1QQB
 
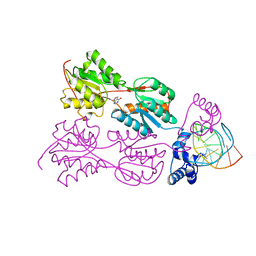 | | PURINE REPRESSOR MUTANT-HYPOXANTHINE-PALINDROMIC OPERATOR COMPLEX | | Descriptor: | 5'-D(*TP*AP*CP*GP*CP*AP*AP*TP*CP*GP*AP*TP*TP*GP*CP*GP*T)-3', HYPOXANTHINE, PURINE NUCLEOTIDE SYNTHESIS REPRESSOR | | Authors: | Glasfeld, A, Koehler, A.N, Schumacher, M.A, Brennan, R.G. | | Deposit date: | 1999-06-01 | | Release date: | 1999-06-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The role of lysine 55 in determining the specificity of the purine repressor for its operators through minor groove interactions.
J.Mol.Biol., 291, 1999
|
|
1G5W
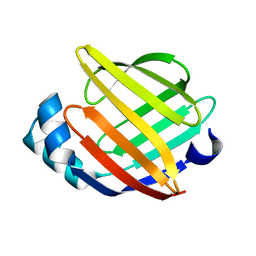 
 | | SOLUTION STRUCTURE OF HUMAN HEART-TYPE FATTY ACID BINDING PROTEIN | | Descriptor: | FATTY ACID-BINDING PROTEIN | | Authors: | Luecke, C, Rademacher, M, Zimmerman, A, van Moerkerk, H.T.B, Veerkamp, J.H, Rueterjans, H. | | Deposit date: | 2000-11-02 | | Release date: | 2001-03-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Spin-system heterogeneities indicate a selected-fit mechanism in fatty acid binding to heart-type fatty acid-binding protein (H-FABP).
Biochem.J., 354, 2001
|
|
4LKV
 
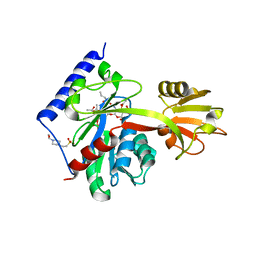 | | Determinants of lipid substrate and membrane binding for the tetraacyldisaccharide-1-phosphate 4 -kinase LpxK | | Descriptor: | (R)-((2R,3S,4R,5R,6R)-3-HYDROXY-2-(HYDROXYMETHYL)-5-((R)-3-HYDROXYTETRADECANAMIDO)-6-(PHOSPHONOOXY)TETRAHYDRO-2H-PYRAN-4-YL) 3-HYDROXYTETRADECANOATE, 2-deoxy-3-O-[(3R)-3-hydroxytetradecanoyl]-2-{[(3R)-3-hydroxytetradecanoyl]amino}-4-O-phosphono-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Emptage, R.P, Tonthat, N.K, York, J.D, Schumacher, M.A, Zhou, P. | | Deposit date: | 2013-07-08 | | Release date: | 2014-07-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.5109 Å) | | Cite: | Structural Basis of Lipid Binding for the Membrane-embedded Tetraacyldisaccharide-1-phosphate 4'-Kinase LpxK.
J.Biol.Chem., 289, 2014
|
|
1PRU
 
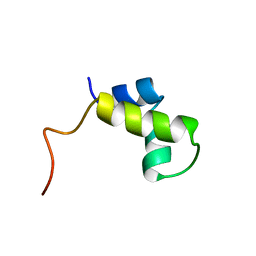 | | PURINE REPRESSOR DNA-BINDING DOMAIN DNA BINDING | | Descriptor: | PURINE REPRESSOR | | Authors: | Nagadoi, A, Morikawa, S, Nakamura, H, Enari, M, Kobayashi, K, Yamamoto, H, Sampei, G, Mizobuchi, K, Schumacher, M.A, Brennan, R.G, Nishimura, Y. | | Deposit date: | 1995-05-08 | | Release date: | 1996-03-08 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural comparison of the free and DNA-bound forms of the purine repressor DNA-binding domain.
Structure, 3, 1995
|
|
1PRV
 
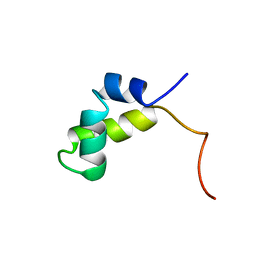 | | PURINE REPRESSOR DNA-BINDING DOMAIN DNA BINDING | | Descriptor: | PURINE REPRESSOR | | Authors: | Nagadoi, A, Morikawa, S, Nakamura, H, Enari, M, Kobayashi, K, Yamamoto, H, Sampei, G, Mizobuchi, K, Schumacher, M.A, Brennan, R.G, Nishimura, Y. | | Deposit date: | 1995-05-08 | | Release date: | 1996-03-08 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural comparison of the free and DNA-bound forms of the purine repressor DNA-binding domain.
Structure, 3, 1995
|
|
4PUR
 
