6P83
 | | E.coli LpxD in complex with compound 1o | | Descriptor: | 4-(2-chlorophenyl)-3-hydroxy-7,7-dimethyl-2-phenyl-2,6,7,8-tetrahydro-5H-pyrazolo[3,4-b]quinolin-5-one, MAGNESIUM ION, UDP-3-O-(3-hydroxymyristoyl)glucosamine N-acyltransferase | | Authors: | Ma, X, Shia, S. | | Deposit date: | 2019-06-06 | | Release date: | 2019-08-28 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and Biological Basis of Small Molecule Inhibition ofEscherichia coliLpxD Acyltransferase Essential for Lipopolysaccharide Biosynthesis.
Acs Infect Dis., 6, 2020
|
|
6P8A
 
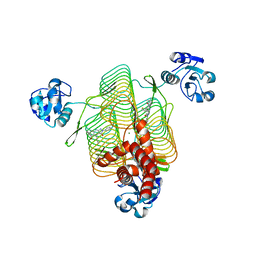 | | E.coli LpxD in complex with compound 8.1 | | Descriptor: | 2-[4,6-dimethyl-3-(1H-pyrrol-1-yl)-1H-pyrazolo[3,4-b]pyridin-1-yl]-N-[3-(morpholin-4-yl)propyl]acetamide, 3-hydroxy-7,7-dimethyl-2-phenyl-4-(thiophen-2-yl)-2,6,7,8-tetrahydro-5H-pyrazolo[3,4-b]quinolin-5-one, MAGNESIUM ION, ... | | Authors: | Ma, X, Shia, S. | | Deposit date: | 2019-06-06 | | Release date: | 2019-08-28 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Biological Basis of Small Molecule Inhibition ofEscherichia coliLpxD Acyltransferase Essential for Lipopolysaccharide Biosynthesis.
Acs Infect Dis., 6, 2020
|
|
4EUU
 
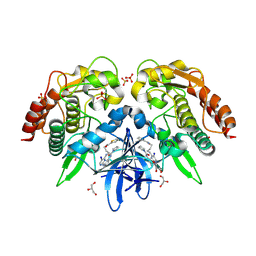 | | Structure of BX-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on Ser172 | | Descriptor: | GLYCEROL, IODIDE ION, N-(3-{[5-iodo-4-({3-[(thiophen-2-ylcarbonyl)amino]propyl}amino)pyrimidin-2-yl]amino}phenyl)pyrrolidine-1-carboxamide, ... | | Authors: | Ma, X, Helgason, E, Phung, Q.T, Quan, C.L, Iyer, R.S, Lee, M.W, Bowman, K.K, Starovasnik, M.A, Dueber, E.C. | | Deposit date: | 2012-04-25 | | Release date: | 2012-05-23 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Molecular basis of Tank-binding kinase 1 activation by transautophosphorylation.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4EUT
 
 | | Structure of BX-795 Complexed with Unphosphorylated Human TBK1 Kinase-ULD Domain | | Descriptor: | IODIDE ION, N-(3-{[5-iodo-4-({3-[(thiophen-2-ylcarbonyl)amino]propyl}amino)pyrimidin-2-yl]amino}phenyl)pyrrolidine-1-carboxamide, SULFATE ION, ... | | Authors: | Ma, X, Helgason, E, Phung, Q.T, Quan, C.L, Iyer, R.S, Lee, M.W, Bowman, K.K, Starovasnik, M.A, Dueber, E.C. | | Deposit date: | 2012-04-25 | | Release date: | 2012-05-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Molecular basis of Tank-binding kinase 1 activation by transautophosphorylation.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3TMP
 
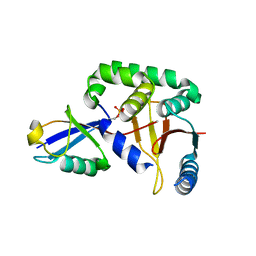 | | The catalytic domain of human deubiquitinase DUBA in complex with ubiquitin aldehyde | | Descriptor: | OTU domain-containing protein 5, Polyubiquitin-C | | Authors: | Ma, X, Yin, J, Hymowitz, S, Starovasnik, M, Cochran, A. | | Deposit date: | 2011-08-31 | | Release date: | 2012-01-11 | | Last modified: | 2012-02-29 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Phosphorylation-dependent activity of the deubiquitinase DUBA.
Nat.Struct.Mol.Biol., 19, 2012
|
|
6IZT
 
