7YEA
 
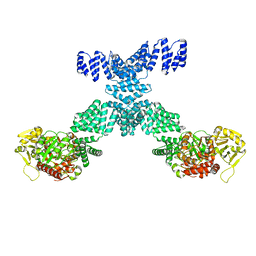 | | Human O-GlcNAc transferase Dimer | | 分子名称: | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase 110 kDa subunit | | 著者 | Gao, H, Lu, P, Liu, Y. | | 登録日 | 2022-07-05 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-01-24 | | 実験手法 | ELECTRON MICROSCOPY (3.82 Å) | | 主引用文献 | Cryo-EM structure of human O-GlcNAcylation enzyme pair OGT-OGA complex.
Nat Commun, 14, 2023
|
|
3P83
 
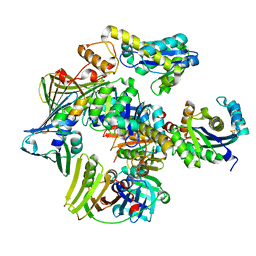 | | Structure of the PCNA:RNase HII complex from Archaeoglobus fulgidus. | | 分子名称: | DNA polymerase sliding clamp, Ribonuclease HII | | 著者 | Bubeck, D, Reijns, M.A, Graham, S.C, Astell, K.R, Jones, E.Y, Jackson, A.P. | | 登録日 | 2010-10-13 | | 公開日 | 2011-02-02 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (3.05 Å) | | 主引用文献 | PCNA directs type 2 RNase H activity on DNA replication and repair substrates.
Nucleic Acids Res., 39, 2011
|
|
7Y77
 
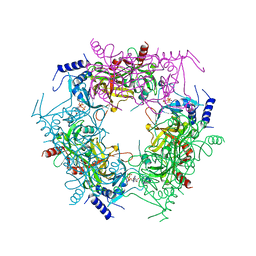 | | Crystal structure of rice NAL1 | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, Protein NARROW LEAF 1 | | 著者 | Yan, J.J, Guan, Z.Y, Yin, P, Xiong, L.Z. | | 登録日 | 2022-06-21 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Serine protease NAL1 exerts pleiotropic functions through degradation of TOPLESS-related corepressor in rice.
Nat.Plants, 9, 2023
|
|
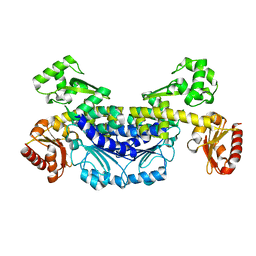
7YIK
 
 | |
6SZY
 
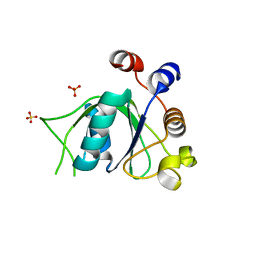 | | Crystal structure of YTHDC1 with fragment 12 (DHU_DC1_150) | | 分子名称: | 4-propan-2-yl-5,6,7,8-tetrahydro-4~{a}~{H}-quinazolin-2-one, SULFATE ION, YTHDC1 | | 著者 | Bedi, R.K, Huang, D, Sledz, P, Caflisch, A. | | 登録日 | 2019-10-02 | | 公開日 | 2020-03-04 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (1.79 Å) | | 主引用文献 | Selectively Disrupting m6A-Dependent Protein-RNA Interactions with Fragments.
Acs Chem.Biol., 15, 2020
|
|
7YIL
 
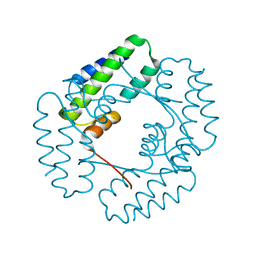 | |
6T04
 
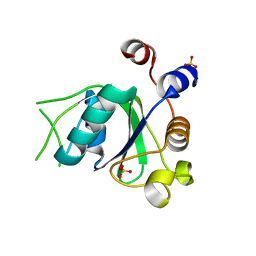 | | Crystal structure of YTHDC1 with fragment 17 (DHU_DC1_042) | | 分子名称: | 2-phenyl-4,5-dihydro-1~{H}-imidazole, SULFATE ION, YTH domain-containing protein 1 | | 著者 | Bedi, R.K, Huang, D, Sledz, P, Caflisch, A. | | 登録日 | 2019-10-02 | | 公開日 | 2020-03-04 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Selectively Disrupting m6A-Dependent Protein-RNA Interactions with Fragments.
Acs Chem.Biol., 15, 2020
|
|
2XTH
 
