7O2Z
 
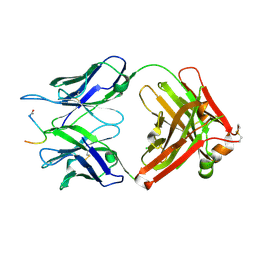 | | Crystal structure of the anti-PAS Fab 2.2 in complex with its epitope peptide | | Descriptor: | CHLORIDE ION, P/A#1 epitope peptide, anti-PAS Fab 2.2 chimeric heavy chain, ... | | Authors: | Schilz, J, Schiefner, A, Skerra, A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Molecular recognition of structurally disordered Pro/Ala-rich sequences (PAS) by antibodies involves an Ala residue at the hot spot of the epitope.
J.Mol.Biol., 433, 2021
|
|
3BX8
 
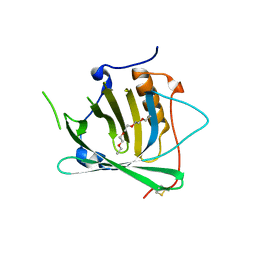 | | Engineered Human Lipocalin 2 (LCN2), apo-form | | Descriptor: | 3,6,9,12,15,18,21,24-OCTAOXAHEXACOSAN-1-OL, ENGINEERED HUMAN LIPOCALIN 2, PENTAETHYLENE GLYCOL | | Authors: | Schonfeld, D.L, Chatwell, L, Skerra, A. | | Deposit date: | 2008-01-11 | | Release date: | 2009-01-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | High affinity molecular recognition and functional blockade of CTLA-4 by an engineered human lipocalin
To be Published
|
|
7O31
 
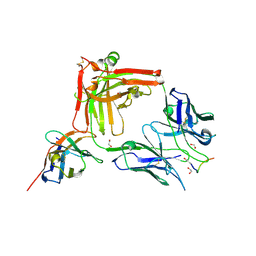 | | Crystal structure of the anti-PAS Fab 1.2 in complex with its epitope peptide and the anti-Kappa VHH domain | | Descriptor: | 1,2-ETHANEDIOL, PAS#1 epitope peptide, anti-Kappa VHH domain, ... | | Authors: | Schilz, J, Schiefner, A, Skerra, A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Molecular recognition of structurally disordered Pro/Ala-rich sequences (PAS) by antibodies involves an Ala residue at the hot spot of the epitope.
J.Mol.Biol., 433, 2021
|
|
7O33
 
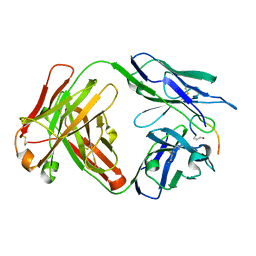 | | Crystal structure of the anti-PAS Fab 3.1 in complex with its epitope peptide | | Descriptor: | APSA epitope peptide, anti-PAS Fab 3.1 chimeric heavy chain, anti-PAS Fab 3.1 chimeric light chain | | Authors: | Schilz, J, Skerra, A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Molecular recognition of structurally disordered Pro/Ala-rich sequences (PAS) by antibodies involves an Ala residue at the hot spot of the epitope.
J.Mol.Biol., 433, 2021
|
|
7O30
 
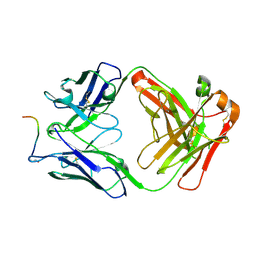 | |
6SUA
 
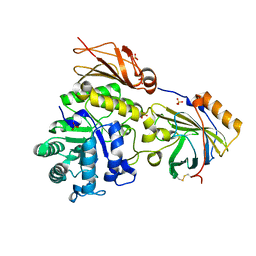 | |
5BPK
 
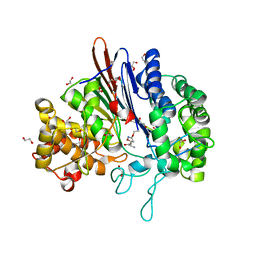 | | Varying binding modes of inhibitors and structural differences in the binding pockets of different gamma-glutamyltranspeptidases | | Descriptor: | (2S)-amino[(5S)-4,5-dihydro-1,2-oxazol-5-yl]acetic acid, 1,2-ETHANEDIOL, Gamma-glutamyltranspeptidase (Ggt) | | Authors: | Bolz, C, Bach, N.C, Meyer, H, Mueller, G, Dawidowski, M, Popowicz, G, Sieber, S.A, Skerra, A, Gerhard, M. | | Deposit date: | 2015-05-28 | | Release date: | 2016-05-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Varying binding modes of inhibitors and structural differences in the binding pockets of different gamma-glutamyltranspeptidases
To Be Published
|
|
1RSU
 
