3M4G
 
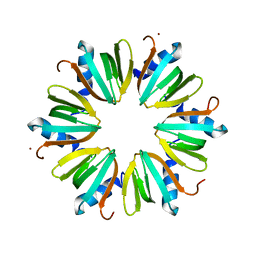 | | H57A HFQ from Pseudomonas Aeruginosa | | Descriptor: | Protein hfq, ZINC ION | | Authors: | Moskaleva, O, Melnik, B, Gabdulkhakov, A, Garber, M, Nikonov, S, Stolboushkina, E, Nikulin, A. | | Deposit date: | 2010-03-11 | | Release date: | 2010-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The structures of mutant forms of Hfq from Pseudomonas aeruginosa reveal the importance of the conserved His57 for the protein hexamer organization.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
3QSY
 
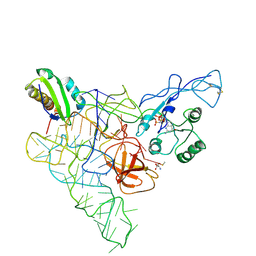 | | Recognition of the methionylated initiator tRNA by the translation initiation factor 2 in Archaea | | Descriptor: | METHIONINE, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, Translation initiation factor 2 subunit alpha, ... | | Authors: | Nikonov, O.S, Stolboushkina, E.A, Zelinskaya, N.V, Nikulin, A.D, Garber, M.B, Nikonov, S.V. | | Deposit date: | 2011-02-22 | | Release date: | 2012-03-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal structure of the archaeal translation initiation factor 2 in complex with a GTP analogue and Met-tRNAf(Met.)
J.Mol.Biol., 425, 2013
|
|
1AD2
 
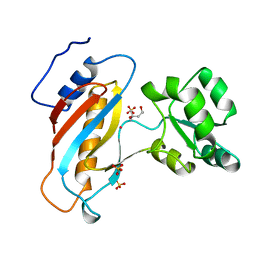 | | RIBOSOMAL PROTEIN L1 MUTANT WITH SERINE 179 REPLACED BY CYSTEINE | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, MERCURY (II) ION, RIBOSOMAL PROTEIN L1, ... | | Authors: | Unge, J, Al-Karadaghi, S, Liljas, A, Jonsson, B.-H, Eliseikina, I, Ossina, N, Nevskaya, N, Fomenkova, N, Garber, M, Nikonov, S. | | Deposit date: | 1997-02-20 | | Release date: | 1997-05-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A mutant form of the ribosomal protein L1 reveals conformational flexibility.
FEBS Lett., 411, 1997
|
|
1BXY
 
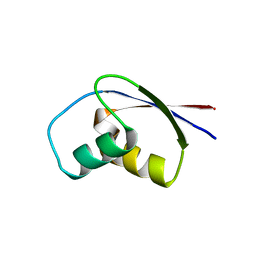 | | CRYSTAL STRUCTURE OF RIBOSOMAL PROTEIN L30 FROM THERMUS THERMOPHILUS AT 1.9 A RESOLUTION: CONFORMATIONAL FLEXIBILITY OF THE MOLECULE. | | Descriptor: | PROTEIN (RIBOSOMAL PROTEIN L30) | | Authors: | Fedorov, R, Nevskaya, N, Khairullina, A, Tishchenko, S, Mikhailov, A, Garber, M, Nikonov, S. | | Deposit date: | 1998-10-09 | | Release date: | 1998-10-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of ribosomal protein L30 from Thermus thermophilus at 1.9 A resolution: conformational flexibility of the molecule.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
1BXE
 
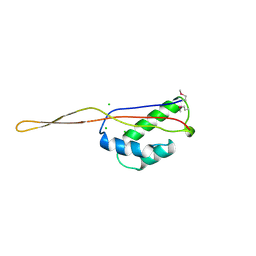 | | RIBOSOMAL PROTEIN L22 FROM THERMUS THERMOPHILUS | | Descriptor: | CHLORIDE ION, PROTEIN (RIBOSOMAL PROTEIN L22) | | Authors: | Unge, J, Aberg, A, Al-Karadaghi, S, Nikulin, A, Nikonov, S, Davydova, N, Nevskaya, N, Garber, M, Liljas, A. | | Deposit date: | 1998-10-02 | | Release date: | 1998-10-07 | | Last modified: | 2018-03-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The crystal structure of ribosomal protein L22 from Thermus thermophilus: insights into the mechanism of erythromycin resistance.
Structure, 6, 1998
|
|
1YGM
 
