7R2E
 
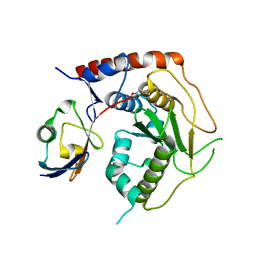 | | Structure of human Senp7 with SUMO2 | | Descriptor: | Sentrin-specific protease 7, Small ubiquitin-related modifier 3, prop-2-en-1-amine | | Authors: | Reverter, D, Li, Y. | | Deposit date: | 2022-02-04 | | Release date: | 2022-12-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Structural Basis for the SUMO2 Isoform Specificity of SENP7.
J.Mol.Biol., 434, 2022
|
|
8RQI
 
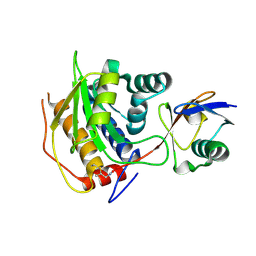 | | Structure of Rhizobium NopD with ubiquitin | | Descriptor: | NopD, Polyubiquitin-B, prop-2-en-1-amine | | Authors: | Reverter, D, Li, Y. | | Deposit date: | 2024-01-18 | | Release date: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Broad-spectrum ubiquitin/ubiquitin-like deconjugation activity of the rhizobial effector NopD from Bradyrhizobium (sp. XS1150).
Commun Biol, 7, 2024
|
|
6EOA
 
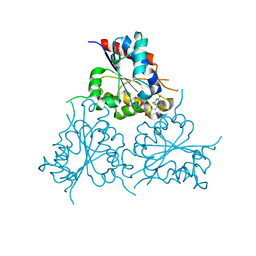 | | Crystal Structure of HAL3 from Cryptococcus neoformans | | Descriptor: | FLAVIN MONONUCLEOTIDE, Phosphopantothenoylcysteine decarboxylase | | Authors: | Reverter, D, Zhang, C, Molero, C, Arino, J. | | Deposit date: | 2017-10-09 | | Release date: | 2018-10-31 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Characterization of the atypical Ppz/Hal3 phosphatase system from the pathogenic fungus Cryptococcus neoformans.
Mol.Microbiol., 111, 2019
|
|
8OI3
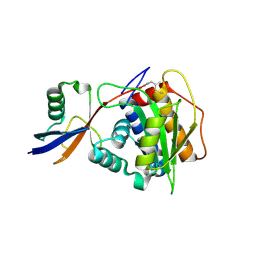 
 | | Structure of NopD with AtSUMO2 | | Descriptor: | Small ubiquitin-related modifier 2, Type III effector, prop-2-en-1-amine | | Authors: | Reverter, D, Li, Y. | | Deposit date: | 2023-03-22 | | Release date: | 2024-04-03 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Broad-spectrum ubiquitin/ubiquitin-like deconjugation activity of the rhizobial effector NopD from Bradyrhizobium (sp. XS1150).
Commun Biol, 7, 2024
|
|
7P99
 
 | | Structure of human USPL1 in complex with SUMO2 | | Descriptor: | SUMO-specific isopeptidase USPL1, Small ubiquitin-related modifier, ZINC ION | | Authors: | Reverter, D, Ying, L. | | Deposit date: | 2021-07-26 | | Release date: | 2022-03-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for the SUMO protease activity of the atypical ubiquitin-specific protease USPL1.
Nat Commun, 13, 2022
|
|
5A6I
 
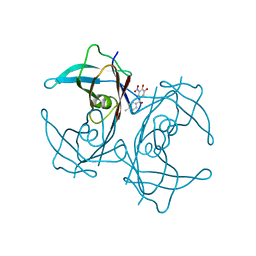 | | V122I Transthyretin structure in complex with Tolcalpone | | Descriptor: | TRANSTHYRETIN, Tolcapone | | Authors: | Reverter, D, Gallego, P, Santana, R, Ventura, S. | | Deposit date: | 2015-06-26 | | Release date: | 2016-02-24 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Repositioning Tolcapone as a Potent Inhibitor of Transthyretin Amyloidogenesis and its Associated Cellular Toxicity
Nat.Commun., 7, 2016
|
|
1DTV
 
