7Q4K
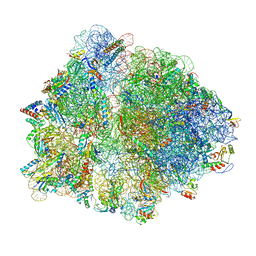 
 | | Erythromycin-stalled Escherichia coli 70S ribosome with streptococcal MsrDL nascent chain | | Descriptor: | 16S rRNA, 23S rRNA, 30S ribosomal protein S10, ... | | Authors: | Fostier, C.R, Ousalem, F, Soufari, H, Leroy, E.C, Ngo, S, Innis, A, Hashem, Y, Boel, G. | | Deposit date: | 2021-10-31 | | Release date: | 2022-11-16 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Regulation of the macrolide resistance ABC-F translation factor MsrD.
Nat Commun, 14, 2023
|
|
7RI3
 
 | | Crystal structure of Albireti Toxin, a diphtheria toxin homolog, from Streptomyces albireticuli | | Descriptor: | ACETATE ION, CALCIUM ION, Diphtheria_T domain-containing protein, ... | | Authors: | Sugiman-Marangos, S.N, Gill, S.K, Melnyk, R.A. | | Deposit date: | 2021-07-19 | | Release date: | 2022-04-20 | | Last modified: | 2022-11-02 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Structures of distant diphtheria toxin homologs reveal functional determinants of an evolutionarily conserved toxin scaffold.
Commun Biol, 5, 2022
|
|
7RB4
 
 | | Crystal structure of Peptono Toxin, a diphtheria toxin homolog, from Seinonella peptonophila | | Descriptor: | Diphtheria toxin, C domain, IODIDE ION | | Authors: | Sugiman-Marangos, S.N, Gill, S.K, Melnyk, R.A. | | Deposit date: | 2021-07-05 | | Release date: | 2022-04-20 | | Last modified: | 2022-11-02 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Structures of distant diphtheria toxin homologs reveal functional determinants of an evolutionarily conserved toxin scaffold.
Commun Biol, 5, 2022
|
|
3L1V
 
 | | Crystal structure of GmhB from E. coli in complex with calcium and phosphate. | | Descriptor: | CALCIUM ION, D,D-heptose 1,7-bisphosphate phosphatase, PHOSPHATE ION, ... | | Authors: | Sugiman-Marangos, S.N, Junop, M.S. | | Deposit date: | 2009-12-14 | | Release date: | 2010-01-05 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.954 Å) | | Cite: | Structural and kinetic characterization of the LPS biosynthetic enzyme D-alpha,beta-D-heptose-1,7-bisphosphate phosphatase (GmhB) from Escherichia coli.
Biochemistry, 49, 2010
|
|
3L1U
 
 | | Crystal structure of Calcium-bound GmhB from E. coli. | | Descriptor: | CALCIUM ION, D,D-heptose 1,7-bisphosphate phosphatase, ZINC ION | | Authors: | Sugiman-Marangos, S.N, Junop, M.S. | | Deposit date: | 2009-12-14 | | Release date: | 2010-01-05 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural and kinetic characterization of the LPS biosynthetic enzyme D-alpha,beta-D-heptose-1,7-bisphosphate phosphatase (GmhB) from Escherichia coli.
Biochemistry, 49, 2010
|
|
5TCO
 
 | | Human p38 MAP Kinase in Complex with Dibenzosuberone Compound 1 | | Descriptor: | 3-[(3-benzamido-4-fluoranyl-phenyl)amino]-~{N}-(2-morpholin-4-ylethyl)-11-oxidanylidene-5,6-dihydrodibenzo[1,2-~{d}:1',2'-~{f}][7]annulene-9-carboxamide, Mitogen-activated protein kinase 14, octyl beta-D-glucopyranoside | | Authors: | Mayer-Wrangowski, S.C, Rauh, D. | | Deposit date: | 2016-09-15 | | Release date: | 2017-04-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Optimized Target Residence Time: Type I1/2 Inhibitors for p38 alpha MAP Kinase with Improved Binding Kinetics through Direct Interaction with the R-Spine.
Angew. Chem. Int. Ed. Engl., 56, 2017
|
|
5D10
 
 | | Kinase domain of cSrc in complex with RL236 | | Descriptor: | N-[4-({4-(4-methylpiperazin-1-yl)-6-[(5-methyl-1H-pyrazol-3-yl)amino]pyrimidin-2-yl}oxy)phenyl]prop-2-enamide, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Becker, C, Mayer-Wrangowski, S.C, Julian, E, Rauh, D. | | Deposit date: | 2015-08-03 | | Release date: | 2015-09-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Targeting Drug Resistance in EGFR with Covalent Inhibitors: A Structure-Based Design Approach.
J.Med.Chem., 58, 2015
|
|
3E35
 