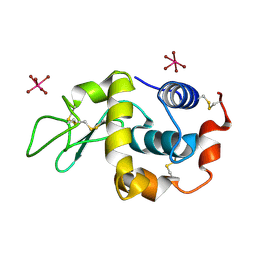 | | K2PtBr6 binding to lysozyme | | 分子名称: | HEXABROMOPLATINATE(IV), LYSOZYME C | | 著者 | Helliwell, J.R, Bell, A.M.T, Bryant, P, Fisher, S, Habash, J, Helliwell, M, Margiolaki, I, Kaenket, S, Watier, Y, Wright, J, Yalamanchili, S.K. | | 登録日 | 2010-10-07 | | 公開日 | 2010-12-08 | | 最終更新日 | 2017-06-28 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Time-Dependent Analysis of K2Ptbr6 Binding to Lysozyme Studied by Protein Powder and Single Crystal X-Ray Analysis
Z.Kristallogr., 225, 2010
|
|
6T12
 
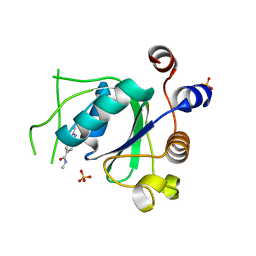 | | Crystal structure of YTHDC1 with fragment 30 (DHU_DC1_220) | | 分子名称: | SULFATE ION, YTHDC1, ~{N},2,3-trimethyl-1~{H}-indole-5-carboxamide | | 著者 | Bedi, R.K, Huang, D, Sledz, P, Caflisch, A. | | 登録日 | 2019-10-03 | | 公開日 | 2020-03-04 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (1.46 Å) | | 主引用文献 | Selectively Disrupting m6A-Dependent Protein-RNA Interactions with Fragments.
Acs Chem.Biol., 15, 2020
|
|
3PD4
 
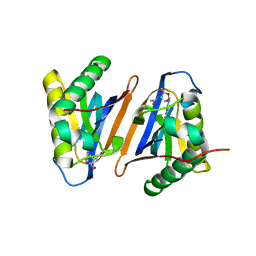 | |
8SZR
 
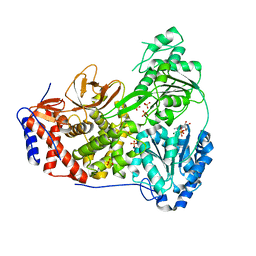 | | Dog DHX9 bound to ADP | | 分子名称: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, RNA helicase, ... | | 著者 | Lee, Y.-T, Sickmier, E.A, Grigoriu, S, Boriack-Sjodin, P.A. | | 登録日 | 2023-05-30 | | 公開日 | 2023-08-02 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.97 Å) | | 主引用文献 | Crystal structures of the DExH-box RNA helicase DHX9.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
2XW9
 
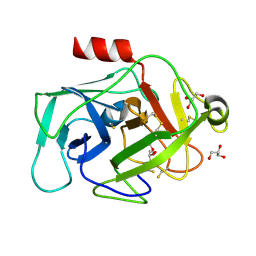 | | Crystal Structure of Complement Factor D mutant S183A | | 分子名称: | COMPLEMENT FACTOR D, GLYCEROL | | 著者 | Forneris, F, Ricklin, D, Wu, J, Tzekou, A, Wallace, R.S, Lambris, J.D, Gros, P. | | 登録日 | 2010-11-01 | | 公開日 | 2011-01-12 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.2 Å) | | 主引用文献 | Structures of C3B in Complex with Factors B and D Give Insight Into Complement Convertase Formation.
Science, 330, 2010
|
|
8IFY
 
