7XWN
 
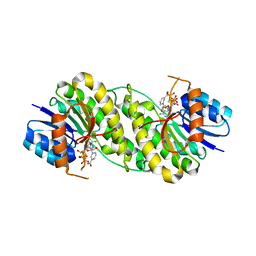 | | structure of patulin-detoxifying enzyme Y155F/V187K with NADPH and substrate | | Descriptor: | (4~{S})-4-oxidanyl-4,6-dihydrofuro[3,2-c]pyran-2-one, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Short-chain dehydrogenase/reductase | | Authors: | Dai, L, Li, H, Hu, Y, Guo, R.T, Chen, C.C. | | Deposit date: | 2022-05-26 | | Release date: | 2022-10-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-based rational design of a short-chain dehydrogenase/reductase for improving activity toward mycotoxin patulin.
Int.J.Biol.Macromol., 222, 2022
|
|
7XWK
 
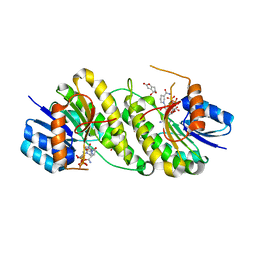 | | structure of patulin-detoxifying enzyme Y155F with NADPH and substrate | | Descriptor: | (4~{S})-4-oxidanyl-4,6-dihydrofuro[3,2-c]pyran-2-one, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Short-chain dehydrogenase/reductase | | Authors: | Dai, L, Li, H, Hu, Y, Guo, R.T, Chen, C.C. | | Deposit date: | 2022-05-26 | | Release date: | 2022-10-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Structure-based rational design of a short-chain dehydrogenase/reductase for improving activity toward mycotoxin patulin.
Int.J.Biol.Macromol., 222, 2022
|
|
1TXX
 
 | |
5U4C
 
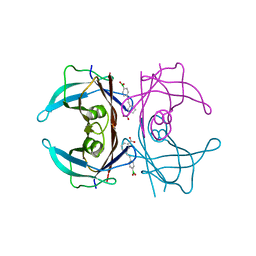 | | Wild-type Transthyretin in complex with 3-[(1E)-2-(4-Boronic acid)ethenyl]benzoic Acid | | Descriptor: | 3-[(E)-2-(4-boronophenyl)ethenyl]benzoic acid, Transthyretin | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
5U4A
 
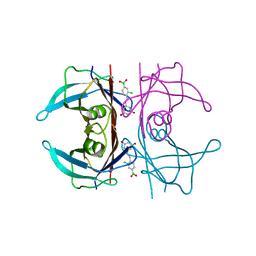 | | Wild-type Transthyretin in complex with 5-[(1E)-2-(2-Chloro-4-boronic acid)ethenyl]-1,3-benzenediol | | Descriptor: | Transthyretin, {3-chloro-4-[(E)-2-(3,5-dihydroxyphenyl)ethenyl]phenyl}boronic acid | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
5U4G
 
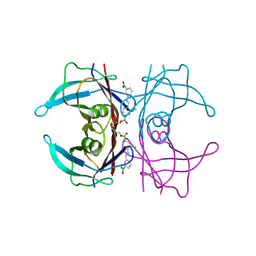 | | Wild-type Transthyretin in complex with 2-Boronic Acid-1-[(1E)-2-(3-boronic acid)ethenyl]-4-chlorobenzene | | Descriptor: | (3-{(E)-2-[2-chloro-4-(hydroxyboranyl)phenyl]ethenyl}phenyl)boronic acid, Transthyretin | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
1V4E
 
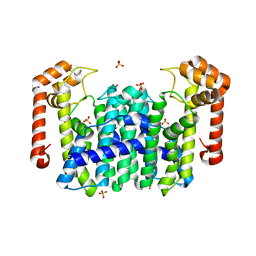 | | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima | | Descriptor: | SULFATE ION, octoprenyl-diphosphate synthase | | Authors: | Guo, R.T, Kuo, C.J, Chou, C.C, Ko, T.P, Shr, H.L, Liang, P.H, Wang, A.H.-J. | | Deposit date: | 2003-11-13 | | Release date: | 2004-03-02 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima and Mechanism of Product Chain Length Determination
J.Biol.Chem., 279, 2004
|
|
1V4I
 