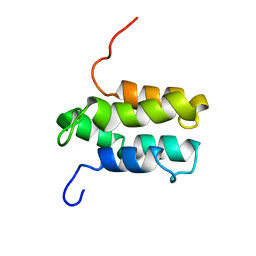 | | NMR structure of Mistic | | Descriptor: | hypothetical protein BSU31320 | | Authors: | Roosild, T.P, Greenwald, J, Vega, M, Castronovo, S, Riek, R, Choe, S. | | Deposit date: | 2005-01-05 | | Release date: | 2005-03-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR structure of Mistic, a membrane-integrating protein for membrane protein expression.
Science, 307, 2005
|
|
1Y4I
 
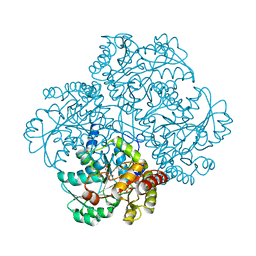 | | Crystal structure of Citrobacter Freundii L-methionine-lyase | | Descriptor: | SULFATE ION, methionine gamma-lyase | | Authors: | Revtovich, S.V, Mamaeva, D.V, Morozova, E.A, Nikulin, A.D, Nikonov, S.V, Garber, M.B, Demidkina, T.V. | | Deposit date: | 2004-12-01 | | Release date: | 2005-06-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of Citrobacter freundii L-methionine gamma-lyase.
Acta Crystallogr.,Sect.F, 61, 2005
|
|
3JSY
 
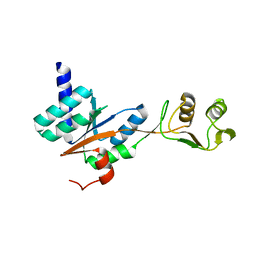 | | N-terminal fragment of ribosomal protein L10 from Methanococcus jannaschii | | Descriptor: | Acidic ribosomal protein P0 homolog | | Authors: | Kravchenko, O, Mitroshin, I, Nikulin, A.D, Piendl, W, Garber, M, Nikonov, S. | | Deposit date: | 2009-09-11 | | Release date: | 2010-05-26 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of a two-domain N-terminal fragment of ribosomal protein L10 from Methanococcus jannaschii reveals a specific piece of the archaeal ribosomal stalk
J.Mol.Biol., 399, 2010
|
|
1AK1
 
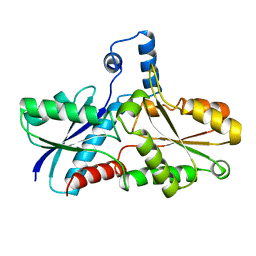 | | FERROCHELATASE FROM BACILLUS SUBTILIS | | Descriptor: | FERROCHELATASE | | Authors: | Al-Karadaghi, S, Hansson, M, Nikonov, S, Jonsson, B, Hederstedt, L. | | Deposit date: | 1997-05-28 | | Release date: | 1997-12-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of ferrochelatase: the terminal enzyme in heme biosynthesis.
Structure, 5, 1997
|
|
2RFV
 
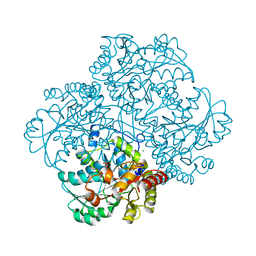 | | High resolution structure of L-methionine gamma-lyase from Citrobacter freundii | | Descriptor: | CHLORIDE ION, Methionine gamma-lyase | | Authors: | Nikulin, A.D, Revtovich, S.V, Morozova, E.A, Nevskaya, N.A, Nikonov, S.V, Garber, M.B, Demidkina, T.V. | | Deposit date: | 2007-10-02 | | Release date: | 2008-08-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.355 Å) | | Cite: | High-resolution structure of methionine gamma-lyase from Citrobacter freundii.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
1DWU
 
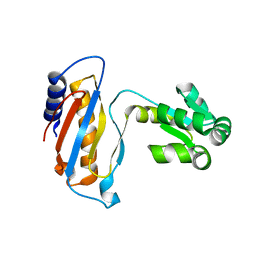 | | Ribosomal protein L1 | | Descriptor: | RIBOSOMAL PROTEIN L1 | | Authors: | Tishchenko, S.V, Nevskaya, N.A, Pavelyev, M.N, Nikonov, S.V, Garber, M.B, Piendl, W. | | Deposit date: | 1999-12-13 | | Release date: | 2000-12-07 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of Ribosomal Protein L1 from Methanococcus Thermolithotrophicus. Functionally Important Structural Invariants on the L1 Surface
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1KUQ
 