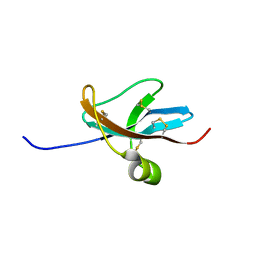 | | NMR STRUCTURE OF THE LEECH CARBOXYPEPTIDASE INHIBITOR (LCI) | | Descriptor: | CARBOXYPEPTIDASE INHIBITOR | | Authors: | Reverter, D, Fernandez-Catalan, C, Bode, W, Holak, T.A, Aviles, F.X. | | Deposit date: | 2000-01-13 | | Release date: | 2000-07-19 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | Structure of a novel leech carboxypeptidase inhibitor determined free in solution and in complex with human carboxypeptidase A2.
Nat.Struct.Biol., 7, 2000
|
|
5O71
 
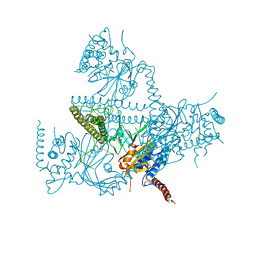 | | Crystal structure of human USP25 | | Descriptor: | Ubiquitin carboxyl-terminal hydrolase 25 | | Authors: | Reverter, D, Liu, B. | | Deposit date: | 2017-06-07 | | Release date: | 2018-06-20 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.283 Å) | | Cite: | A quaternary tetramer assembly inhibits the deubiquitinating activity of USP25.
Nat Commun, 9, 2018
|
|
6RO9
 | | Human spectrin SH3 domain D48G, E7V, K60V | | Descriptor: | Spectrin alpha, non-erythrocytic 1 | | Authors: | Reverter, D, Navarro, S, Ventura, S. | | Deposit date: | 2019-05-10 | | Release date: | 2020-06-03 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.814 Å) | | Cite: | Human spectrin SH3 domain D48G, E7V, K60V
To Be Published
|
|
7AL8
 | |
1DTD
 
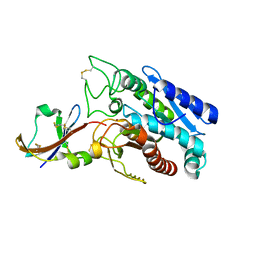 | | CRYSTAL STRUCTURE OF THE COMPLEX BETWEEN THE LEECH CARBOXYPEPTIDASE INHIBITOR AND THE HUMAN CARBOXYPEPTIDASE A2 (LCI-CPA2) | | Descriptor: | CARBOXYPEPTIDASE A2, GLUTAMIC ACID, METALLOCARBOXYPEPTIDASE INHIBITOR, ... | | Authors: | Reverter, D, Fernandez-Catalan, C, Bode, W, Holak, T.A, Aviles, F.X. | | Deposit date: | 2000-01-12 | | Release date: | 2000-07-12 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure of a novel leech carboxypeptidase inhibitor determined free in solution and in complex with human carboxypeptidase A2.
Nat.Struct.Biol., 7, 2000
|
|
1Z5S
 
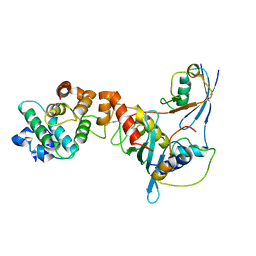 | | Crystal structure of a complex between UBC9, SUMO-1, RANGAP1 and NUP358/RANBP2 | | Descriptor: | Ran GTPase-activating protein 1, Ran-binding protein 2, Ubiquitin-conjugating enzyme E2 I, ... | | Authors: | Reverter, D, Lima, C.D. | | Deposit date: | 2005-03-19 | | Release date: | 2005-06-07 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | Insights into E3 ligase activity revealed by a SUMO-RanGAP1-Ubc9-Nup358 complex.
Nature, 435, 2005
|
|
1XT9
 
