1VZ2
 
 | |
1VZ3
 
 | | PROLYL OLIGOPEPTIDASE FROM PORCINE BRAIN, T597C MUTANT | | Descriptor: | GLYCEROL, PROLYL ENDOPEPTIDASE | | Authors: | Rea, D, Fulop, V. | | Deposit date: | 2004-05-14 | | Release date: | 2004-07-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Concerted Structural Changes in the Peptidase and the Propeller Domains of Prolyl Oligopeptidase are Required for Substrate Binding
J.Mol.Biol., 340, 2004
|
|
1H2Z
 
 | |
1H2Y
 
 | |
2VWT
 
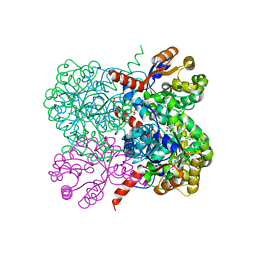 | | Crystal structure of YfaU, a metal ion dependent class II aldolase from Escherichia coli K12 - Mg-pyruvate product complex | | Descriptor: | GLYCEROL, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Rea, D, Rakus, J.F, Gerlt, J.A, Fulop, V, Bugg, T.D.H, Roper, D.I. | | Deposit date: | 2008-06-26 | | Release date: | 2008-09-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Crystal Structure and Functional Assignment of Yfau, a Metal Ion Dependent Class II Aldolase from Escherichia Coli K12.
Biochemistry, 47, 2008
|
|
2V5J
 
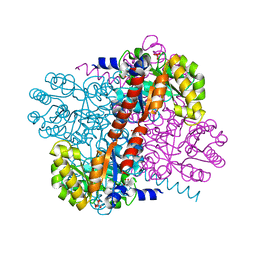 | | Apo Class II aldolase HpcH | | Descriptor: | 2,4-DIHYDROXYHEPT-2-ENE-1,7-DIOIC ACID ALDOLASE, GLYCEROL, PHOSPHATE ION | | Authors: | Rea, D, Fulop, V, Bugg, T.D.H, Roper, D.I. | | Deposit date: | 2007-07-05 | | Release date: | 2007-10-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure and Mechanism of Hpch: A Metal Ion Dependent Class II Aldolase from the Homoprotocatechuate Degradation Pathway of Escherichia Coli.
J.Mol.Biol., 373, 2007
|
|
2VWS
 
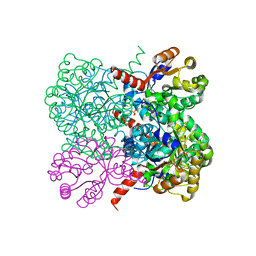 | | Crystal structure of YfaU, a metal ion dependent class II aldolase from Escherichia coli K12 | | Descriptor: | GLYCEROL, PHOSPHATE ION, YFAU, ... | | Authors: | Rea, D, Rakus, J.F, Gerlt, J.A, Fulop, V, Bugg, T.D.H, Roper, D.I. | | Deposit date: | 2008-06-26 | | Release date: | 2008-09-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Crystal Structure and Functional Assignment of Yfau, a Metal Ion Dependent Class II Aldolase from Escherichia Coli K12.
Biochemistry, 47, 2008
|
|
2V5K
 
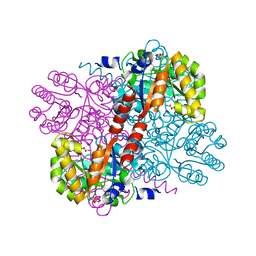 | | Class II aldolase HpcH - magnesium - oxamate complex | | Descriptor: | 2,4-DIHYDROXYHEPT-2-ENE-1,7-DIOIC ACID ALDOLASE, MAGNESIUM ION, OXAMIC ACID, ... | | Authors: | Rea, D, Fulop, V, Bugg, T.D.H, Roper, D.I. | | Deposit date: | 2007-07-06 | | Release date: | 2007-10-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure and Mechanism of Hpch: A Metal Ion Dependent Class II Aldolase from the Homoprotocatechuate Degradation Pathway of Escherichia Coli.
J.Mol.Biol., 373, 2007
|
|
1UOP
 
 | |
1UOQ
 
 | |
1UOO
 
 | |
1O6G
 
 | |
1O6F
 
 | |
8PO0
 
 | |
8PO3
 
 | |
8PNZ
 
 | |
8PO1
 
 | |
8PO4
 
 | |
7AEM
 
 | | Studies Towards a Reversible EGFR C797S Triple Mutant Inhibitor Series | | Descriptor: | 5-chloro-N~4~-[2-(dimethylphosphoryl)phenyl]-N~2~-{2-methoxy-4-[4-(4-methylpiperazin-1-yl)piperidin-1-yl]phenyl}pyrimidine-2,4-diamine, Epidermal growth factor receptor | | Authors: | Hargreaves, D. | | Deposit date: | 2020-09-17 | | Release date: | 2021-04-07 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Abstract 4451: Evaluation of the therapeutic potential of phosphine oxide pyrazole inhibitors in tumors harboring EGFR C797S mutation
Cancer Res., 79, 2019
|
|
7AEI
 