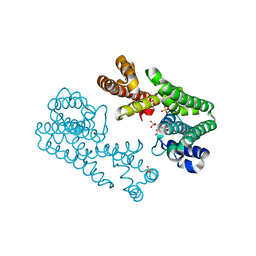 | | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima F132A mutant | | Descriptor: | SULFATE ION, octoprenyl-diphosphate synthase | | Authors: | Guo, R.T, Kuo, C.J, Chou, C.C, Ko, T.P, Shr, H.L, Liang, P.H, Wang, A.H.-J. | | Deposit date: | 2003-11-14 | | Release date: | 2004-03-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima and Mechanism of Product Chain Length Determination
J.Biol.Chem., 279, 2004
|
|
5TXV
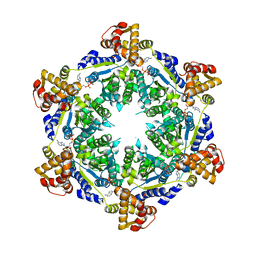 
 | | HslU P21 cell with 4 hexamers | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ATP-dependent protease ATPase subunit HslU | | Authors: | Grant, R.A, Chen, J, Glynn, S.E, Sauer, R.T. | | Deposit date: | 2016-11-17 | | Release date: | 2017-03-01 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (7.086 Å) | | Cite: | Covalently linked HslU hexamers support a probabilistic mechanism that links ATP hydrolysis to protein unfolding and translocation.
J. Biol. Chem., 292, 2017
|
|
5U48
 
 | | Wild-type Transthyretin in complex with 5-[(1E)-2-(4-Boronic acid)ethenyl]-1,3-benzenediol | | Descriptor: | Transthyretin, {4-[(E)-2-(3,5-dihydroxyphenyl)ethenyl]phenyl}boronic acid | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
5U4F
 
 | | Wild-type Transthyretin in complex with 1,1'-(1E)-(1,2-Ethenediyl)bis[2-chloro-4-boronic acid]benzene | | Descriptor: | Transthyretin, [(E)-ethene-1,2-diylbis(3-chloro-4,1-phenylene)]diboronic acid | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
5U4B
 
 | | Wild-type Transthyretin in complex with 3-[(1E)-2-(4-Hydroxyphenyl)ethenyl]benzoic Acid | | Descriptor: | 3-[(E)-2-(4-hydroxyphenyl)ethenyl]benzoic acid, Transthyretin | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.448 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
5TXT
 
 | | Structure of asymmetric apo/holo ALAS dimer from S. cerevisiae | | Descriptor: | 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, 5-aminolevulinate synthase, mitochondrial, ... | | Authors: | Brown, B.L, Grant, R.A, Kardon, J.R, Sauer, R.T, Baker, T.A. | | Deposit date: | 2016-11-17 | | Release date: | 2018-03-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of the Mitochondrial Aminolevulinic Acid Synthase, a Key Heme Biosynthetic Enzyme.
Structure, 26, 2018
|
|
1V4J
 
 | | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima V73Y mutant | | Descriptor: | SULFATE ION, octoprenyl-diphosphate synthase | | Authors: | Guo, R.T, Kuo, C.J, Chou, C.C, Ko, T.P, Shr, H.L, Liang, P.H, Wang, A.H.-J. | | Deposit date: | 2003-11-14 | | Release date: | 2004-03-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima and Mechanism of Product Chain Length Determination
J.Biol.Chem., 279, 2004
|
|
5U49
 
 | | Wild-type Transthyretin in complex with 5-[(1E)-2-(2-Chloro-4-hydroxyphenyl)ethenyl]-1,3-benzenediol | | Descriptor: | 5-[(E)-2-(2-chloro-4-hydroxyphenyl)ethenyl]benzene-1,3-diol, Transthyretin | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
5U4E
 
