5J0U
 
 | |
5J0Q
 
 | |
5J0T
 
 | |
5J2I
 
 | |
5J2D
 
 | |
5J2B
 
 | |
5J2H
 
 | |
7W2W
 
 | | Crystal structure of TxGH116 R786K mutant from Thermoanaerobacterium xylanolyticum with glucose | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, GLYCEROL, ... | | Authors: | Huang, M, Pengthaisong, S, Charoenwattanasatien, R, Jitonnom, J, Ketudat Cairns, J.R. | | Deposit date: | 2021-11-24 | | Release date: | 2022-04-06 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Systematic Functional and Computational Analysis of Glucose-Binding Residues in Glycoside Hydrolase Family GH116.
Catalysts, 12, 2022
|
|
3I7Z
 
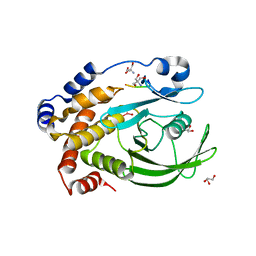 | | Protein Tyrosine Phosphatase 1B - Transition state analog for the first catalytic step | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, EGFR receptor fragment, GLYCEROL, ... | | Authors: | Brandao, T.A.S, Johnson, S.J, Hengge, A.C. | | Deposit date: | 2009-07-09 | | Release date: | 2010-03-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Insights into the reaction of protein-tyrosine phosphatase 1B: crystal structures for transition state analogs of both catalytic steps.
J.Biol.Chem., 285, 2010
|
|
4BFG
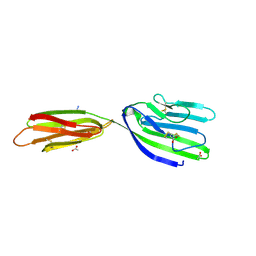 
 | | Structure of the extracellular portion of mouse CD200R | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, AZIDE ION, ... | | Authors: | Hatherley, D, Lea, S.M, Johnson, S, Barclay, A.N. | | Deposit date: | 2013-03-18 | | Release date: | 2013-05-01 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Structures of Cd200/Cd200 Receptor Family and Implications for Topology, Regulation, and Evolution
Structure, 21, 2013
|
|
4BFI
 
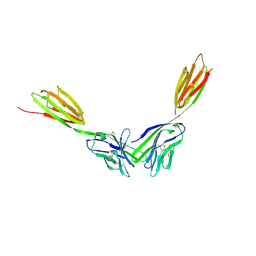 | | Structure of the complex of the extracellular portions of mouse CD200R and mouse CD200 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, CELL SURFACE GLYCOPROTEIN CD200 RECEPTOR 1, ... | | Authors: | Hatherley, D, Lea, S.M, Johnson, S, Barclay, A.N. | | Deposit date: | 2013-03-19 | | Release date: | 2013-05-01 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.22 Å) | | Cite: | Structures of Cd200/Cd200 Receptor Family and Implications for Topology, Regulation, and Evolution
Structure, 21, 2013
|
|
5VH9
 
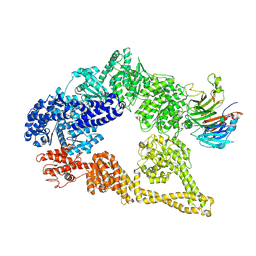 | | Cryo-EM structure of yeast cytoplasmic dynein-1 with Lis1 and ATP | | Descriptor: | Dynein heavy chain, cytoplasmic, Nuclear distribution protein PAC1 | | Authors: | Cianfrocco, M.A, DeSantis, M.E, Htet, Z.M, Tran, P.T, Reck-Peterson, S.L, Leschziner, A.E. | | Deposit date: | 2017-04-12 | | Release date: | 2017-09-06 | | Last modified: | 2020-01-01 | | Method: | ELECTRON MICROSCOPY (7.7 Å) | | Cite: | Lis1 Has Two Opposing Modes of Regulating Cytoplasmic Dynein.
Cell, 170, 2017
|
|
5VLJ
 
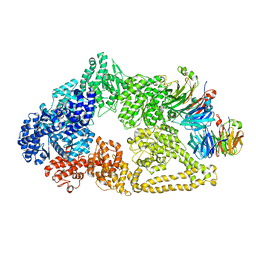 | | Cryo-EM structure of yeast cytoplasmic dynein with Walker B mutation at AAA3 in presence of ATP-VO4 | | Descriptor: | Dynein heavy chain, cytoplasmic, Nuclear distribution protein PAC1 | | Authors: | Cianfrocco, M.A, DeSantis, M.E, Htet, Z.M, Tran, P.T, Reck-Peterson, S.L, Leschziner, A.E. | | Deposit date: | 2017-04-25 | | Release date: | 2017-09-06 | | Last modified: | 2020-01-01 | | Method: | ELECTRON MICROSCOPY (10.5 Å) | | Cite: | Lis1 Has Two Opposing Modes of Regulating Cytoplasmic Dynein.
Cell, 170, 2017
|
|
3I80
 
