAM181A, SULFATE ION, Transcriptional enhancer factor TEF-3 | | Authors: | Scheufler, C, Villard, F. | | Deposit date: | 2019-07-30 | | Release date: | 2019-11-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Identification of FAM181A and FAM181B as new interactors with the TEAD transcription factors.
Protein Sci., 29, 2020
|
|
6Y20
 
 | |
8A8Q
 
 | |
6SEO
 
 | | TEAD4 bound to a FAM181B peptide | | Descriptor: | Protein FAM181B, Transcriptional enhancer factor TEF-3 | | Authors: | Scheufler, C, Villard, F. | | Deposit date: | 2019-07-30 | | Release date: | 2019-11-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Identification of FAM181A and FAM181B as new interactors with the TEAD transcription factors.
Protein Sci., 29, 2020
|
|
6Z10
 | | Crystal structure of a humanized (K18E, K269N) rat succinate receptor SUCNR1 (GPR91) in complex with a nanobody and antagonist | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 2-[2-[[3-[4-chloranyl-2-fluoranyl-5-[(3~{R})-piperidin-3-yl]oxy-phenyl]-2-fluoranyl-phenyl]carbonylamino]-5-fluoranyl-phenyl]ethanoic acid, CHLORIDE ION, ... | | Authors: | Haffke, M, Villard, F. | | Deposit date: | 2020-05-11 | | Release date: | 2020-09-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.269 Å) | | Cite: | Discovery and Optimization of Novel SUCNR1 Inhibitors: Design of Zwitterionic Derivatives with a Salt Bridge for the Improvement of Oral Exposure.
J.Med.Chem., 63, 2020
|
|
8P0M
 
 | | Crystal structure of TEAD3 in complex with IAG933 | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, 4-[(2~{S})-5-chloranyl-6-fluoranyl-2-phenyl-2-[(2~{S})-pyrrolidin-2-yl]-3~{H}-1-benzofuran-4-yl]-5-fluoranyl-6-(2-hydroxyethyloxy)-~{N}-methyl-pyridine-3-carboxamide, DIMETHYL SULFOXIDE, ... | | Authors: | Scheufler, C, Villard, F, Chau, S. | | Deposit date: | 2023-05-10 | | Release date: | 2024-04-10 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.961 Å) | | Cite: | Direct and selective pharmacological disruption of the YAP-TEAD interface by IAG933 inhibits Hippo-dependent and RAS-MAPK-altered cancers.
Nat Cancer, 2024
|
|