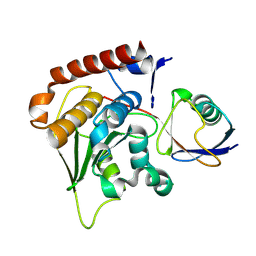 | | Crystal Structure of Den1 in complex with Nedd8 | | Descriptor: | Neddylin, Sentrin-specific protease 8 | | Authors: | Reverter, D, Wu, K, Erdene, T.G, Pan, Z.Q, Wilkinson, K.D, Lima, C.D. | | Deposit date: | 2004-10-21 | | Release date: | 2004-12-21 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of a Complex between Nedd8 and the Ulp/Senp Protease Family Member Den1.
J.Mol.Biol., 345, 2005
|
|
1TH0
 
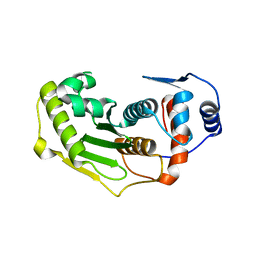 | | Structure of human Senp2 | | Descriptor: | Sentrin-specific protease 2 | | Authors: | Reverter, D, Lima, C.D. | | Deposit date: | 2004-05-31 | | Release date: | 2004-09-14 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A basis for SUMO protease specificity provided by analysis of human Senp2 and a
Senp2-SUMO complex
Structure, 12, 2004
|
|
1TGZ
 
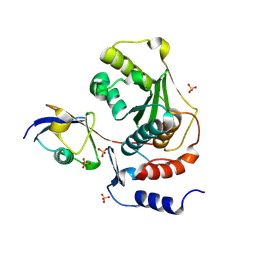 | | Structure of human Senp2 in complex with SUMO-1 | | Descriptor: | SULFATE ION, Sentrin-specific protease 2, Ubiquitin-like protein SMT3C | | Authors: | Reverter, D, Lima, C.D. | | Deposit date: | 2004-05-31 | | Release date: | 2004-09-14 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A basis for SUMO protease specificity provided by analysis of human Senp2 and a
Senp2-SUMO complex
Structure, 12, 2004
|
|
2IO2
 
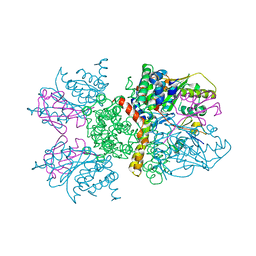 | | Crystal structure of human Senp2 in complex with RanGAP1-SUMO-1 | | Descriptor: | Ran GTPase-activating protein 1, Sentrin-specific protease 2, Small ubiquitin-related modifier 1 | | Authors: | Reverter, D, Lima, C.D. | | Deposit date: | 2006-10-09 | | Release date: | 2006-11-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates.
Nat.Struct.Mol.Biol., 13, 2006
|
|
2IO1
 
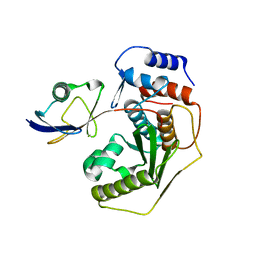 | | Crystal structure of human Senp2 in complex with preSUMO-3 | | Descriptor: | Sentrin-specific protease 2, Small ubiquitin-related modifier 3 precursor | | Authors: | Reverter, D, Lima, C.D. | | Deposit date: | 2006-10-09 | | Release date: | 2006-11-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates.
Nat.Struct.Mol.Biol., 13, 2006
|
|
2IO0
 
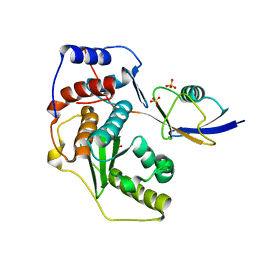 | | Crystal structure of human Senp2 in complex with preSUMO-2 | | Descriptor: | SULFATE ION, Sentrin-specific protease 2, Small ubiquitin-related modifier 2 precursor | | Authors: | Reverter, D, Lima, C.D. | | Deposit date: | 2006-10-09 | | Release date: | 2006-11-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates.
Nat.Struct.Mol.Biol., 13, 2006
|
|
2IO3
 