 | | Actinobacteria-specific protein of unknown function, SCO1997 | | Descriptor: | MAGNESIUM ION, Uncharacterized protein SCO1997 | | Authors: | Gao, B, Gupta, R.S, Sugiman-Marangos, S, Junop, M.S. | | Deposit date: | 2008-08-06 | | Release date: | 2009-06-23 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and phylogenetic analysis of a conserved actinobacteria-specific protein (ASP1; SCO1997) from Streptomyces coelicolor.
Bmc Struct.Biol., 9, 2009
|
|
7P3I
 
 | | Crystal structure of human CD40/TNFRSF5 in complex with the anti-CD40 DARPin protein | | Descriptor: | Darpin, SODIUM ION, Tumor necrosis factor receptor superfamily member 5 | | Authors: | Malvezzi, F, Mangold, S, Hospodarsch, T, Reichen, C, Iss, C, Lammens, A, Krapp, S, Domke, C. | | Deposit date: | 2021-07-07 | | Release date: | 2022-04-06 | | Last modified: | 2022-05-11 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | A Multispecific Anti-CD40 DARPin Construct Induces Tumor-Selective CD40 Activation and Tumor Regression.
Cancer Immunol Res, 10, 2022
|
|
3UVQ
 
 | | Human p38 MAP Kinase in Complex with a Dibenzosuberone Derivative | | Descriptor: | Mitogen-activated protein kinase 14, N-{5-[(7-{[(2R)-2,3-dihydroxypropyl]oxy}-5-oxo-10,11-dihydro-5H-dibenzo[a,d][7]annulen-2-yl)amino]-2-fluorophenyl}benzamide, octyl beta-D-glucopyranoside | | Authors: | Mayer-Wrangowski, S.C, Richters, A, Gruetter, C, Rauh, D. | | Deposit date: | 2011-11-30 | | Release date: | 2012-12-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Dibenzosuberones as p38 mitogen-activated protein kinase inhibitors with low ATP competitiveness and outstanding whole blood activity.
J.Med.Chem., 56, 2013
|
|
3ESQ
 
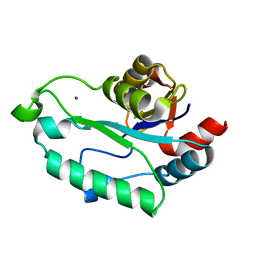 | |
3ESR
 
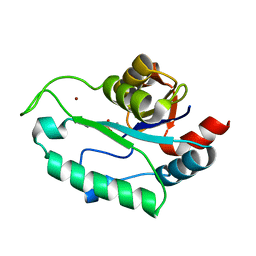 | | Crystal Structure of D,D-heptose1.7-bisphosphate phosphatase from E. coli in complex with calcium and phosphate | | Descriptor: | CALCIUM ION, D,D-heptose 1,7-bisphosphate phosphatase, PHOSPHATE ION, ... | | Authors: | Sugiman-Marangos, S.N, Junop, M.S. | | Deposit date: | 2008-10-06 | | Release date: | 2008-10-14 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of D,D-heptose 1.7-bisphosphate phosphatase from E. Coli.
To be Published
|
|
3QUE
 
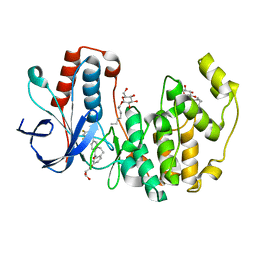 | | Human p38 MAP Kinase in Complex with Skepinone-L | | Descriptor: | 2-[(2,4-difluorophenyl)amino]-7-{[(2R)-2,3-dihydroxypropyl]oxy}-10,11-dihydro-5H-dibenzo[a,d][7]annulen-5-one, Mitogen-activated protein kinase 14, octyl beta-D-glucopyranoside | | Authors: | Gruetter, C, Mayer-Wrangowski, S, Richters, A, Rauh, D. | | Deposit date: | 2011-02-23 | | Release date: | 2012-01-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Skepinone-L is a selective p38 mitogen-activated protein kinase inhibitor.
Nat.Chem.Biol., 8, 2012
|
|
4IQZ
 
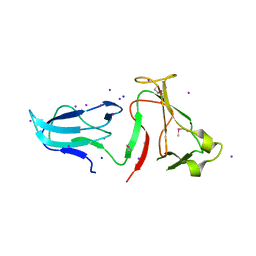 | | The crystal structure of a large insert in RNA polymerase (RpoC) subunit from E. coli | | Descriptor: | DNA-directed RNA polymerase subunit beta', IODIDE ION, SODIUM ION | | Authors: | Bhandari, V, Sugiman-Marangos, S.N, Naushad, H.S, Gupta, R.S, Junop, M.S. | | Deposit date: | 2013-01-14 | | Release date: | 2013-02-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The crystal structure of a large insert in RNA polymerase (RpoC) subunit from E. coli
To be Published
|
|
4IPV
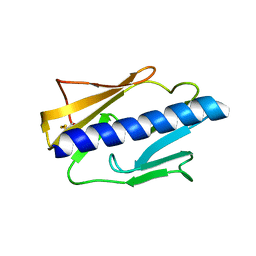 
 | |
4EXW
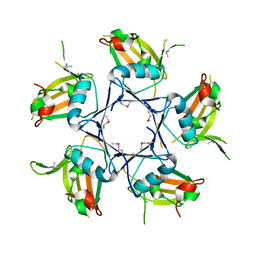 
 | |
3Q9D
 