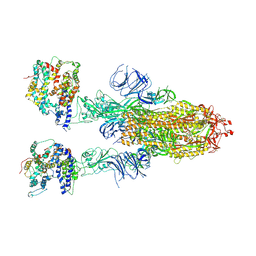 | | Cryo-EM structure of SARS-CoV-2 Omicron BA.4/5 spike protein in complex with white-tailed deer ACE2 | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme, ... | | 著者 | Han, P, Meng, Y.M, Qi, J.X. | | 登録日 | 2023-02-20 | | 公開日 | 2023-08-30 | | 最終更新日 | 2023-10-25 | | 実験手法 | ELECTRON MICROSCOPY (2.55 Å) | | 主引用文献 | Structural basis of white-tailed deer, Odocoileus virginianus , ACE2 recognizing all the SARS-CoV-2 variants of concern with high affinity.
J.Virol., 97, 2023
|
|
3PD3
 
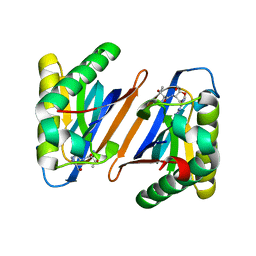 | |
8SHH
 
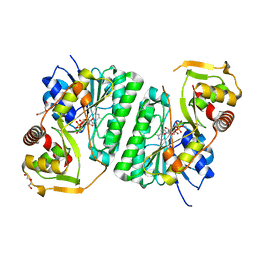 | | Crystal structure of EvdS6 decarboxylase in ligand free state | | 分子名称: | DI(HYDROXYETHYL)ETHER, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, dTDP-glucose 4,6-dehydratase | | 著者 | Sharma, P, Frigo, L, Dulin, C.C, Bachmann, B.O, Iverson, T.M. | | 登録日 | 2023-04-14 | | 公開日 | 2023-08-09 | | 最終更新日 | 2024-05-22 | | 実験手法 | X-RAY DIFFRACTION (1.93 Å) | | 主引用文献 | EvdS6 is a bifunctional decarboxylase from the everninomicin gene cluster.
J.Biol.Chem., 299, 2023
|
|
5LM4
 
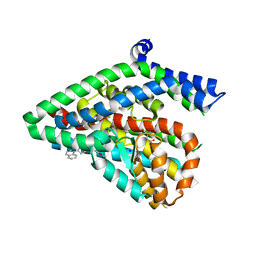 | | Structure of the thermostalilized EAAT1 cryst-II mutant in complex with L-ASP and the allosteric inhibitor UCPH101 | | 分子名称: | 2-Amino-5,6,7,8-tetrahydro-4-(4-methoxyphenyl)-7-(naphthalen-1-yl)-5-oxo-4H-chromene-3-carbonitrile, ASPARTIC ACID, Excitatory amino acid transporter 1,Neutral amino acid transporter B(0),Excitatory amino acid transporter 1, ... | | 著者 | Canul-Tec, J, Assal, R, Legrand, P, Reyes, N. | | 登録日 | 2016-07-29 | | 公開日 | 2017-04-19 | | 最終更新日 | 2024-01-10 | | 実験手法 | X-RAY DIFFRACTION (3.1 Å) | | 主引用文献 | Structure and allosteric inhibition of excitatory amino acid transporter 1.
Nature, 544, 2017
|
|
6Y71
 
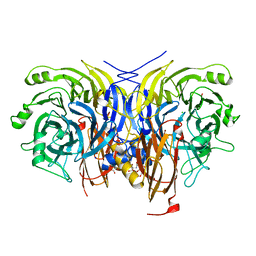 | | Pseudomonas stutzeri nitrous oxide reductase mutant, H130A | | 分子名称: | 2-[3-(2-HYDROXY-1,1-DIHYDROXYMETHYL-ETHYLAMINO)-PROPYLAMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, CHLORIDE ION, ... | | 著者 | Zhang, L, Kroneck, P.M.H, Einsle, O. | | 登録日 | 2020-02-27 | | 公開日 | 2021-01-27 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (1.64 Å) | | 主引用文献 | A [3Cu:2S] cluster provides insight into the assembly and function of the Cu Z site of nitrous oxide reductase.
Chem Sci, 12, 2021
|
|
2XYW
 