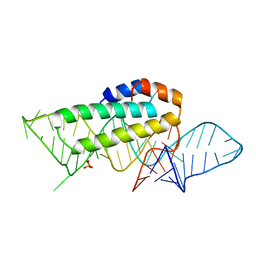 | | CRYSTAL STRUCTURE OF T3C MUTANT S15 RIBOSOMAL PROTEIN IN COMPLEX WITH 16S RRNA | | Descriptor: | 16S RIBOSOMAL RNA FRAGMENT, 30S RIBOSOMAL PROTEIN S15, SULFATE ION | | Authors: | Nikulin, A.D, Tishchenko, S, Revtovich, S, Ehresmann, B, Ehresmann, C, Dumas, P, Garber, M, Nikonov, S, Nevskaya, N. | | Deposit date: | 2002-01-22 | | Release date: | 2003-06-24 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Role of N-terminal helix in interaction of ribosomal protein S15 with 16S rRNA.
Biochemistry Mosc., 69, 2004
|
|
3A17
 
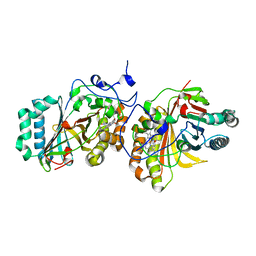 | | Crystal Structure of Aldoxime Dehydratase (OxdRE) in Complex with Butyraldoxime (Co-crystal) | | Descriptor: | (1Z)-butanal oxime, Aldoxime dehydratase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Sawai, H, Sugimoto, H, Kato, Y, Asano, Y, Shiro, Y, Aono, S. | | Deposit date: | 2009-03-26 | | Release date: | 2009-09-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-ray crystal structure of michaelis complex of aldoxime dehydratase
J.Biol.Chem., 284, 2009
|
|
3U56
 
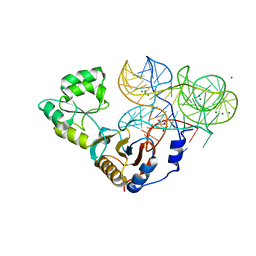 | |
2OUM
 
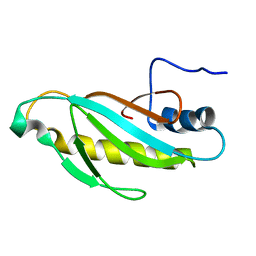 | | The first domain of L1 from Thermus thermophilus | | Descriptor: | 50S ribosomal protein L1 | | Authors: | Kljashtorny, V, Tishchenko, S, Nevskaya, N, Nikonov, S, Davydova, N, Garber, M. | | Deposit date: | 2007-02-12 | | Release date: | 2008-02-26 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Domain I of ribosomal protein L1 is sufficient for specific RNA binding.
Nucleic Acids Res., 35, 2007
|
|
2OV7
 
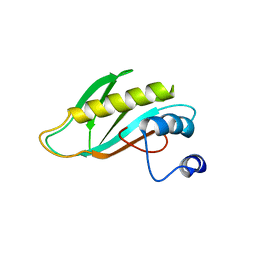 | | The first domain of the ribosomal protein L1 from Thermus thermophilus | | Descriptor: | 50S ribosomal protein L1 | | Authors: | Kljashtorny, V, Tishchenko, S, Nevskaya, N, Nikonov, S, Davydova, N, Garber, M. | | Deposit date: | 2007-02-13 | | Release date: | 2007-12-25 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Domain I of ribosomal protein L1 is sufficient for specific RNA binding.
Nucleic Acids Res., 35, 2007
|
|
2AIY
 
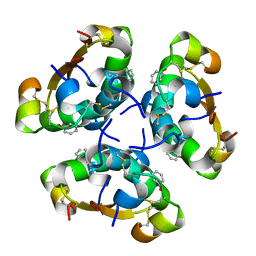 | | R6 HUMAN INSULIN HEXAMER (SYMMETRIC), NMR, 20 STRUCTURES | | Descriptor: | PHENOL, PROTEIN (INSULIN) | | Authors: | O'Donoghue, S.I, Chang, X, Abseher, R, Nilges, M, Led, J.J. | | Deposit date: | 1998-12-28 | | Release date: | 2000-02-28 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Unraveling the symmetry ambiguity in a hexamer: calculation of the R6 human insulin structure.
J.Biomol.NMR, 16, 2000
|
|
3UMY
 
