2AGQ
 
 | | Fidelity of Dpo4: effect of metal ions, nucleotide selection and pyrophosphorolysis | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, 5'-D(*GP*GP*CP*TP*AP*CP*AP*GP*GP*AP*CP*TP*(DOC))-3', 5'-D(*TP*CP*AP*TP*GP*AP*GP*TP*CP*CP*TP*GP*TP*AP*GP*CP*C)-3', ... | | Authors: | Ling, H, Yang, W. | | Deposit date: | 2005-07-27 | | Release date: | 2005-09-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Fidelity of Dpo4: effect of metal ions, nucleotide selection and pyrophosphorolysis.
Embo J., 24, 2005
|
|
2AGO
 
 | | Fidelity of Dpo4: effect of metal ions, nucleotide selection and pyrophosphorolysis | | Descriptor: | CALCIUM ION, DNA (5'-D(*GP*GP*GP*GP*GP*AP*AP*GP*GP*AP*TP*TP*CP*G)-3'), DNA (5'-D(*TP*TP*TP*TP*GP*AP*AP*TP*CP*CP*TP*TP*CP*CP*CP*CP*C)-3'), ... | | Authors: | Ling, H, Yang, W. | | Deposit date: | 2005-07-27 | | Release date: | 2005-09-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Fidelity of Dpo4: effect of metal ions, nucleotide selection and pyrophosphorolysis.
Embo J., 24, 2005
|
|
2AGP
 
 | | Fidelity of Dpo4: effect of metal ions, nucleotide selection and pyrophosphorolysis | | Descriptor: | 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, CALCIUM ION, DNA (5'-D(*GP*GP*GP*GP*GP*AP*AP*GP*GP*AP*TP*TP*(DOC)-3'), ... | | Authors: | Ling, H, Yang, W. | | Deposit date: | 2005-07-27 | | Release date: | 2005-09-06 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Fidelity of Dpo4: effect of metal ions, nucleotide selection and pyrophosphorolysis.
Embo J., 24, 2005
|
|
1N48
 
 | | Y-family DNA polymerase Dpo4 in complex with DNA containing abasic lesion | | Descriptor: | 5'-D(*CP*AP*(3DR)P*TP*AP*GP*TP*CP*CP*TP*TP*CP*CP*CP*CP*C)-3', 5'-D(*GP*GP*GP*GP*GP*AP*AP*GP*GP*AP*CP*TP*A)-3', ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Ling, H, Boudsocq, F, Woodgate, R, Yang, W. | | Deposit date: | 2002-10-30 | | Release date: | 2004-02-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A Y-family polymerase complexed with abasic lesions: catching DNA with a loaded nucleoside triphosphate
To be Published
|
|
2IMW
 
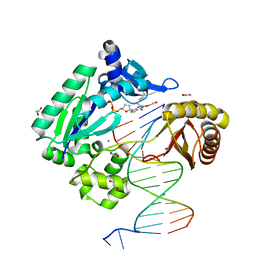 | | Mechanism of Template-Independent Nucleotide Incorporation Catalyzed by a Template-Dependent DNA Polymerase | | Descriptor: | 1,2-ETHANEDIOL, 2',3'-dideoxyadenosine triphosphate, 5'-D(*GP*GP*GP*GP*GP*AP*AP*GP*GP*AP*TP*TP*C)-3', ... | | Authors: | Ling, H, Yang, W. | | Deposit date: | 2006-10-05 | | Release date: | 2007-01-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Mechanism of Template-independent Nucleotide Incorporation Catalyzed by a Template-dependent DNA Polymerase.
J.Mol.Biol., 365, 2007
|
|
1N56
 
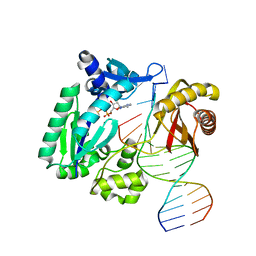 | | Y-family DNA polymerase Dpo4 in complex with DNA containing abasic lesion | | Descriptor: | 5'-D(*GP*GP*GP*GP*GP*AP*AP*GP*GP*AP*CP*TP*AP*A)-3', 5'-D(*TP*CP*AP*TP*(3DR)P*AP*GP*TP*CP*CP*TP*TP*CP*CP*CP*CP*C)-3', ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Ling, H, Boudsocq, F, Woodgate, R, Yang, W. | | Deposit date: | 2002-11-04 | | Release date: | 2004-02-24 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Snapshots of replication through an abasic lesion; structural basis for base substitutions and frameshifts.
Mol.Cell, 13, 2004
|
|
2B5E
 
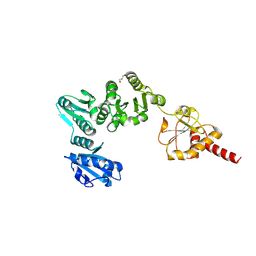 | | Crystal Structure of Yeast Protein Disulfide Isomerase | | Descriptor: | BARIUM ION, GLYCEROL, Protein disulfide-isomerase | | Authors: | Schindelin, H, Tian, G. | | Deposit date: | 2005-09-28 | | Release date: | 2006-01-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The crystal structure of yeast protein disulfide isomerase suggests cooperativity between its active sites.
Cell(Cambridge,Mass.), 124, 2006
|
|
1ORL
 
