1WQ6
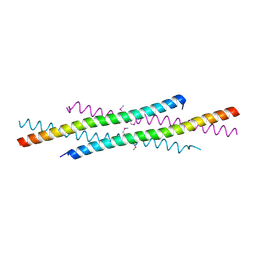 
 | | The tetramer structure of the nervy homolgy two (NHR2) domain of AML1-ETO is critical for AML1-ETO'S activity | | Descriptor: | AML1-ETO | | Authors: | Liu, Y, Cheney, M.D, Chruszcz, M, Lukasik, S.M, Hartman, K.L, Laue, T.M, Dauter, Z, Minor, W, Speck, N.A, Bushweller, J.H. | | Deposit date: | 2004-09-23 | | Release date: | 2005-10-04 | | Last modified: | 2022-04-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The tetramer structure of the Nervy homology two domain, NHR2, is critical for AML1/ETO's activity
Cancer Cell, 9, 2006
|
|
1W9H
 
 | | The Structure of a Piwi protein from Archaeoglobus fulgidus. | | Descriptor: | CADMIUM ION, CHLORIDE ION, HYPOTHETICAL PROTEIN AF1318, ... | | Authors: | Parker, J.S, Roe, S.M, Barford, D. | | Deposit date: | 2004-10-13 | | Release date: | 2005-01-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of a Piwi Protein Suggests Mechanisms for Sirna Recognition and Slicer Activity
Embo J., 23, 2004
|
|
1W4S
 
 | |
1UW3
 
 | | The crystal structure of the globular domain of sheep prion protein | | Descriptor: | GLUTATHIONE, PHOSPHATE ION, PRION PROTEIN | | Authors: | Haire, L.F, Whyte, S.M, Vasisht, N, Gill, A.C, Verma, C, Dodson, E.J, Dodson, G.G, Bayley, P.M. | | Deposit date: | 2004-01-29 | | Release date: | 2004-03-25 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The Crystal Structure of the Globular Domain of Sheep Prion Protein
J.Mol.Biol., 336, 2004
|
|
1UWH
 
 | | The complex of wild type B-RAF and BAY439006. | | Descriptor: | 4-{4-[({[4-CHLORO-3-(TRIFLUOROMETHYL)PHENYL]AMINO}CARBONYL)AMINO]PHENOXY}-N-METHYLPYRIDINE-2-CARBOXAMIDE, B-RAF PROTO-ONCOGENE SERINE/THREONINE-PROTEIN KINASE, CHLORIDE ION | | Authors: | Barford, D, Roe, S.M, Wan, P.T.C, Cancer Genome Project | | Deposit date: | 2004-02-05 | | Release date: | 2004-03-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Mechanism of Activation of the Raf-Erk Signaling Pathway by Oncogenic Mutations of B-Raf
Cell(Cambridge,Mass.), 116, 2004
|
|
1UNN
 
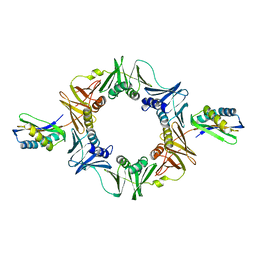 | |
1UOV
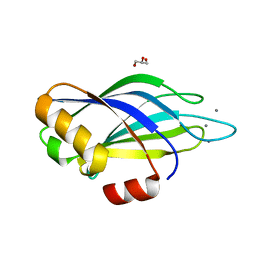 
 | | Calcium binding domain C2B | | Descriptor: | CALCIUM ION, GLYCEROL, SYNAPTOTAGMIN I | | Authors: | Cheng, Y, Sequeira, S.M, Sollner, T.H, Patel, D.J. | | Deposit date: | 2003-09-24 | | Release date: | 2004-09-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystallographic Identification of Ca2+ and Sr2+ Coordination Sites in Synaptotagmin I C2B Domain
Protein Sci., 13, 2004
|
|
1UWJ
 
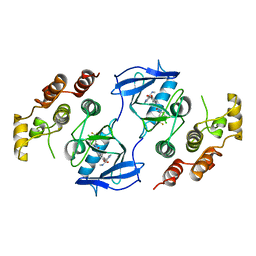 | | The complex of mutant V599E B-RAF and BAY439006. | | Descriptor: | 4-{4-[({[4-CHLORO-3-(TRIFLUOROMETHYL)PHENYL]AMINO}CARBONYL)AMINO]PHENOXY}-N-METHYLPYRIDINE-2-CARBOXAMIDE, B-RAF PROTO-ONCOGENE SERINE/THREONINE-PROTEIN KINASE | | Authors: | Barford, D, Roe, S.M, Wan, P.T.C, Cancer Genome Project | | Deposit date: | 2004-02-05 | | Release date: | 2004-03-19 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Mechanism of Activation of the Raf-Erk Signaling Pathway by Oncogenic Mutations of B-Raf
Cell(Cambridge,Mass.), 116, 2004
|
|
1VCJ
 
