5A4C
 
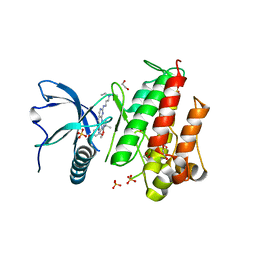 | | FGFR1 ligand complex | | Descriptor: | 1,2-ETHANEDIOL, 1-tert-butyl-3-[2-[3-(diethylamino)propylamino]-6-(3,5-dimethoxyphenyl)pyrido[2,3-d]pyrimidin-7-yl]urea, FIBROBLAST GROWTH FACTOR RECEPTOR 1 (FMS-RELATED TYROSINE KINASE 2, ... | | Authors: | Klein, T, Vajpai, N, Phillips, J.J, Davies, G, Holdgate, G.A, Phillips, C, Tucker, J.A, Norman, R.A, Scott, A.S, Higazi, D.R, Lowe, D, Thompson, G.S, Breeze, A.L. | | Deposit date: | 2015-06-05 | | Release date: | 2015-08-05 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structural and Dynamic Insights Into the Energetics of Activation Loop Rearrangement in Fgfr1 Kinase.
Nat.Commun., 6, 2015
|
|
5A46
 
 | | FGFR1 in complex with dovitinib | | Descriptor: | 1,2-ETHANEDIOL, 4-amino-5-fluoro-3-[5-(4-methylpiperazin-1-yl)-1H-benzimidazol-2-yl]quinolin-2(1H)-one, FIBROBLAST GROWTH FACTOR RECEPTOR 1 (FMS-RELATED TYROSINE KINASE 2, ... | | Authors: | Klein, T, Vajpai, N, Phillips, J.J, Davies, G, Holdgate, G.A, Phillips, C, Tucker, J.A, Norman, R.A, Scott, A.S, Higazi, D.R, Lowe, D, Thompson, G.S, Breeze, A.L. | | Deposit date: | 2015-06-05 | | Release date: | 2015-08-05 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Structural and Dynamic Insights Into the Energetics of Activation Loop Rearrangement in Fgfr1 Kinase.
Nat.Commun., 6, 2015
|
|
3ZD0
 
 | | The Solution Structure of Monomeric Hepatitis C Virus p7 Yields Potent Inhibitors of Virion Release | | Descriptor: | P7 PROTEIN | | Authors: | Foster, T.L, Sthompson, G, Kalverda, A.P, Kankanala, J, Thompson, J, Barker, A.M, Clarke, D, Noerenberg, M, Pearson, A.R, Rowlands, D.J, Homans, S.W, Harris, M, Foster, R, Griffin, S.D.C. | | Deposit date: | 2012-11-23 | | Release date: | 2013-09-04 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Structure-Guided Design Affirms Inhibitors of Hepatitis C Virus P7 as a Viable Class of Antivirals Targeting Virion Release
Hepatology, 59, 2014
|
|
2XKS
 
 | | Prion-like conversion during amyloid formation at atomic resolution | | Descriptor: | BETA-2-MICROGLOBULIN | | Authors: | Eichner, T, Kalverda, A.P, Thompson, G.S, Radford, S.E, Homans, S.W. | | Deposit date: | 2010-07-12 | | Release date: | 2011-02-16 | | Last modified: | 2020-01-15 | | Method: | SOLUTION NMR | | Cite: | Conformational Conversion During Amyloid Formation at Atomic Resolution.
Mol.Cell, 41, 2011
|
|
2XKU
 
 | | Prion-like conversion during amyloid formation at atomic resolution | | Descriptor: | BETA-2-MICROGLOBULIN | | Authors: | Eichner, T, Kalverda, A.P, Thompson, G.S, Homans, S.W, Radford, S.E. | | Deposit date: | 2010-07-12 | | Release date: | 2011-02-09 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Conformational Conversion During Amyloid Formation at Atomic Resolution.
Mol.Cell, 41, 2011
|
|
2MZW
 
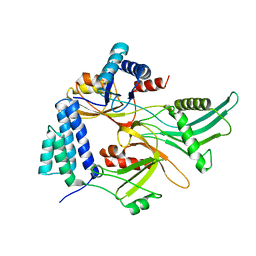 | | Staphylococcus aureus FusB:EF-GC3 complex | | Descriptor: | Elongation factor G, Far1, ZINC ION | | Authors: | Tomlinson, J.H, Thompson, G.S, Kalverda, A.P, Zhuravleva, A, O'Neill, A. | | Deposit date: | 2015-02-25 | | Release date: | 2016-01-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | A target-protection mechanism of antibiotic resistance at atomic resolution: insights into FusB-type fusidic acid resistance.
Sci Rep, 6, 2016
|
|
2K36
 
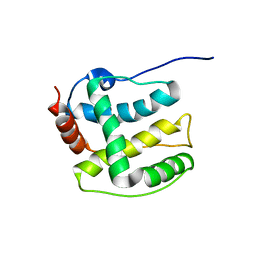 | | Structure ensemble Backbone and side-chain 1H, 13C, and 15N Chemical Shift Assignments, 1H-15N RDCs and 1H-1H nOe restraints for protein K7 from the Vaccinia virus | | Descriptor: | Protein K7 | | Authors: | Kalverda, A.P, Thompson, G.S, Vogel, A, Schr der, M, Bowie, A.G, Khan, A.R, Homans, S.W. | | Deposit date: | 2008-04-22 | | Release date: | 2008-10-28 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Poxvirus K7 protein adopts a Bcl-2 fold: biochemical mapping of its interactions with human DEAD box RNA helicase DDX3.
J.Mol.Biol., 385, 2009
|
|
2KYA
 
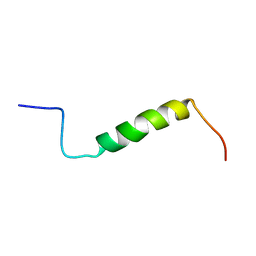 | | Solution structure of the leader sequence of the patellamide precursor peptide, PatE1-34 | | Descriptor: | Patellamide protein | | Authors: | Houssen, W.E, Wright, S.H, Kalverda, A.P, Thompson, G.S, Kelly, S.M, Jaspars, M. | | Deposit date: | 2010-05-21 | | Release date: | 2010-09-01 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Leader Sequence of the Patellamide Precursor Peptide, PatE(1-34).
Chembiochem, 11, 2010
|
|