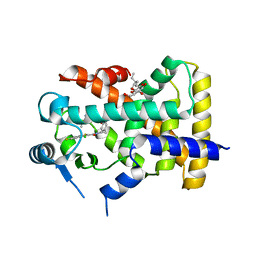 | |
8T5X
 
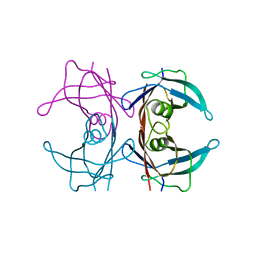 | | Probing the dissociation pathway of a kinetically labile transthyretin mutant (A25T) | | 分子名称: | Transthyretin | | 著者 | Ferguson, J.A, Sun, X, Leach, B.I, Stanfield, R.L, Dyson, H.J, Wright, P.E. | | 登録日 | 2023-06-14 | | 公開日 | 2023-08-02 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (1.63 Å) | | 主引用文献 | Probing the Dissociation Pathway of a Kinetically Labile Transthyretin Mutant.
J.Am.Chem.Soc., 146, 2024
|
|
8SK0
 
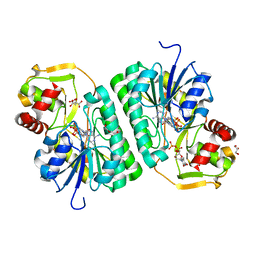 | | Crystal structure of EvdS6 decarboxylase in ligand bound state | | 分子名称: | CITRATE ANION, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | 著者 | Sharma, P, Frigo, L, Dulin, C.C, Bachmann, B.O, Iverson, T.M. | | 登録日 | 2023-04-18 | | 公開日 | 2023-08-09 | | 最終更新日 | 2024-05-22 | | 実験手法 | X-RAY DIFFRACTION (1.51 Å) | | 主引用文献 | EvdS6 is a bifunctional decarboxylase from the everninomicin gene cluster.
J.Biol.Chem., 299, 2023
|
|
7YII
 
 | |
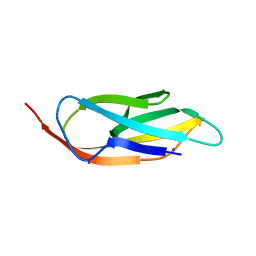
3PE9
 | |
5WLQ
 
 | | Crystal Structure of Amino Acids 1677-1755 of Human Beta Cardiac Myosin Fused to Gp7 and Eb1 | | 分子名称: | Capsid assembly scaffolding protein,Myosin-7,Microtubule-associated protein RP/EB family member 1, SULFATE ION, trimethylamine oxide | | 著者 | Andreas, M.P, Ajay, G, Gellings, J, Rayment, I. | | 登録日 | 2017-07-27 | | 公開日 | 2017-08-09 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (3.104 Å) | | 主引用文献 | Design considerations in coiled-coil fusion constructs for the structural determination of a problematic region of the human cardiac myosin rod.
J. Struct. Biol., 200, 2017
|
|
5WME
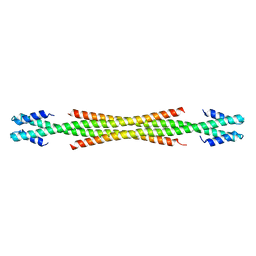 
 | | Crystal Structure of Amino Acids 1729-1786 of Human Beta Cardiac Myosin Fused to Gp7 as Anti-Parallel Four-Helix Bundle | | 分子名称: | Capsid assembly scaffolding protein,Myosin-7 | | 著者 | Andreas, M.P, Ajay, G, Gellings, J, Rayment, I. | | 登録日 | 2017-07-28 | | 公開日 | 2017-08-23 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Design considerations in coiled-coil fusion constructs for the structural determination of a problematic region of the human cardiac myosin rod.
J. Struct. Biol., 200, 2017
|
|
2XQZ
 
 | | Neutron structure of the perdeuterated Toho-1 R274N R276N double mutant beta-lactamase | | 分子名称: | BETA-LACTAMSE TOHO-1 | | 著者 | Tomanicek, S.J, Wang, K.K, Weiss, K.L, Blakeley, M.P, Cooper, J, Chen, Y, Coates, L. | | 登録日 | 2010-09-08 | | 公開日 | 2010-12-22 | | 最終更新日 | 2024-05-08 | | 実験手法 | NEUTRON DIFFRACTION (2.1 Å) | | 主引用文献 | The Active Site Protonation States of Perdeuterated Toho-1 Beta-Lactamase Determined by Neutron Diffraction Support a Role for Glu166 as the General Base in Acylation.
FEBS Lett., 585, 2011
|
|