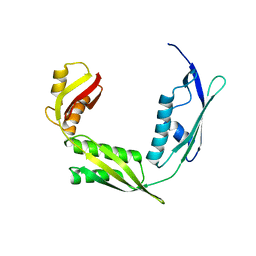 | | Crystal structure of Haemophilus Influenzae BamA POTRA3-5 | | Descriptor: | Outer membrane protein assembly factor BamA | | Authors: | Ma, X, Wang, Q, Li, Y, Tan, P, Wu, H, Wang, P, Dong, X, Hong, L, Meng, G. | | Deposit date: | 2018-12-20 | | Release date: | 2019-10-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | How BamA recruits OMP substratesviapoly-POTRAs domain.
Faseb J., 33, 2019
|
|
6IZS
 | | Crystal structure of Haemophilus influenzae BamA POTRA4 | | Descriptor: | Outer membrane protein assembly factor BamA | | Authors: | Ma, X, Wang, Q, Li, Y, Tan, P, Wu, H, Wang, P, Dong, X, Hong, L, Meng, G. | | Deposit date: | 2018-12-20 | | Release date: | 2019-10-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | How BamA recruits OMP substratesviapoly-POTRAs domain.
Faseb J., 33, 2019
|
|
6J09
 
 | | Crystal structure of Haemophilus Influenzae BamA POTRA1-4 | | Descriptor: | Outer membrane protein assembly factor BamA | | Authors: | Ma, X, Wang, Q, Li, Y, Tan, P, Wu, H, Wang, P, Dong, X, Hong, L, Meng, G. | | Deposit date: | 2018-12-21 | | Release date: | 2019-10-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | How BamA recruits OMP substratesviapoly-POTRAs domain.
Faseb J., 33, 2019
|
|
2O09
 
 | |
2O0G
 
 | |
2O0C
 
 | |
3HLS
 
 | |
3P9W
 
 | |
5BVE
 
 | | Tetrahydropyrrolo-diazepenones as inhibitors of ERK2 kinase | | Descriptor: | 2-[(1S)-1-(3-chlorophenyl)-2-hydroxyethyl]-8-[2-(tetrahydro-2H-pyran-4-ylamino)pyrimidin-4-yl]-2,3,4,5-tetrahydro-1H-pyrrolo[1,2-a][1,4]diazepin-1-one, Mitogen-activated protein kinase 1, SULFATE ION | | Authors: | Ma, X, Steven, S. | | Deposit date: | 2015-06-05 | | Release date: | 2015-09-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Tetrahydropyrrolo-diazepenones as inhibitors of ERK2 kinase.
Bioorg.Med.Chem.Lett., 25, 2015
|
|
5BVF
 
 | | Tetrahydropyrrolo-diazepenones as inhibitors of ERK2 kinase | | Descriptor: | 2-[(1S)-1-(3-chlorophenyl)-2-hydroxyethyl]-8-(2-{[(1S,3R)-3-hydroxycyclopentyl]amino}pyrimidin-4-yl)-2,3,4,5-tetrahydro-1H-pyrrolo[1,2-a][1,4]diazepin-1-one, Mitogen-activated protein kinase 1, SULFATE ION | | Authors: | Ma, X, Steven, S. | | Deposit date: | 2015-06-05 | | Release date: | 2015-09-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Tetrahydropyrrolo-diazepenones as inhibitors of ERK2 kinase.
Bioorg.Med.Chem.Lett., 25, 2015
|
|
4K3J
 
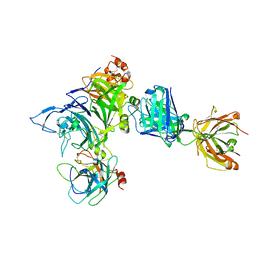 | | Crystal structure of Onartuzumab Fab in complex with MET and HGF-beta | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Hepatocyte growth factor, Hepatocyte growth factor beta chain, ... | | Authors: | Ma, X, Starovasnik, M.A. | | Deposit date: | 2013-04-10 | | Release date: | 2013-08-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Monovalent antibody design and mechanism of action of onartuzumab, a MET antagonist with anti-tumor activity as a therapeutic agent.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
5BVD
 
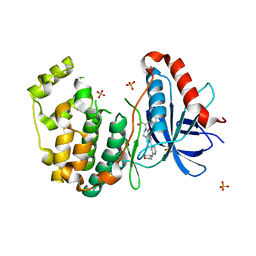 | | Tetrahydropyrrolo-diazepenones as inhibitors of ERK2 kinase | | Descriptor: | 2-[(1S)-1-(3-chlorophenyl)-2-hydroxyethyl]-7-[2-(tetrahydro-2H-pyran-4-ylamino)pyrimidin-4-yl]-3,4-dihydropyrrolo[1,2-a]pyrazin-1(2H)-one, Mitogen-activated protein kinase 1, SULFATE ION | | Authors: | Ma, X, Steven, S. | | Deposit date: | 2015-06-05 | | Release date: | 2015-09-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Tetrahydropyrrolo-diazepenones as inhibitors of ERK2 kinase.
Bioorg.Med.Chem.Lett., 25, 2015
|
|
7XT6
 