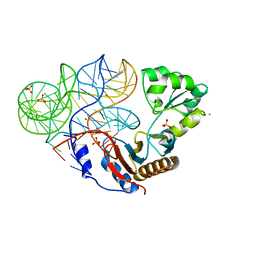 | |
2SIC
 
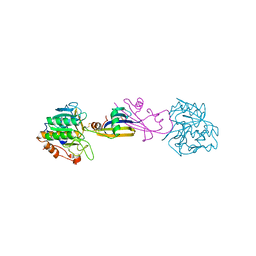 | | REFINED CRYSTAL STRUCTURE OF THE COMPLEX OF SUBTILISIN BPN' AND STREPTOMYCES SUBTILISIN INHIBITOR AT 1.8 ANGSTROMS RESOLUTION | | Descriptor: | CALCIUM ION, STREPTOMYCES SUBTILISIN INHIBITOR (SSI), SUBTILISIN BPN' | | Authors: | Mitsui, Y, Takeuchi, Y, Hirono, S, Akagawa, H, Nakamura, K.T. | | Deposit date: | 1991-04-01 | | Release date: | 1993-04-15 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Refined crystal structure of the complex of subtilisin BPN' and Streptomyces subtilisin inhibitor at 1.8 A resolution.
J.Mol.Biol., 221, 1991
|
|
1ONK
 
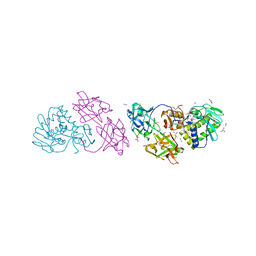 | | Mistletoe lectin I from viscum album | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, AZIDE ION, Beta-galactoside specific lectin I A chain, ... | | Authors: | Gabdoulkhakov, A.G, Savoshkina, Y, Krauspenhaar, R, Stoeva, S, Konareva, N, Kornilov, V, Kornev, A.N, Voelter, W, Nikonov, S.V, Betzel, C, Mikhailov, A.M. | | Deposit date: | 2003-02-28 | | Release date: | 2004-02-28 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Mistletoe lectin I from viscum album
To be Published
|
|
2BH8
 
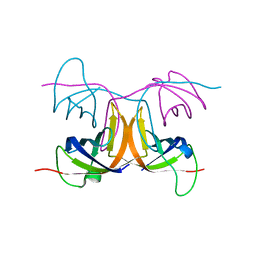 | | Combinatorial Protein 1b11 | | Descriptor: | 1B11 | | Authors: | De Bono, S, Riechmann, L, Girard, E, Williams, R.L, Winter, G. | | Deposit date: | 2005-01-07 | | Release date: | 2005-02-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A Segment of Cold Shock Protein Directs the Folding of a Combinatorial Protein
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1ZCF
 
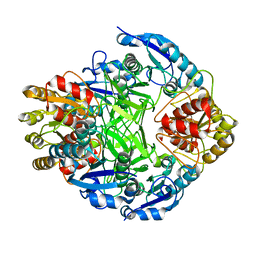 | |
3WD4
 
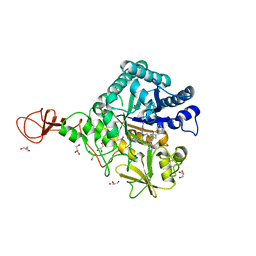 | | Serratia marcescens Chitinase B complexed with azide inhibitor and quinoline compound | | Descriptor: | (E)-N-(prop-2-en-1-yloxy)-1-(quinolin-4-yl)methanimine, Chitinase B, GLYCEROL, ... | | Authors: | Hirose, T, Maita, N, Gouda, H, Koseki, J, Yamamoto, T, Sugawara, A, Nakano, H, Hirono, S, Shiomi, K, Watanabe, T, Taniguchi, H, Sharpless, K.B, Omura, S, Sunazuka, T. | | Deposit date: | 2013-06-06 | | Release date: | 2013-09-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Observation of the controlled assembly of preclick components in the in situ click chemistry generation of a chitinase inhibitor
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
2ORA
 
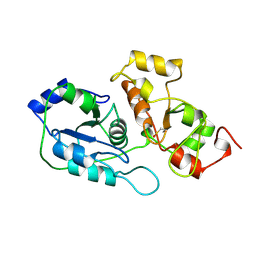 | | RHODANESE (THIOSULFATE: CYANIDE SULFURTRANSFERASE) | | Descriptor: | OXIDIZED RHODANESE | | Authors: | Gliubich, F, Gazerro, M, Zanotti, G, Delbono, S, Berni, R. | | Deposit date: | 1996-02-22 | | Release date: | 1996-08-01 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Active site structural features for chemically modified forms of rhodanese.
J.Biol.Chem., 271, 1996
|
|
3U4M
 
 | | Crystal structure of ribosomal protein tthl1 in complex with 80nt 23s rna from thermus thermophilus | | Descriptor: | 50S ribosomal protein L1, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Gabdulkhakov, A.G, Nevskaya, N.A, NIkonov, S.V. | | Deposit date: | 2011-10-10 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | High-resolution crystal structure of the isolated ribosomal L1 stalk.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