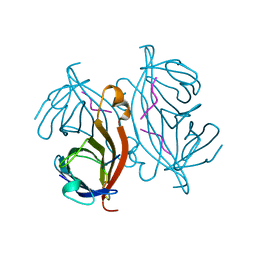 | |
1RST
 
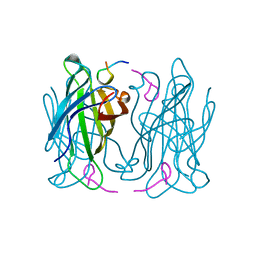 | |
5E9O
 | | Spirochaeta thermophila X module - CBM64 - mutant G504A | | Descriptor: | 5-amino-2,4,6-triiodobenzene-1,3-dicarboxylic acid, Cellulase, glycosyl hydrolase family 5, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2015-10-15 | | Release date: | 2016-03-02 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis for cellulose binding by the type A carbohydrate-binding module 64 of Spirochaeta thermophila.
Proteins, 84, 2016
|
|
5E9P
 
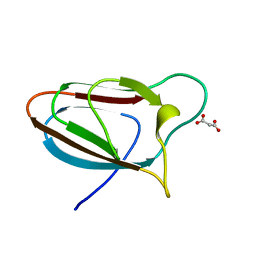 | | Spirochaeta thermophila X module - CBM64 - wildtype | | Descriptor: | Cellulase, glycosyl hydrolase family 5, TPS linker, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2015-10-15 | | Release date: | 2016-03-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structural basis for cellulose binding by the type A carbohydrate-binding module 64 of Spirochaeta thermophila.
Proteins, 84, 2016
|
|
2VO9
 
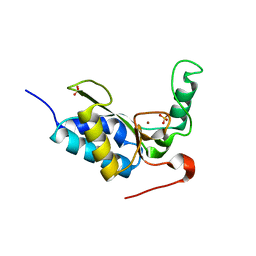 | | CRYSTAL STRUCTURE OF THE ENZYMATICALLY ACTIVE DOMAIN OF THE LISTERIA MONOCYTOGENES BACTERIOPHAGE 500 ENDOLYSIN PLY500 | | Descriptor: | L-ALANYL-D-GLUTAMATE PEPTIDASE, SULFATE ION, ZINC ION | | Authors: | Korndoerfer, I.P, Kanitz, A, Skerra, A. | | Deposit date: | 2008-02-09 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Analysis of the L-Alanoyl-D-Glutamate Endopeptidase Domain of Listeria Bacteriophage Endolysin Ply500 Reveals a New Member of the Las Peptidase Family.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
3BX7
 
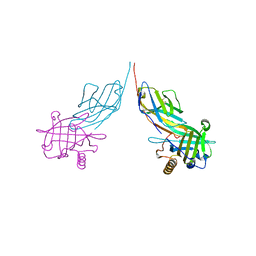 | |
1XKI
 
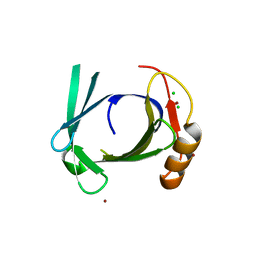 | | Crystal structure of human tear lipocalin/von Ebners gland protein | | Descriptor: | CHLORIDE ION, Von Ebner's gland protein, ZINC ION | | Authors: | Breustedt, D.A, Korndoerfer, I.P, Redl, B, Skerra, A. | | Deposit date: | 2004-09-29 | | Release date: | 2004-10-19 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The 1.8-A crystal structure of human tear lipocalin reveals an extended branched cavity with capacity for multiple ligands
J.Biol.Chem., 280, 2005
|
|
1XOV
 