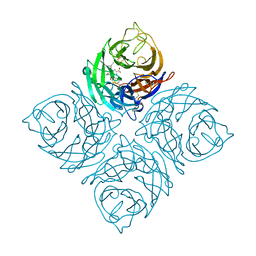 | | Influenza B virus neuraminidase complexed with 1-(4-Carboxy-2-(3-pentylamino)phenyl)-5-aminomethyl-5-hydroxymethyl-pyrrolidin-2-one | | Descriptor: | 4-[(2R)-2-(AMINOMETHYL)-2-(HYDROXYMETHYL)-5-OXOPYRROLIDIN-1-YL]-3-[(1-ETHYLPROPYL)AMINO]BENZOIC ACID, NEURAMINIDASE | | Authors: | Lommer, B.S, Ali, S.M, Bajpai, S.N, Brouillette, W.J, Air, G.M, Luo, M. | | Deposit date: | 2004-03-09 | | Release date: | 2004-03-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | A benzoic acid inhibitor induces a novel conformational change in the active site of Influenza B virus neuraminidase.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1UPN
 
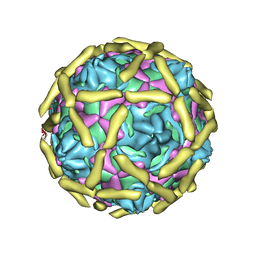 | | COMPLEX OF ECHOVIRUS TYPE 12 WITH DOMAINS 3 AND 4 OF ITS RECEPTOR DECAY ACCELERATING FACTOR (CD55) BY CRYO ELECTRON MICROSCOPY AT 16 A | | Descriptor: | COMPLEMENT DECAY-ACCELERATING FACTOR, ECHOVIRUS 11 COAT PROTEIN VP1, ECHOVIRUS 11 COAT PROTEIN VP2, ... | | Authors: | Bhella, D, Goodfellow, I.G, Roversi, P, Pettigrew, D, Chaudry, Y, Evans, D.J, Lea, S.M. | | Deposit date: | 2003-10-08 | | Release date: | 2004-01-07 | | Last modified: | 2017-08-30 | | Method: | ELECTRON MICROSCOPY (16 Å) | | Cite: | The Structure of Echovirus Type 12 Bound to a Two-Domain Fragment of its Cellular Attachment Protein Decay-Accelerating Factor (Cd 55)
J.Biol.Chem., 279, 2004
|
|
1UWX
 
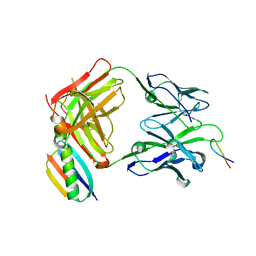 | | P1.2 serosubtype antigen derived from N. meningitidis PorA in complex with Fab fragment | | Descriptor: | ANTIBODY, CLASS 1 OUTER MEMBRANE PROTEIN VARIABLE REGION 2, PROTEIN G-PRIME | | Authors: | Tzitzilonis, C, Prince, S.M, Collins, R.F, Maiden, M.C.J, Feavers, I.M, Derrick, J.P. | | Deposit date: | 2004-02-12 | | Release date: | 2005-06-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural Variation and Immune Recognition of the P1.2 Subtype Meningococcal Antigen.
Proteins: Struct., Funct., Bioinf., 62, 2005
|
|
1UV5
 
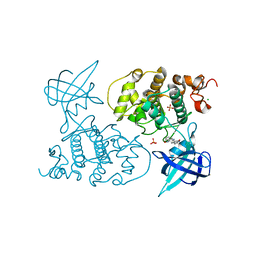 | | GLYCOGEN SYNTHASE KINASE 3 BETA COMPLEXED WITH 6-BROMOINDIRUBIN-3'-OXIME | | Descriptor: | 6-BROMOINDIRUBIN-3'-OXIME, CHLORIDE ION, COBALT (II) ION, ... | | Authors: | Dajani, R, Pearl, L.H, Roe, S.M. | | Deposit date: | 2004-01-14 | | Release date: | 2004-01-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Gsk-3-Selective Inhibitors Derived from Tyrian Purple Indurubins
Chem.Biol., 10, 2003
|
|
1US7
 