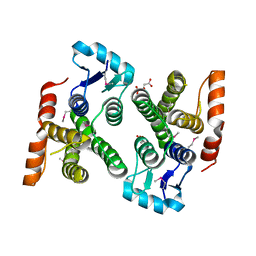 | | Crystal structure of MglA from Francisella tularensis | | Descriptor: | D-MALATE, Macrophage growth locus, subunit A | | Authors: | Cuthbert, B.J, Schumacher, M.A, Brennan, R.G. | | Deposit date: | 2014-03-13 | | Release date: | 2015-06-24 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural and Biochemical Characterization of the Francisella tularensis Pathogenicity Regulator, Macrophage Locus Protein A (MglA).
Plos One, 10, 2015
|
|
1JJJ
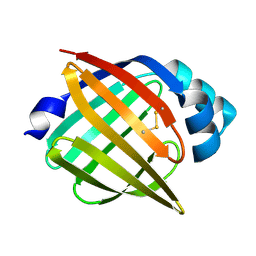 
 | | SOLUTION STRUCTURE OF RECOMBINANT HUMAN EPIDERMAL-TYPE FATTY ACID BINDING PROTEIN | | Descriptor: | EPIDERMAL-TYPE FATTY ACID BINDING PROTEIN (E-FABP) | | Authors: | Gutierrez-Gonzalez, L.H, Ludwig, C, Hohoff, C, Rademacher, M, Hanhoff, T, Rueterjans, H, Spener, F, Luecke, C. | | Deposit date: | 2001-07-06 | | Release date: | 2002-06-19 | | Last modified: | 2024-11-06 | | Method: | SOLUTION NMR | | Cite: | Solution structure and backbone dynamics of human epidermal-type fatty
acid-binding protein (E-FABP)
BIOCHEM.J., 364, 2002
|
|
4PYA
 
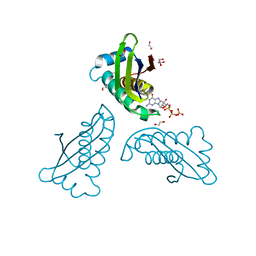 | | MoaC K51A in complex with 3',8-cH2GTP | | Descriptor: | (8S)-3',8-cyclo-7,8-dihydroguanosine 5'-triphosphate, 1,2-ETHANEDIOL, Molybdenum cofactor biosynthesis protein MoaC | | Authors: | Tonthat, N.K, Hover, B.M, Yokoyama, K, Schumacher, M.A. | | Deposit date: | 2014-03-26 | | Release date: | 2015-04-29 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.789 Å) | | Cite: | Mechanism of pyranopterin ring formation in molybdenum cofactor biosynthesis.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4PYD
 
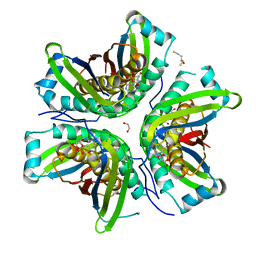 | | MoaC in complex with cPMP crystallized in space group P212121 | | Descriptor: | (2R,4AR,5AR,11AR,12AS)-8-AMINO-2-HYDROXY-4A,5A,9,11,11A,12A-HEXAHYDRO[1,3,2]DIOXAPHOSPHININO[4',5':5,6]PYRANO[3,2-G]PTERIDINE-10,12(4H,6H)-DIONE 2-OXIDE, 1,2-ETHANEDIOL, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, ... | | Authors: | Tonthat, N.K, Hover, B.M, Yokoyama, K, Schumacher, M.A. | | Deposit date: | 2014-03-26 | | Release date: | 2015-04-29 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.186 Å) | | Cite: | Mechanism of pyranopterin ring formation in molybdenum cofactor biosynthesis.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
1RKW
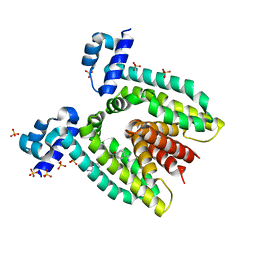 
 | | CRYSTAL STRUCTURE OF THE MULTIDRUG BINDING TRANSCRIPTIONAL REPRESSOR QACR BOUND TO PENTAMADINE | | Descriptor: | 1,5-BIS(4-AMIDINOPHENOXY)PENTANE, SULFATE ION, Transcriptional regulator qacR | | Authors: | Murray, D.S, Schumacher, M.A, Brennan, R.G. | | Deposit date: | 2003-11-23 | | Release date: | 2004-06-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.62 Å) | | Cite: | Crystal structures of QacR-diamidine complexes reveal additional multidrug-binding modes and a novel mechanism of drug charge neutralization.
J.Biol.Chem., 279, 2004
|
|
1RPW
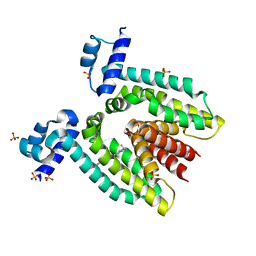 
 | | Crystal Structure Of The Multidrug Binding Protein Qacr Bound To The Diamidine Hexamidine | | Descriptor: | 4,4'[1,6-HEXANEDIYLBIS(OXY)]BISBENZENECARBOXIMIDAMIDE, SULFATE ION, Transcriptional regulator qacR | | Authors: | Murray, D.S, Schumacher, M.A, Brennan, R.G. | | Deposit date: | 2003-12-03 | | Release date: | 2004-06-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structures of QacR-diamidine complexes reveal additional multidrug-binding modes and a novel mechanism of drug charge neutralization.
J.Biol.Chem., 279, 2004
|
|