 | |
3UVP
 
 | | Human p38 MAP Kinase in Complex with a Benzamide Substituted Benzosuberone | | Descriptor: | Mitogen-activated protein kinase 14, N-{2-fluoro-5-[(5-oxo-6,7,8,9-tetrahydro-5H-benzo[7]annulen-2-yl)amino]phenyl}benzamide, octyl beta-D-glucopyranoside | | Authors: | Mayer-Wrangowski, S.C, Richters, A, Gruetter, C, Rauh, D. | | Deposit date: | 2011-11-30 | | Release date: | 2012-11-07 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Targeting the Hinge Glycine Flip and the Activation Loop: Novel Approach to Potent p38 alpha Inhibitors.
J.Med.Chem., 55, 2012
|
|
4IPU
 
 | |
4NOA
 
 | |
4NPH
 
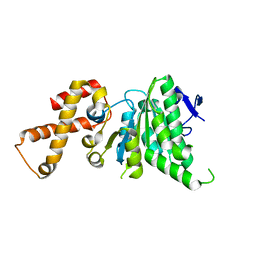 | | Crystal structure of SsaN from Salmonella enterica | | Descriptor: | Probable secretion system apparatus ATP synthase SsaN | | Authors: | Sugiman-Marangos, S.N, Zhang, K, Cooper, C.A, Coombes, B.K, Junop, M.S. | | Deposit date: | 2013-11-21 | | Release date: | 2014-07-23 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Crystal structure of SsaN from Salmonella enterica
J.Biol.Chem., 2014
|
|
4HQB
 
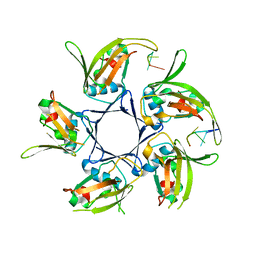 | |
4MBQ
 
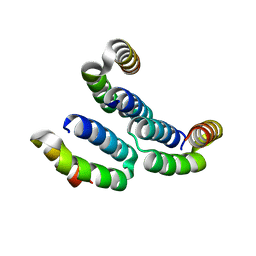 | | TPR3 of FimV from P. aeruginosa (PAO1) | | Descriptor: | Motility protein FimV | | Authors: | Nguyen, Y, Zhang, K, Daniel-Ivad, M, Robinson, H, Wolfram, F, Sugiman-Marangos, S.N, Junop, M.S, Burrows, L.L, Howell, P.L. | | Deposit date: | 2013-08-19 | | Release date: | 2014-08-20 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.006 Å) | | Cite: | Crystal structure of TPR2 from FimV
To be Published
|
|
4NOE
 
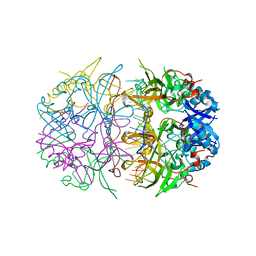 | | Crystal structure of DdrB bound to 30b ssDNA | | Descriptor: | 5'-D(*TP*TP*GP*CP*GP*CP*TP*TP*GP*CP*GP*CP*TP*TP*GP*CP*GP*CP*TP*TP*GP*CP*GP*CP*TP*TP*GP*CP*GP*CP)-3', CALCIUM ION, Single-stranded DNA-binding protein DdrB | | Authors: | Sugiman-Marangos, S.N, Weiss, Y.M, Junop, M.S. | | Deposit date: | 2013-11-19 | | Release date: | 2015-05-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mechanism for accurate, protein-assisted DNA annealing by Deinococcus radiodurans DdrB.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
1K1G
 
 | | STRUCTURAL BASIS FOR RECOGNITION OF THE INTRON BRANCH SITE RNA BY SPLICING FACTOR 1 | | Descriptor: | 5'-R(*UP*AP*UP*AP*CP*UP*AP*AP*CP*AP*A)-3', SF1-Bo isoform | | Authors: | Liu, Z, Luyten, I, Bottomley, M.J, Messias, A.C, Houngninou-Molango, S, Sprangers, R, Zanier, K, Kramer, A, Sattler, M. | | Deposit date: | 2001-09-25 | | Release date: | 2001-11-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for recognition of the intron branch site RNA by splicing factor 1.
Science, 294, 2001
|
|