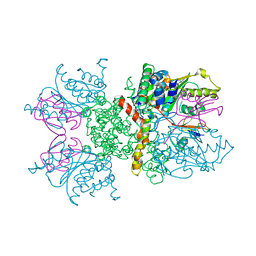 | | Crystal structure of human Senp2 in complex with RanGAP1-SUMO-2 | | Descriptor: | Ran GTPase-activating protein 1, Sentrin-specific protease 2, Small ubiquitin-related modifier 2 | | Authors: | Reverter, D, Lima, C.D. | | Deposit date: | 2006-10-09 | | Release date: | 2006-11-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates.
Nat.Struct.Mol.Biol., 13, 2006
|
|
7P47
 
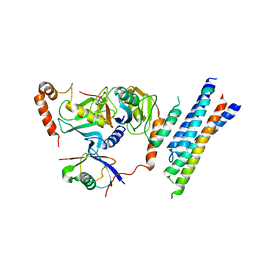 | | Structure of the E3 ligase Smc5/Nse2 in complex with Ubc9-SUMO thioester mimetic | | Descriptor: | E3 SUMO-protein ligase MMS21, SUMO-conjugating enzyme UBC9, Structural maintenance of chromosomes protein 5, ... | | Authors: | Lascorz, J, Varejao, N, Reverter, D. | | Deposit date: | 2021-07-09 | | Release date: | 2021-11-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.314 Å) | | Cite: | Structural basis for the E3 ligase activity enhancement of yeast Nse2 by SUMO-interacting motifs.
Nat Commun, 12, 2021
|
|
6FWW
 | | GFP/KKK. A redesigned GFP with improved solubility | | Descriptor: | Green fluorescent protein | | Authors: | Varejao, N, Lascorz, J, Gil-Garcia, M, Diaz-Caballero, M, Navarro, S, Ventura, S, Reverter, D. | | Deposit date: | 2018-03-07 | | Release date: | 2018-08-01 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.131 Å) | | Cite: | Combining Structural Aggregation Propensity and Stability Predictions To Redesign Protein Solubility.
Mol. Pharm., 15, 2018
|
|
8PM8
 
 | | V30M Transthyretin structure in complex with Tolcalpone | | Descriptor: | Tolcapone, Transthyretin | | Authors: | Varejao, N, Pinheiro, F, Pallares, I, Ventura, S, Reverter, D. | | Deposit date: | 2023-06-28 | | Release date: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | PITB: A high affinity transthyretin aggregation inhibitor with optimal pharmacokinetic properties.
Eur.J.Med.Chem., 261, 2023
|
|
8PMO
 
 | | Crystal structure of human V122I transthyretin in complex with PITB (Pharmacokinetically Improved TTR Binder) | | Descriptor: | (3-fluoranyl-5-oxidanyl-phenyl)-(3-methoxy-5-nitro-4-oxidanyl-phenyl)methanone, Transthyretin | | Authors: | Varejao, N, Pinheiro, F, Pallares, I, Ventura, S, Reverter, D. | | Deposit date: | 2023-06-29 | | Release date: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | PITB: A high affinity transthyretin aggregation inhibitor with optimal pharmacokinetic properties.
Eur.J.Med.Chem., 261, 2023
|
|
8PMA
 
 | | Crystal structure of human V30M transthyretin in complex with PITB (Pharmacokinetically Improved TTR Binder) | | Descriptor: | (3-fluoranyl-5-oxidanyl-phenyl)-(3-methoxy-5-nitro-4-oxidanyl-phenyl)methanone, Transthyretin | | Authors: | Varejao, N, Pinheiro, F, Pallares, I, Ventura, S, Reverter, D. | | Deposit date: | 2023-06-28 | | Release date: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | PITB: A high affinity transthyretin aggregation inhibitor with optimal pharmacokinetic properties.
Eur.J.Med.Chem., 261, 2023
|
|
8BQ3
 | |