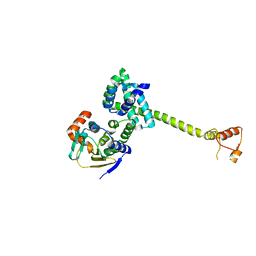 | | Complex of Hsp90 and P50 | | Descriptor: | HEAT SHOCK PROTEIN HSP82, HSP90 CO-CHAPERONE CDC37 | | Authors: | Roe, S.M, Ali, M.M.U, Pearl, L.H. | | Deposit date: | 2003-11-20 | | Release date: | 2004-01-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Mechanism of Hsp90 Regulation by the Protein Kinase-Specific Cochaperone p50(Cdc37)
Cell(Cambridge,Mass.), 116, 2004
|
|
1USV
 
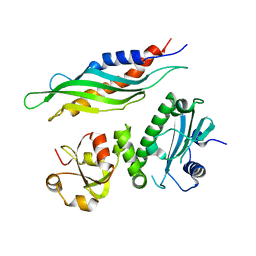 | |
1UOW
 
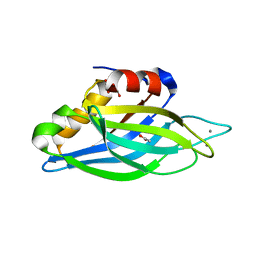 | | Calcium binding domain C2B | | Descriptor: | ACETATE ION, CALCIUM ION, GLYCEROL, ... | | Authors: | Cheng, Y, Sequeira, S.M, Sollner, T.H, Patel, D.J. | | Deposit date: | 2003-09-24 | | Release date: | 2004-09-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Crystallographic Identification of Ca2+ and Sr2+ Coordination Sites in Synaptotagmin I C2B Domain
Protein Sci., 13, 2004
|
|
1WYZ
 
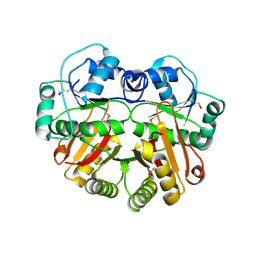 | | X-Ray structure of the putative methyltransferase from Bacteroides thetaiotaomicron VPI-5482 at the resolution 2.5 A. Norteast Structural Genomics Consortium target Btr28 | | Descriptor: | putative S-adenosylmethionine-dependent methyltransferase | | Authors: | Kuzin, A.P, Chen, Y, Forouhar, F, Vorobiev, S.M, Acton, T, Ma, L.-C, Xiao, R, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2005-02-21 | | Release date: | 2005-03-08 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-Ray structure of the putative methyltransferase from Bacteroides thetaiotaomicron VPI-5482 at the resolution 2.5 A. Norteast Structural Genomics Consortium target Btr28
To be Published
|
|
1USU
 
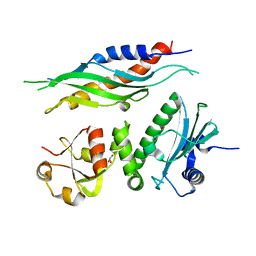 | |
1UNP
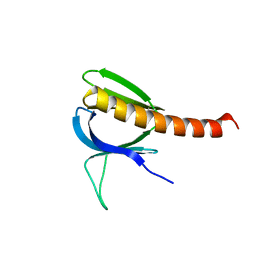 
 | | Crystal structure of the pleckstrin homology domain of PKB alpha | | Descriptor: | RAC-ALPHA SERINE/THREONINE KINASE | | Authors: | Milburn, C.C, Deak, M, Kelly, S.M, Price, N.C, Alessi, D.R, van Aalten, D.M.F. | | Deposit date: | 2003-09-12 | | Release date: | 2004-09-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Binding of phosphatidylinositol 3,4,5-trisphosphate to the pleckstrin homology domain of protein kinase B induces a conformational change.
Biochem. J., 375, 2003
|
|
1TOQ
 
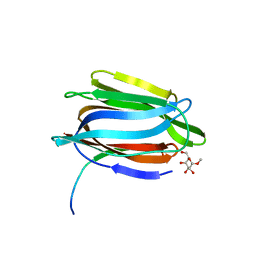 | | CRYSTAL STRUCTURE OF A GALACTOSE SPECIFIC LECTIN FROM ARTOCARPUS HIRSUTA IN COMPLEX WITH METHYL-a-D-GALACTOSE | | Descriptor: | AGGLUTININ ALPHA CHAIN, AGGLUTININ BETA CHAIN, methyl alpha-D-galactopyranoside | | Authors: | Rao, K.N, Suresh, C.G, Katre, U.V, Gaikwad, S.M, Khan, M.I. | | Deposit date: | 2004-06-15 | | Release date: | 2004-08-03 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Two orthorhombic crystal structures of a galactose-specific lectin from Artocarpus hirsuta in complex with methyl-alpha-D-galactose.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1TTZ
 