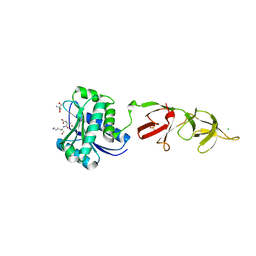 | |
6S8V
 
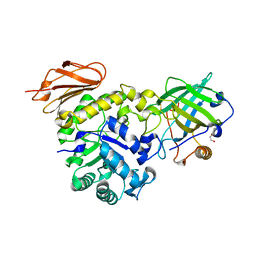 | | Structure of the high affinity Anticalin P3D11 in complex with the human CD98 heavy chain ectodomain | | Descriptor: | 1,2-ETHANEDIOL, 4F2 cell-surface antigen heavy chain, Neutrophil gelatinase-associated lipocalin | | Authors: | Schiefner, A, Deuschle, F.-C, Skerra, A. | | Deposit date: | 2019-07-10 | | Release date: | 2020-03-04 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Development of a high affinity Anticalin®directed against human CD98hc for theranostic applications.
Theranostics, 10, 2020
|
|
1XK4
 
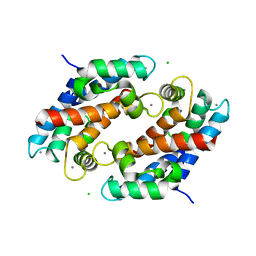 | | Crystal structure of human calprotectin(S100A8/S100A9) | | Descriptor: | CALCIUM ION, CHLORIDE ION, CITRATE ANION, ... | | Authors: | Korndoerfer, I.P, Brueckner, F, Skerra, A. | | Deposit date: | 2004-09-26 | | Release date: | 2005-10-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of the human (S100A8/S100A9)2 heterotetramer, calprotectin, illustrates how conformational changes of interacting alpha-helices can determine specific association of two EF-hand proteins
J.Mol.Biol., 370, 2007
|
|
1KXO
 | | ENGINEERED LIPOCALIN DIGA16 : APO-FORM | | Descriptor: | DigA16 | | Authors: | Korndoerfer, I.P, Skerra, A. | | Deposit date: | 2002-02-01 | | Release date: | 2003-06-10 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural mechanism of specific ligand recognition by a lipocalin tailored for the complexation of digoxigenin.
J.Mol.Biol., 330, 2003
|
|
1LKE
 
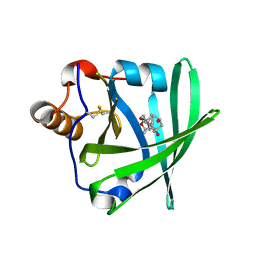 | |
1LNM
 
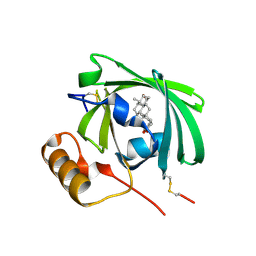 | |
4H8I
 
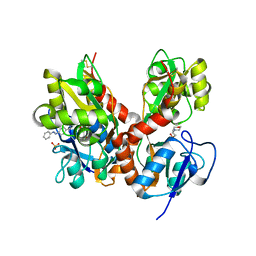 | | Structure of GluK2-LBD in complex with GluAzo | | Descriptor: | (4R)-4-[(2E)-3-{4-[(E)-phenyldiazenyl]phenyl}prop-2-en-1-yl]-L-glutamic acid, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CHLORIDE ION, ... | | Authors: | Reiter, A, Skerra, A, Trauner, D, Schiefner, A. | | Deposit date: | 2012-09-22 | | Release date: | 2013-09-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A photoswitchable neurotransmitter analogue bound to its receptor.
Biochemistry, 52, 2013
|
|
2HZQ
 
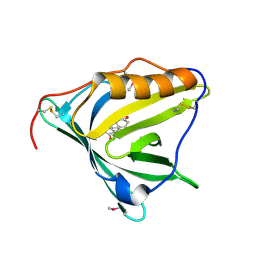 | |
2HZR
 
 | | Crystal structure of human apolipoprotein D (ApoD) | | Descriptor: | Apolipoprotein D | | Authors: | Eichinger, A, Skerra, A. | | Deposit date: | 2006-08-09 | | Release date: | 2007-08-14 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural insight into the dual ligand specificity and mode of high density lipoprotein association of apolipoprotein d.
J.Biol.Chem., 282, 2007
|
|
1SG2
 
 | |
4H8J
 
 | | Structure of GluA2-LBD in complex with MES | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Glutamate receptor 2, ... | | Authors: | Reiter, A, Skerra, A, Trauner, D, Schiefner, A. | | Deposit date: | 2012-09-22 | | Release date: | 2013-09-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis of an artificial photoreceptor
To be Published
|
|