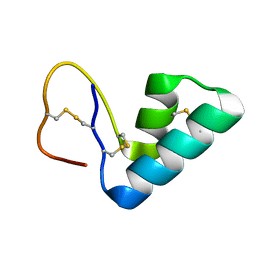 | | 1H NMR structure determination of Viscotoxin C1 | | Descriptor: | Viscotoxin C1 | | Authors: | Molinari, H, Romagnoli, S, Fogolari, F, Catalano, M, Urech, K, Giannattasio, M, Ragona, L. | | Deposit date: | 2003-03-14 | | Release date: | 2003-04-01 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of viscotoxin C1 from Viscum album species Coloratum ohwi: toward a structure-function analysis of viscotoxins.
Biochemistry, 42, 2003
|
|
1P9H
 
 | |
1Q7L
 
 | | Zn-binding domain of the T347G mutant of human aminoacylase-I | | Descriptor: | Aminoacylase-1, GLYCINE, ZINC ION | | Authors: | Lindner, H.A, Lunin, V.V, Alary, A, Hecker, R, Cygler, M, Menard, R. | | Deposit date: | 2003-08-19 | | Release date: | 2004-01-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Essential roles of zinc ligation and enzyme dimerization for catalysis in the aminoacylase-1/M20 family.
J.Biol.Chem., 278, 2003
|
|
1CSP
 | |
1D1I
 
 | |
1D1K
 
 | |
1CQF
 
 | |
1EKS
 
 | | ASP128ALA VARIANT OF MOAC PROTEIN FROM E. COLI | | Descriptor: | L(+)-TARTARIC ACID, MOLYBDENUM COFACTOR BIOSYNTHESIS PROTEIN C | | Authors: | Schindelin, H, Liu, M.T.W, Wuebbens, M.M, Rajagopalan, K.V. | | Deposit date: | 2000-03-09 | | Release date: | 2000-09-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Insights into molybdenum cofactor deficiency provided by the crystal structure of the molybdenum cofactor biosynthesis protein MoaC.
Structure Fold.Des., 8, 2000
|
|
1EKR
 
 | | MOAC PROTEIN FROM E. COLI | | Descriptor: | MOLYBDENUM COFACTOR BIOSYNTHESIS PROTEIN C | | Authors: | Schindelin, H, Liu, M.T.W, Wuebbens, M.M, Rajagopalan, K.V. | | Deposit date: | 2000-03-09 | | Release date: | 2000-09-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insights into molybdenum cofactor deficiency provided by the crystal structure of the molybdenum cofactor biosynthesis protein MoaC.
Structure Fold.Des., 8, 2000
|
|
5H9S
 
 | | Crystal Structure of Human Galectin-7 in Complex with TAZTDG | | Descriptor: | (2~{S},3~{R},4~{S},5~{R},6~{R})-2-[(2~{S},3~{R},4~{S},5~{R},6~{R})-4-[4-(3-fluorophenyl)-1,2,3-triazol-1-yl]-6-(hydroxymethyl)-3,5-bis(oxidanyl)oxan-2-yl]sulfanyl-6-(hydroxymethyl)oxane-3,4,5-triol, Galectin-7 | | Authors: | Hsieh, T.J, Lin, H.Y, Lin, C.H. | | Deposit date: | 2015-12-29 | | Release date: | 2016-06-29 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.821 Å) | | Cite: | Dual thio-digalactoside-binding modes of human galectins as the structural basis for the design of potent and selective inhibitors
Sci Rep, 6, 2016
|
|
5H9Q
 
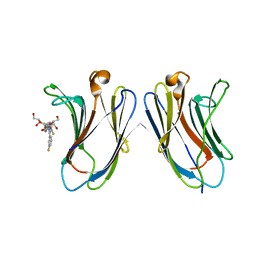 | | Crystal Structure of Human Galectin-7 in Complex with TD139 | | Descriptor: | 3-deoxy-3-[4-(3-fluorophenyl)-1H-1,2,3-triazol-1-yl]-beta-D-galactopyranosyl 3-deoxy-3-[4-(3-fluorophenyl)-1H-1,2,3-triazol-1-yl]-1-thio-beta-D-galactopyranoside, Galectin-7 | | Authors: | Hsieh, T.J, Lin, H.Y, Lin, C.H. | | Deposit date: | 2015-12-29 | | Release date: | 2016-06-29 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.931 Å) | | Cite: | Dual thio-digalactoside-binding modes of human galectins as the structural basis for the design of potent and selective inhibitors
Sci Rep, 6, 2016
|
|
5GGE
 | | Fatty Acid-Binding Protein in Brain Tissue of Drosophila melanogaster | | Descriptor: | CITRIC ACID, Fatty acid bindin protein, isoform B | | Authors: | Cheng, Y.-Y, Huang, Y.-F, Lin, H.-H, Chang, W.W, Lyu, P.-C. | | Deposit date: | 2016-06-15 | | Release date: | 2017-06-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.861 Å) | | Cite: | The ligand-mediated affinity of brain-type fatty acid-binding protein for membranes determines the directionality of lipophilic cargo transport.
Biochim Biophys Acta Mol Cell Biol Lipids, 1864, 2019
|
|
9CBT
 