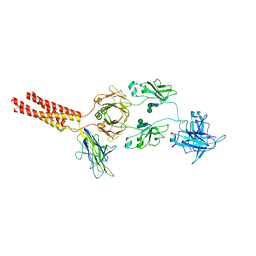 | | Structure of a membrane protein M3 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, B-cell antigen receptor complex-associated protein alpha chain, ... | | Authors: | Ma, X, Zhu, Y, Chen, Y, Huang, Z. | | Deposit date: | 2022-05-16 | | Release date: | 2022-08-31 | | Method: | ELECTRON MICROSCOPY (3.63 Å) | | Cite: | Cryo-EM structures of two human B cell receptor isotypes.
Science, 377, 2022
|
|
5WL0
 
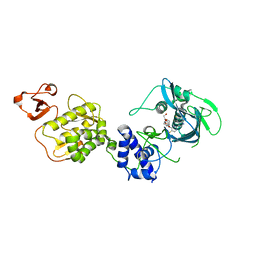 | | Co-crystal structure of Influenza A H3N2 PB2 (241-741) bound to VX-787 | | Descriptor: | (2S,3S)-3-[[5-fluoranyl-2-(5-fluoranyl-1H-pyrrolo[2,3-b]pyridin-3-yl)pyrimidin-4-yl]amino]bicyclo[2.2.2]octane-2-carboxylic acid, DI(HYDROXYETHYL)ETHER, Polymerase basic protein 2 | | Authors: | Ma, X, Shia, S. | | Deposit date: | 2017-07-25 | | Release date: | 2017-09-06 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for therapeutic inhibition of influenza A polymerase PB2 subunit.
Sci Rep, 7, 2017
|
|
5Z55
 
 | |
7Y47
 
 | | Crystal structure of bifunctional miltiradiene synthase from selaginella moellendorffii that complexed with GGPP | | Descriptor: | Bifunctional diterpene synthase, chloroplastic, GERANYLGERANYL DIPHOSPHATE, ... | | Authors: | Ma, X, Tao, Y, Jiang, T. | | Deposit date: | 2022-06-14 | | Release date: | 2022-10-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and mechanistic insights into the precise product synthesis by a bifunctional miltiradiene synthase.
Plant Biotechnol J, 21, 2023
|
|
7WSO
 | | Structure of a membrane protein G | | Descriptor: | B-cell antigen receptor complex-associated protein alpha chain, B-cell antigen receptor complex-associated protein beta chain, Immunoglobulin heavy constant gamma 1 | | Authors: | Ma, X, Zhu, Y, Chen, Y, Huang, Z. | | Deposit date: | 2022-01-30 | | Release date: | 2022-08-31 | | Method: | ELECTRON MICROSCOPY (3.03 Å) | | Cite: | Cryo-EM structures of two human B cell receptor isotypes.
Science, 377, 2022
|
|
7U8F
 
 | | Ternary complex structure of Cereblon-DDB1 bound to IKZF2(ZF2) and the molecular glue DKY709 | | Descriptor: | (3S)-3-[5-(1-benzylpiperidin-4-yl)-1-oxo-1,3-dihydro-2H-isoindol-2-yl]piperidine-2,6-dione, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, DNA damage-binding protein 1, ... | | Authors: | Ma, X, Ornelas, E, Clifton, M.C. | | Deposit date: | 2022-03-08 | | Release date: | 2023-03-08 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Discovery and characterization of a selective IKZF2 glue degrader for cancer immunotherapy.
Cell Chem Biol, 30, 2023
|
|
7WAT
 
 | |
7WSP
 
 | | Structure of a membrane protein M | | Descriptor: | B-cell antigen receptor complex-associated protein alpha chain, B-cell antigen receptor complex-associated protein beta chain, Isoform 2 of Immunoglobulin heavy constant mu | | Authors: | Ma, X, Zhu, Y, Chen, Y, Huang, Z. | | Deposit date: | 2022-01-30 | | Release date: | 2023-02-01 | | Method: | ELECTRON MICROSCOPY (4.09 Å) | | Cite: | Structure of a membrane protein M
To Be Published
|
|