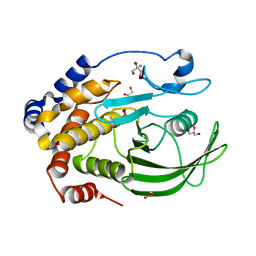 | | Protein Tyrosine Phosphatase 1B - Transition state analog for the second catalytic step | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLYCEROL, Tyrosine-protein phosphatase non-receptor type 1, ... | | Authors: | Brandao, T.A.S, Johnson, S.J, Hengge, A.C. | | Deposit date: | 2009-07-09 | | Release date: | 2010-03-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Insights into the reaction of protein-tyrosine phosphatase 1B: crystal structures for transition state analogs of both catalytic steps.
J.Biol.Chem., 285, 2010
|
|
4KLI
 
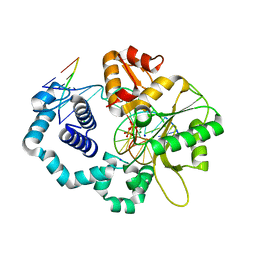 | | DNA polymerase beta matched product complex with Mg2+, 90 s | | Descriptor: | 5'-D(*CP*CP*GP*AP*CP*GP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*CP*C)-3', 5'-D(P*GP*TP*CP*GP*G)-3', ... | | Authors: | Freudenthal, B.D, Beard, W.A, Shock, D.D, Wilson, S.H. | | Deposit date: | 2013-05-07 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.597 Å) | | Cite: | Observing a DNA polymerase choose right from wrong.
Cell(Cambridge,Mass.), 154, 2013
|
|
4CMM
 
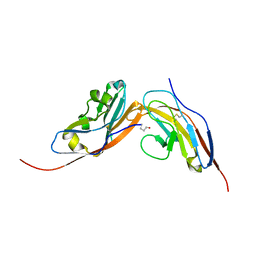 | | Structure of human CD47 in complex with human Signal Regulatory Protein (SIRP) alpha v1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, LEUKOCYTE SURFACE ANTIGEN CD47, TYROSINE-PROTEIN PHOSPHATASE NON-RECEPTOR TYPE SUBSTRATE 1 | | Authors: | Hatherley, D, Lea, S.M, Johnson, S, Barclay, A.N. | | Deposit date: | 2014-01-16 | | Release date: | 2014-02-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Polymorphisms in the Human Inhibitory Signal-Regulatory Protein Alpha Do not Affect Binding to its Ligand Cd47.
J.Biol.Chem., 289, 2014
|
|
7LV9
 
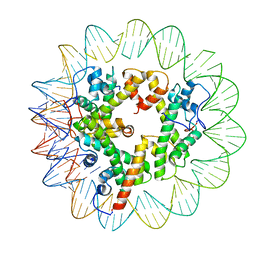 | | Marseillevirus heterotrimeric (hexameric) nucleosome | | Descriptor: | DNA (96-MER), Histone doublet Beta-Alpha (Alpha), Histone doublet Beta-Alpha (Beta), ... | | Authors: | Valencia-Sanchez, M.I, Abini-Agbomson, S, Armache, K.-J. | | Deposit date: | 2021-02-24 | | Release date: | 2021-05-05 | | Last modified: | 2021-05-26 | | Method: | ELECTRON MICROSCOPY (4.5 Å) | | Cite: | The structure of a virus-encoded nucleosome.
Nat.Struct.Mol.Biol., 28, 2021
|
|
7LV8
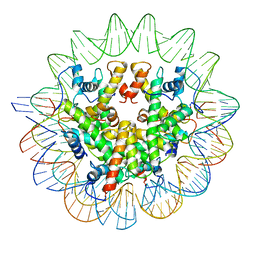 
 | | Structure of the Marseillevirus nucleosome | | Descriptor: | DNA (123-MER), Histone doublet Beta-Alpha (Alpha), Histone doublet Beta-Alpha (Beta), ... | | Authors: | Valencia-Sanchez, M.I, Abini-Agbomson, S, Armache, K.-J. | | Deposit date: | 2021-02-24 | | Release date: | 2021-05-05 | | Last modified: | 2021-05-26 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | The structure of a virus-encoded nucleosome.
Nat.Struct.Mol.Biol., 28, 2021
|
|
4BF3
 