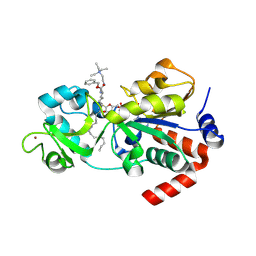 | | Crystal structure of human sirtuin 3 fragment (residues 118-399) bound to intermediates from reaction with NAD and inhibitor NH6-10 | | Descriptor: | (phenylmethyl) ~{N}-[(2~{S})-6-[[(2~{R},3~{a}~{R},5~{R},6~{R},6~{a}~{R})-5-[[[[(2~{R},3~{S},4~{R},5~{R})-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl]oxy-oxidanyl-phosphoryl]oxymethyl]-6-oxidanyl-2-tridecyl-3~{a},5,6,6~{a}-tetrahydrofuro[2,3-d][1,3]oxathiol-2-yl]amino]-1-oxidanylidene-1-[2-(triethyl-$l^{4}-azanyl)ethylamino]hexan-2-yl]carbamate, 2-{[(2S)-6-[(Z)-(1-{[(2R,3R,4R,5R)-5-({[(R)-{[(R)-{[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}methyl)-4-hydroxy-2-sulfanyloxolan-3-yl]oxy}tetradecylidene)amino]-2-{[(benzyloxy)carbonyl]amino}hexanoyl]amino}-N,N,N-triethylethan-1-aminium (non-preferred name), NAD-dependent protein deacetylase sirtuin-3, ... | | Authors: | Fenwick, M.K, Young, H.J, Lin, H. | | Deposit date: | 2024-06-20 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of human sirtuin 3 fragment (residues 118-399) bound to intermediates from reaction with NAD and inhibitor NH6-10
To Be Published
|
|
6DO4
 
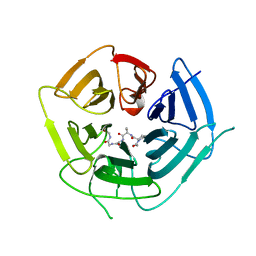 | | KLHDC2 ubiquitin ligase in complex with SelS C-end degron | | Descriptor: | Kelch domain-containing protein 2, SELS C-END DEGRON | | Authors: | Rusnac, D.V, Lin, H.C, Yen, H.C.S, Zheng, N. | | Deposit date: | 2018-06-08 | | Release date: | 2018-12-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Recognition of the Diglycine C-End Degron by CRL2KLHDC2Ubiquitin Ligase.
Mol. Cell, 72, 2018
|
|
6DO3
 
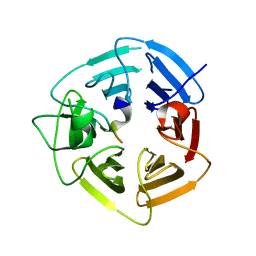 | | KLHDC2 ubiquitin ligase in complex with SelK C-end degron | | Descriptor: | Kelch domain-containing protein 2, SelK C-end Degron | | Authors: | Rusnac, D.V, Lin, H.C, Yen, H.C.S, Zheng, N. | | Deposit date: | 2018-06-08 | | Release date: | 2018-12-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.165 Å) | | Cite: | Recognition of the Diglycine C-End Degron by CRL2KLHDC2Ubiquitin Ligase.
Mol. Cell, 72, 2018
|
|
8WQM
 
 | | Complex structure of AtHPPD with Atovaquone | | Descriptor: | 2-[trans-4-(4-chlorophenyl)cyclohexyl]-3-hydroxynaphthalene-1,4-dione, 4-hydroxyphenylpyruvate dioxygenase, COBALT (II) ION | | Authors: | Yang, G.-F, Lin, H.-Y, Dong, J. | | Deposit date: | 2023-10-11 | | Release date: | 2024-03-13 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.796 Å) | | Cite: | Complex structure of AtHPPD with Atovaquone
To Be Published
|
|
6JZ7
 
 | |
6JZ6
 
 | | b-glucuronidase from Ruminococcus gnavus in complex with C6-substituted uronic isofagomine | | Descriptor: | (2~{S},3~{S},4~{R},5~{R})-4,5-bis(oxidanyl)-2-propyl-piperidine-3-carboxylic acid, Beta-glucuronidase | | Authors: | Dashnyam, P, Lin, H.Y. | | Deposit date: | 2019-04-30 | | Release date: | 2020-06-10 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.605 Å) | | Cite: | Substituent Position of Iminocyclitols Determines the Potency and Selectivity for Gut Microbial Xenobiotic-Reactivating Enzymes.
J.Med.Chem., 63, 2020
|
|