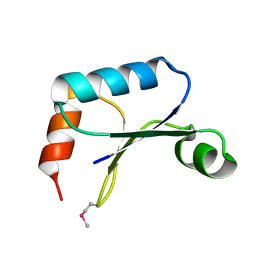 | | X-ray structure of Northeast Structural Genomics target protein XcR50 from X. campestris | | Descriptor: | conserved hypothetical protein | | Authors: | Kuzin, A.P, Vorobiev, S.M, Lee, I, Acton, T.B, Ho, C.K, Cooper, B, Ma, L.-C, Xiao, R, Montelione, G, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-06-23 | | Release date: | 2004-07-13 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | X-ray structure of Northeast Structural Genomics target protein XcR50 from X. campestris
To be Published
|
|
1TO0
 
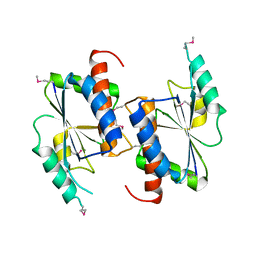 | | X-ray structure of Northeast Structural Genomics target protein sr145 from Bacillus subtilis | | Descriptor: | Hypothetical UPF0247 protein yyda | | Authors: | Kuzin, A.P, Edstrom, W, Vorobiev, S.M, Shastry, R, Ma, L.-C, Xiao, R, Acton, T, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-06-11 | | Release date: | 2004-07-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-ray structure of Northeast Structural Genomics target protein sr145 from Bacillus subtilis
To be Published
|
|
1TZA
 
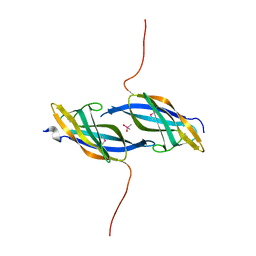 | | X-ray structure of Northeast Structural Genomics Consortium target SoR45 | | Descriptor: | CACODYLATE ION, apaG protein | | Authors: | Kuzin, A.P, Vorobiev, S.M, Edstrom, W, Acton, T.B, Shastry, R, Ma, L.-C, Cooper, B, Xiao, R, Montelione, G, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-07-09 | | Release date: | 2004-07-27 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | X-ray structure of Northeast Structural Genomics Consortium
target SoR45
To be Published
|
|
1XBH
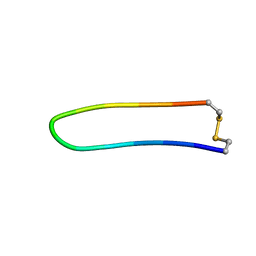 
 | | A BETA-HAIRPIN MIMIC FROM FCERI-ALPHA-CYCLO(L-262) | | Descriptor: | PROTEIN (CYCLO(L-262)) | | Authors: | Mcdonnell, J.M, Fushman, D, Cahill, S.M, Sutton, B.J, Cowburn, D. | | Deposit date: | 1999-02-17 | | Release date: | 1999-02-21 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution Structures of FceRI Alpha-Chain Mimics: A Beta-Hairpin Peptide and Its Retroenantiomer
J.Am.Chem.Soc., 119, 1997
|
|
1X9Q
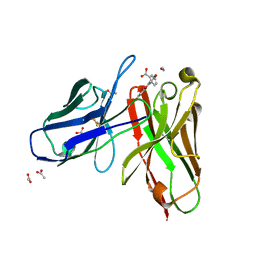 
 | | 4m5.3 anti-fluorescein single chain antibody fragment (scFv) | | Descriptor: | 2-(6-HYDROXY-3-OXO-3H-XANTHEN-9-YL)-BENZOIC ACID, 4m5.3 anti-fluorescein single chain antibody fragment, ACETATE ION | | Authors: | Midelfort, K.S, Hernandez, H.H, Lippow, S.M, Tidor, B, Drennan, C.L, Wittrup, K.D. | | Deposit date: | 2004-08-24 | | Release date: | 2005-03-22 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Substantial energetic improvement with minimal structural perturbation in a high affinity mutant antibody
J.Mol.Biol., 343, 2004
|
|
1XQ4
 
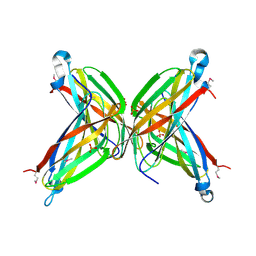 | | Crystal Structure of the Putative ApaA Protein from Bordetella pertussis, Northeast Structural Genomics Target BeR40 | | Descriptor: | PHOSPHATE ION, Protein apaG | | Authors: | Forouhar, F, Yong, W, Vorobiev, S.M, Xiao, R, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-10-11 | | Release date: | 2004-11-09 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of the Putative ApaA Protein from Bordetella pertussis, Northeast Structural Genomics Target BeR40
To be Published
|
|