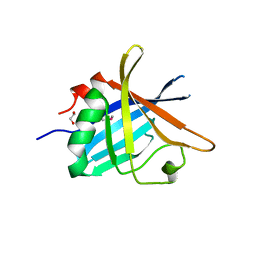 | | ErpC, a member of the complement regulator acquiring family of surface proteins from Borrelia burgdorfei, possesses an architecture previously unseen in this protein family. | | Descriptor: | 1,2-ETHANEDIOL, ERPC | | Authors: | Caesar, J.J.E, Johnson, S, Kraiczy, P, Lea, S.M. | | Deposit date: | 2013-03-14 | | Release date: | 2013-06-05 | | Last modified: | 2013-10-09 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Erpc, a Member of the Complement Regulator-Acquiring Family of Surface Proteins from Borrelia Burgdorferi, Possesses an Architecture Previously Unseen in This Protein Family.
Acta Crystallogr.,Sect.F, 69, 2013
|
|
4KLT
 
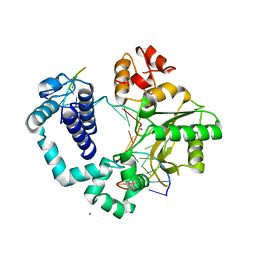 | | DNA polymerase beta mismatched product complex with Mn2+, 30 min | | Descriptor: | 5'-D(*CP*CP*GP*AP*CP*GP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*CP*A)-3', 5'-D(P*GP*TP*CP*GP*G)-3', ... | | Authors: | Freudenthal, B.D, Beard, W.A, Shock, D.D, Wilson, S.H. | | Deposit date: | 2013-05-07 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.979 Å) | | Cite: | Observing a DNA polymerase choose right from wrong.
Cell(Cambridge,Mass.), 154, 2013
|
|
4KLJ
 
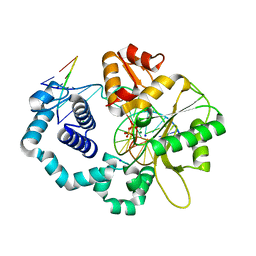 | | DNA polymerase beta matched product complex with Mg2+, 5 min | | Descriptor: | 5'-D(*CP*CP*GP*AP*CP*GP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*CP*C)-3', 5'-D(P*GP*TP*CP*GP*G)-3', ... | | Authors: | Freudenthal, B.D, Beard, W.A, Shock, D.D, Wilson, S.H. | | Deposit date: | 2013-05-07 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.799 Å) | | Cite: | Observing a DNA polymerase choose right from wrong.
Cell(Cambridge,Mass.), 154, 2013
|
|
4KLO
 
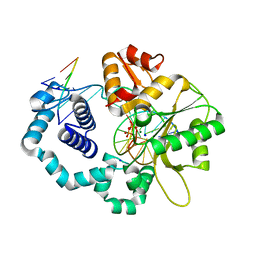 | | DNA polymerase beta matched nick complex with Mg2+ and PPi, 30 min | | Descriptor: | 5'-D(*CP*CP*GP*AP*CP*GP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*CP*C)-3', 5'-D(P*GP*TP*CP*GP*G)-3', ... | | Authors: | Freudenthal, B.D, Beard, W.A, Shock, D.D, Wilson, S.H. | | Deposit date: | 2013-05-07 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.845 Å) | | Cite: | Observing a DNA polymerase choose right from wrong.
Cell(Cambridge,Mass.), 154, 2013
|
|
4KLS
 
 | | DNA polymerase beta mismatched reactant complex with Mn2+, 10 min | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, 5'-D(*CP*CP*GP*AP*CP*GP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*CP*A)-3', ... | | Authors: | Freudenthal, B.D, Beard, W.A, Shock, D.D, Wilson, S.H. | | Deposit date: | 2013-05-07 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.978 Å) | | Cite: | Observing a DNA polymerase choose right from wrong.
Cell(Cambridge,Mass.), 154, 2013
|
|
4KLM
 
 | | DNA polymerase beta matched product complex with Mg2+, 11 h | | Descriptor: | 5'-D(*CP*CP*GP*AP*CP*GP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*CP*C)-3', 5'-D(P*GP*TP*CP*GP*G)-3', ... | | Authors: | Freudenthal, B.D, Beard, W.A, Shock, D.D, Wilson, S.H. | | Deposit date: | 2013-05-07 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.747 Å) | | Cite: | Observing a DNA polymerase choose right from wrong.
Cell(Cambridge,Mass.), 154, 2013
|
|
8DYV
 
 | | Structure of human cytoplasmic dynein-1 bound to one Lis1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Cytoplasmic dynein 1 heavy chain 1, ... | | Authors: | Reimer, J.M, DeSantis, M, Reck-Peterson, S.L, Leschziner, A.E. | | Deposit date: | 2022-08-05 | | Release date: | 2023-02-01 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.97 Å) | | Cite: | Structures of human dynein in complex with the lissencephaly 1 protein, LIS1.
Elife, 12, 2023
|
|