 | | Studies Towards a Reversible EGFR C797S Triple Mutant Inhibitor Series | | Descriptor: | 5-chloranyl-~{N}2-[4-[4-(dimethylamino)piperidin-1-yl]-2-methoxy-5-(1-methylpyrazol-4-yl)phenyl]-~{N}4-(2-dimethylphosphorylphenyl)pyrimidine-2,4-diamine, Epidermal growth factor receptor | | Authors: | Hargreaves, D. | | Deposit date: | 2020-09-17 | | Release date: | 2021-06-02 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Abstract 4451: Evaluation of the therapeutic potential of phosphine oxide pyrazole inhibitors in tumors harboring EGFR C797S mutation
Cancer Res., 79, 2019
|
|
8PO2
 
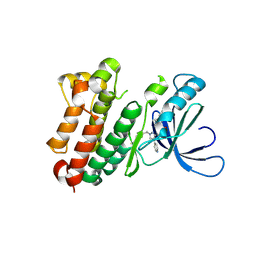 | |
1M1F
 
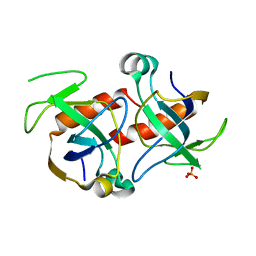 | | Kid toxin protein from E.coli plasmid R1 | | Descriptor: | Kid toxin protein, PHOSPHATE ION | | Authors: | Hargreaves, D, Santos-Sierra, S, Giraldo, R, Sabariegos-Jareno, R, de la Cueva-Mendez, G, Boelens, R, Diaz-Orejas, R, Rafferty, J.B. | | Deposit date: | 2002-06-19 | | Release date: | 2002-11-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural and Functional Analysis of the Kid toxin protein from E.coli plasmid R1
Structure, 10, 2002
|
|
9FKZ
 
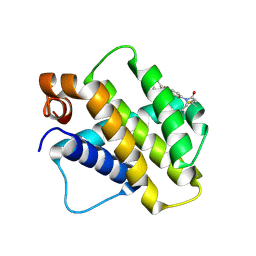 | | Discovery of a Series of Covalent, Cell Active Bfl-1 Inhibitors | | Descriptor: | Bcl-2-related protein A1, ~{N}-[4-[(1~{R},3~{R})-3-azanylcyclopentyl]oxyphenyl]-~{N}-[(4-chlorophenyl)methyl]prop-2-enamide | | Authors: | Hargreaves, D. | | Deposit date: | 2024-06-04 | | Release date: | 2024-10-02 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.684 Å) | | Cite: | Structure-Based Optimization of a Series of Covalent, Cell Active Bfl-1 Inhibitors.
J.Med.Chem., 67, 2024
|
|
9FKY
 
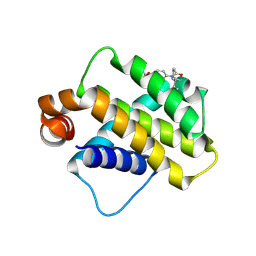 | | Discovery of a Series of Covalent, Cell Active Bfl-1 Inhibitors | | Descriptor: | Bcl-2-related protein A1, ~{N}-[4-[(1~{R},3~{R})-3-azanylcyclopentyl]oxyphenyl]-~{N}-[(1~{S})-1-[3-cyano-4-(trifluoromethyl)phenyl]ethyl]propanamide | | Authors: | Hargreaves, D. | | Deposit date: | 2024-06-04 | | Release date: | 2024-10-02 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.557 Å) | | Cite: | Structure-Based Optimization of a Series of Covalent, Cell Active Bfl-1 Inhibitors.
J.Med.Chem., 67, 2024
|
|
9FL0
 
 | | Discovery of a Series of Covalent, Cell Active Bfl-1 Inhibitors | | Descriptor: | Bcl-2-related protein A1, ~{N}-[(4-chlorophenyl)methyl]-~{N}-(4-fluorophenyl)prop-2-enamide | | Authors: | Hargreaves, D. | | Deposit date: | 2024-06-04 | | Release date: | 2024-10-02 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Structure-Based Optimization of a Series of Covalent, Cell Active Bfl-1 Inhibitors.
J.Med.Chem., 67, 2024
|
|