 | | Wild-type Transthyretin in complex with 3-[(1E)-2-(2-Chloro-4-boronic acid)ethenyl]benzoic Acid | | Descriptor: | 3-[(E)-2-(4-borono-2-chlorophenyl)ethenyl]benzoic acid, Transthyretin | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
5U4D
 
 | | Wild-type Transthyretin in complex with 3-[(1E)-2-(2-Chloro-4-hydroxyphenyl)ethenyl]benzoic Acid | | Descriptor: | 3-[(E)-2-(2-chloro-4-hydroxyphenyl)ethenyl]benzoic acid, Transthyretin | | Authors: | Windsor, I.W, Smith, T.P, Raines, R.T, Forest, K.T. | | Deposit date: | 2016-12-03 | | Release date: | 2017-09-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Stilbene Boronic Acids Form a Covalent Bond with Human Transthyretin and Inhibit Its Aggregation.
J. Med. Chem., 60, 2017
|
|
1V4H
 
 | | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima F52A mutant | | Descriptor: | SULFATE ION, octoprenyl-diphosphate synthase | | Authors: | Guo, R.T, Kuo, C.J, Chou, C.C, Ko, T.P, Shr, H.L, Liang, P.H, Wang, A.H.-J. | | Deposit date: | 2003-11-14 | | Release date: | 2004-03-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima and Mechanism of Product Chain Length Determination
J.Biol.Chem., 279, 2004
|
|
5TXR
 
 | | Structure of ALAS from S. cerevisiae non-covalently bound to PLP cofactor | | Descriptor: | 5-aminolevulinate synthase, mitochondrial, FORMIC ACID, ... | | Authors: | Brown, B.L, Grant, R.A, Kardon, J.R, Sauer, R.T, Baker, T.A. | | Deposit date: | 2016-11-17 | | Release date: | 2018-03-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the Mitochondrial Aminolevulinic Acid Synthase, a Key Heme Biosynthetic Enzyme.
Structure, 26, 2018
|
|
1V4K
 
 | | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima S77F mutant | | Descriptor: | SULFATE ION, octoprenyl-diphosphate synthase | | Authors: | Guo, R.T, Kuo, C.J, Chou, C.C, Ko, T.P, Shr, H.L, Liang, P.H, Wang, A.H.-J. | | Deposit date: | 2003-11-14 | | Release date: | 2004-03-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal Structure of Octaprenyl Pyrophosphate Synthase from Hyperthermophilic Thermotoga maritima and Mechanism of Product Chain Length Determination
J.Biol.Chem., 279, 2004
|
|
5V8C
 
 | | LytR-Csp2A-Psr enzyme from Actinomyces oris | | Descriptor: | 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, COBALT (II) ION, PHOSPHATE ION, ... | | Authors: | Amer, B.R, Sawaya, M.R, Liauw, B, Fu, J.Y, Ton-That, H, Clubb, R.T. | | Deposit date: | 2017-03-21 | | Release date: | 2018-04-11 | | Last modified: | 2020-01-08 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Structure and Mechanism of LcpA, a Phosphotransferase That Mediates Glycosylation of a Gram-Positive Bacterial Cell Wall-Anchored Protein.
MBio, 10, 2019
|
|
5V1Y
 
 | | Crystal structure of the ternary RPN13 PRU-RPN2 (940-953)-ubiquitin complex | | Descriptor: | 26S proteasome non-ATPase regulatory subunit 1, Proteasomal ubiquitin receptor ADRM1, Ubiquitin | | Authors: | Hemmis, C.W, VanderLinden, R.T, Yao, T, Robinson, H, Hill, C.P. | | Deposit date: | 2017-03-02 | | Release date: | 2017-05-03 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.421 Å) | | Cite: | Structure and energetics of pairwise interactions between proteasome subunits RPN2, RPN13, and ubiquitin clarify a substrate recruitment mechanism.
J. Biol. Chem., 292, 2017
|
|
4ERL
 
 | |
2KW8
 
 | |
4E7A
 
 | |
